Abstract
Beech is one of the most important trees in the temperate and subtropical forests of the Northern Hemisphere. Despite Chinese beeches have the particularity that only grow in subtropical areas, they have received few phylogeographic research. In this study, we sampled 25 populations of the northernmost-distributed Chinese beech, Fagus engleriana, and detected six haplotypes across 350 individuals by using sequences of two chloroplast intergenic spacers. The chloroplast genetic diversity was relatively low (h T = 0.659), with most genetic variance residing among populations (G ST = 0.831, N ST = 0.855, G ST≈N ST). SAMOVA analysis indicated that populations clustered into six groups with little admixture among them (most groups were characterized by a unique hapotype). Pairwise difference among haplotypes and Fu’s Fs statistic indicated that populations of F. engleriana have not experienced recent sudden expansions. Both the phylogeographic and demographic patterns found in this study suggest that F. engleriana remained fragmented in multiple refugia throughout the Pleistocene climatic changes, and experienced limited both glacial and interglacial/postglacial expansion. The results of this study imply that long-term isolation among multiple refugia, coupled with little admixture among populations of different refugia provided numerous opportunities for population divergence and allopatric speciation, which might be a driving factor for the exceptionally broad temperate species diversity in southern China.


Similar content being viewed by others
References
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Bennett KD, Provan J (2008) What do we mean by ‘refugia’? Quat Sci Rev 27:2449–2455
Cao KF, Peters R, Oldeman RAA (1995) Climatic range and distribution of Chinese Fagus species. J Veget Sci 6:317–324
Cao KF, Peters R (1998) Structure and stem growth of multi-stemmed trees of Fagus engleriana in China. Plant Ecol 139:211–220
Chen K, Abbott RJ, Milne RI, Tian X-M, Liu J (2008) Phylogeography of Pinus tabulaeformis Carr. (Pinaceae), a dominant species of coniferous forest in northern China. Mol Ecol 17:4276–4288
Comes HP, Kadereit JW (1998) The effect of Pleistocene climatic changes on plant distribution and evolution. Trends Plant Sci 3:432–438
Denk T (2003) Phylogeny of Fagus L. (Fagaceae) based on morphological data. Plant Syst Evol 240:55–81
Denk T, Grimm GW, Hemleben V (2005) Patterns of molecular and morphological differentiation in Fagus (Fagaceae): phylogenetic implications. Am J Bot 92:1006–1016
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Fang J, Lechowicz MJ (2006) Climatic limits for the present distribution of beech (Fagus L.) species in the world. J Biogeogr 33:1804–1819
Frascaria N, Maggia L, Michaud M, Bousquet J (1993) The rbcL gene sequence from chestnut indicates a slow rate of evolution in the Fagaceae. Genome 36:668–671
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Funk DJ, Omland KE (2003) Species-level paraphyly and polyphyly: frequency, causes, and consequences, with insights from animal mitochondrial DNA. Annu Rev Ecol Evol System 34:397–423
Gao LM, Möller M, Zhang XM, Hollingsworth ML, Liu J, Mill RR, Gibby M, Li DZ (2007) High variation and strong phylogeographic pattern among cpDNA haplotypes in Taxus wallichiana (Taxaceae) in China and North Vietnam. Mol Ecol 16:4684–4698
Gong W, Chen C, Dobeš C, Fu C-X, Koch MA (2008) Phylogeography of a living fossil: Pleistocene glaciations forced Ginkgo biloba L. (Ginkgoaceae) into two refuge areas in China with limited subsequent postglacial expansion. Mol Phylogenet Evol 48:1094–1105
Gugger P, González-Rodríguez A, Rodríguez-Correa H, Sugita S, Cavender-Bares J (2011) Southward Pleistocene migration of douglas-fir into Mexico: phylogeography, ecological niche modeling, and conservation of ‘rear edge’ populations. New Phytol 189:1185–1199
Guo K, Werger MJA (2010) Effect of prevailing monsoons on the distribution of beeches in continental East Asia. Forest Ecol Manag 259:2197–2203
Hampe A, Petit RJ (2005) Conserving biodiversity under climate change: the rear edge matters. Ecol Lett 8:461–467
Harpending HC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 66:591–600
Harrison SP, Yu G, Takahara H, Prentice IC (2001) Diversity of temperate plants in east Asia. Nature 413:129–130
Hewitt GM (2004) Genetic consequences of climatic oscillations in the Pleistocene. Philos Trans R Soc London (Biol) 359:183–195
Hiraoka K, Tomaru N (2009) Population genetic structure of Fagus japonica revealed by nuclear microsatellite markers. Int J Plant Sci 170:748–758
Hu FS, Hampe A, Petit RJ (2009) Paleoecology meets genetics: deciphering past vegetational dynamics. Front Ecol Environ 7:371–379
Jakob SS, Blattner FR (2006) A chloroplast genealogy of Hordeum (Poaceae): long-term persisting haplotypes, incomplete lineage sorting, regional extinction, and the consequences for phylogenetic inference. Mol Biol Evol 23:1602–1612
Johnson WC, Adkisson CS (1985) Dispersal of beech nuts by blue jays in fragmented landscapes. Am Midl Nat 113:319–324
Ju L, Wang H, Jiang D (2007) Simulation of the Last Glacial Maximum climate over East Asia with a regional climate model nested in a general circulation model. Palaeogeogr Palaeoclim Palaeoecol 248:376–390
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism. Bioinformatics 25:1451–1452
Liu Y-S, Wang W-M, Arata M (1998) China’s beech forests in the Pre-Pleistocene. Foss Rec 1:151–165
Liu H, Xing Q, Ji Z, Xu L, Tian Y (2003) An outline of Pleistocene development of Fagus forest in China: palynological and ecological perspectives. Flora 198:249–259
Liu M-H (2008) Phylogeography of Fagus longipetiolata: insights from nuclear DNA microsatellites and chloroplast DNA variation. Dissertation, East China Normal University, Shanghai
López-Pujol J, Zhang F-M, Sun H-Q, Ying T-S, Ge S (2011) Centres of plant endemism in China: places for survival or for speciation? J Biogeogr 38:1267–1280
Magri D, Vendramin GG, Comps B et al (2006) A new scenario for the quaternary history of European beech populations: palaeobotanical evidence and genetic consequences. New Phytol 171:199–221
Morris AB, Graham CH, Soltis DE, Soltis PS (2010) Reassessment of phylogeographical structure in an eastern North American tree using Monmonier's algorithm and ecological niche modelling. J Biogeogr 37:1657–1667
Ni J, Yu G, Harrison SP, Prentice IC (2010) Palaeovegetation in China during the late Pleistocene: biome reconstructions based on a global scheme of plant functional types. Palaeogeogr Palaeoclim Palaeoecol 289:44–61
Okaura T, Harada K (2002) Phylogeographical structure revealed by chloroplast DNA variation in Japanese Beech (Fagus crenata Blume). Heredity 88:322–329
Ohkawa T, Kitamura K, Takasu H, Kawano S (2006) Genetic variation in Fagus multinervis Nakai (Fagaceae), a beech species endemic to Ullung Island, South Korea. Plant Species Biol 21:135–145
Palme AE, Su Q, Palsson