Abstract
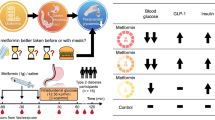
Oral glucose tolerance tests (OGTT) are valuable tools for diagnosis of impaired glucose tolerance and insulin resistance. Protein intake modulates insulin secretion and sensitivity and thereby can alter the metabolic response. To assess these effects in the plasma metabolome, 15 healthy human volunteers underwent in random order an OGTT with and without 30 g of a protein-hydrolysate (OGTT + PL) followed by metabolite profiling via LC–MS/MS. Compared to the OGTT alone, plasma glucose response was reduced and insulin clearance retarded after OGTT + PL while suppression of catabolic markers such as free fatty acids, glycerol, lactate or acylcarnitines showed no difference between the two challenges. Clusters of plasma amino acids characterized by subgroups with similar physico-chemical characteristics revealed long-lasting declines after the OGTT and coherent increases after the OGTT + PL despite the fact that the protein-hydrolysate provided different amounts of individual amino acids. The amino acid clusters identified appear to derive from common transport processes involved in intestinal absorption and the insulin-dependent extraction by target tissues and suggest that these processes may be of high diagnostic value to assess insulin sensitivity.





Similar content being viewed by others
References
Adibi, S. A., Gray, S. J., & Menden, E. (1967). The kinetics of amino acid absorption and alteration of plasma composition of free amino acids after intestinal perfusion of amino acid mixtures. American Journal of Clinical Nutrition, 20, 24–33.
Baggio, L. L., & Drucker, D. J. (2007). Biology of incretins: GLP-1 and GIP. Gastroenterology, 132, 2131–2157.
Blazquez, R., & Lopez Quijada, C. (1970). The effect of a high-protein diet on plasma glucose concentration, insulin sensitivity and plasma insulin in rats. Journal of Endocrinology, 46, 445–451.
Deo, R. C., Hunter, L., Lewis, G. D., Pare, G., Vasan, R. S., Chasman, D., et al. (2010). Interpreting metabolomic profiles using unbiased pathway models. PLoS Computational Biology, 6, e1000692.
Felig, P., Marliss, E., Pozefsky, T., & Cahill, G. F., Jr. (1970). Amino acid metabolism in the regulation of gluconeogenesis in man. American Journal of Clinical Nutrition, 23, 986–992.
Floyd, J. C., Jr., Fajans, S. S., Conn, J. W., Knopf, R. F., & Rull, J. (1966). Stimulation of insulin secretion by amino acids. Journal of Clinical Investigation, 45, 1487–1502.
Fukuhara, D., Kanai, Y., Chairoungdua, A., Babu, E., Bessho, F., Kawano, T., et al. (2007). Protein characterization of NA+-independent system l amino acid transporter 3 in mice: A potential role in supply of branched-chain amino acids under nutrient starvation. American Journal of Pathology, 170, 888–898.
Gual, P., Le Marchand-Brustel, Y., & Tanti, J. F. (2005). Positive and negative regulation of insulin signaling through IRS-1 phosphorylation. Biochimie, 87, 99–109.
Harrell, F. J. (2009). Hmisc: Harrel Miscellaneous, http://CRAN.R-project.org/package=Hmisc
Hatanaka, T., Hatanaka, Y., Tsuchida, J., Ganapathy, V., & Setou, M. (2006). Amino acid transporter ATA2 is stored at the trans-Golgi network and released by insulin stimulus in adipocytes. Journal of Biological Chemistry, 281, 39273–39284.
Holl, M. G., & Allen, L. H. (1988). Comparative effects of meals high in protein, sucrose, or starch on human mineral metabolism and insulin secretion. American Journal of Clinical Nutrition, 48, 1219–1225.
Holst, J. J. (2007). The physiology of glucagon-like peptide 1. Physiological Reviews, 87, 1409–1439.
Husson, F., Josse, J., Le, S., & Mazet, J. (2009). FactoMineR: Factor analysis and data mining with R. R package version 1.12, URL http://CRAN.R-project.org/package=FactoMineR
Jaki, T., & Wolfsegger, M. J. (2009). A theoretical framework for estimation of AUCs in complete and incomplete sampling designs. Statistics in Biopharmaceutical Research, 1, 176–184.
Kashiwagi, H., Yamazaki, K., Takekuma, Y., Ganapathy, V., & Sugawara, M. (2009). Regulatory mechanisms of SNAT2, an amino acid transporter, in L6 rat skeletal muscle cells by insulin, osmotic shock and amino acid deprivation. Amino Acids, 36, 219–230.
Manders, R. J., Wagenmakers, A. J., Koopman, R., Zorenc, A. H., Menheere, P. P., Schaper, N. C., et al. (2005). Co-ingestion of a protein hydrolysate and amino acid mixture with carbohydrate improves plasma glucose disposal in patients with type 2 diabetes. American Journal of Clinical Nutrition, 82, 76–83.
Meier, C., Ristic, Z., Klauser, S., & Verrey, F. (2002). Activation of system L heterodimeric amino acid exchangers by intracellular substrates. EMBO Journal, 21, 580–589.
Nobukuni, T., Joaquin, M., Roccio, M., Dann, S. G., Kim, S. Y., Gulati, P., et al. (2005). Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proceedings of the National Academy of Sciences of the United States of America, 102, 14238–14243.
Palacin, M., Estevez, R., Bertran, J., & Zorzano, A. (1998). Molecular biology of mammalian plasma membrane amino acid transporters. Physiological Reviews, 78, 969–1054.
R Development Core Team. (2009). R: A language and environment for statistical computing. Vienna. Austria: R Foundation for Statistical Computing.
Roberge, J. N., & Brubaker, P. L. (1991). Secretion of proglucagon-derived peptides in response to intestinal luminal nutrients. Endocrinology, 128, 3169–3174.
Sarkar, D. (2008). Lattice: Multivariate data visualization with R. New York: Springer.
Schulze, M. B., Weikert, C., Pischon, T., Bergmann, M. M., Al-Hasani, H., Schleicher, E., et al. (2009). Use of multiple metabolic and genetic markers to improve the prediction of type 2 diabetes: The EPIC-Potsdam Study. Diabetes Care, 32, 2116–2119.
Shaham, O., Wei, R., Wang, T. J., Ricciardi, C., Lewis, G. D., Vasan, R. S., et al. (2008). Metabolic profiling of the human response to a glucose challenge reveals distinct axes of insulin sensitivity. Molecular Systems Biology, 4, 214.
Simmons, W. W., Closs, E. I., Cunningham, J. M., Smith, T. W., & Kelly, R. A. (1996). Cytokines and insulin induce cationic amino acid transporter (CAT) expression in cardiac myocytes. Regulation of l-arginine transport and no production by CAT-1, CAT-2A, and CAT-2B. Journal of Biological Chemistry, 271, 11694–11702.
Smyth, G. K. (2005). Limma: Linear models for microarray data. In R. Gentleman, V. Carey, S. Dudoit, R. Irizarry, & W. Huber (Eds.), Bioinformatics and computational biology solutions using R and bioconductor (pp. 397–420). New York: Springer.
Spector, P. (2008). Data manipulation with R. New York: Springer.
Stump, C. S., Short, K. R., Bigelow, M. L., Schimke, J. M., & Nair, K. S. (2003). Effect of insulin on human skeletal muscle mitochondrial ATP production, protein synthesis, and mRNA transcripts. Proceedings of the National Academy of Sciences of the United States of America, 100, 7996–8001.
Suzuki, R., & Shimodaira, H. (2009). pvclust: Hierarchical clustering with P-values via multiscale bootstrap resampling. R package version 1.2-1, URL http://www.is.titech.ac.jp/~shimo/prog/pvclust/
Taylor, P. M. (2009). Amino acid transporters: Eminences grises of nutrient signalling mechanisms? Biochemical Society Transactions, 37, 237–241.
Tolhurst, G., Reimann, F., & Gribble, F. M. (2009). Nutritional regulation of glucagon-like peptide-1 secretion. Journal of Physiology, 587, 27–32.
Tremblay, F., & Marette, A. (2001). Amino acid and insulin signaling via the mTOR/p70 S6 kinase pathway. A negative feedback mechanism leading to insulin resistance in skeletal muscle cells. Journal of Biological Chemistry, 276, 38052–38060.
Um, S. H., D’Alessio, D., & Thomas, G. (2006). Nutrient overload, insulin resistance, and ribosomal protein S6 kinase 1, S6K1. Cell Metabolism, 3, 393–402.
van Loon, L. J., Saris, W. H., Verhagen, H., & Wagenmakers, A. J. (2000). Plasma insulin responses after ingestion of different amino acid or protein mixtures with carbohydrate. American Journal of Clinical Nutrition, 72, 96–105.
Vary, T. C., Jefferson, L. S., & Kimball, S. R. (1999). Amino acid-induced stimulation of translation initiation in rat skeletal muscle. American Journal of Physiology, 277, E1077–E1086.
Zhao, X., Peter, A., Fritsche, J., Elcnerova, M., Fritsche, A., Haring, H. U., et al. (2009). Changes of the plasma metabolome during an oral glucose tolerance test: Is there more than glucose to look at? American Journal of Physiology Endocrinology and Metabolism, 296, E384–E393.
Acknowledgments
We specifically would like to thank Kerstin Geillinger for the insulin measurements, Manuela Sailer for the measurements of phospholipids and acylcarnitines. We also acknowledge Johanna Welzhofer and Manuela Hubersberger for their excellent technical assistance. GLP-1 was kindly measured by Jens Holst, University of Copenhagen, Denmark. This work was supported in part by the Else Kröner-Fresenius-Foundation, Bad Hombrug v.d. Höhe, Germany.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Thomas Skurk and Isabel Rubio-Aliaga contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
Time–course profiling of the amino acids considering insulin extraction. In black, the fold-change time–course of the amino acids after a glucose and a protein-hydrolysate load (OGTT-PL). In grey, the calculated fold-change time–course by removing the clearance effect caused by insulin, i.e. the time–course due only to the protein-hydrolysate load added (PLcalculated). The calculation was performed for each variable and individual, as FCtime = i;treatment = PL = FCtime = i;treatment = OGTT + PL +( 1 – FCtime = i;treatment = OGTT). Data depicted as mean ± SEM (TIFF 773 kb)
Rights and permissions
About this article
Cite this article
Skurk, T., Rubio-Aliaga, I., Stamfort, A. et al. New metabolic interdependencies revealed by plasma metabolite profiling after two dietary challenges. Metabolomics 7, 388–399 (2011). https://doi.org/10.1007/s11306-010-0258-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-010-0258-z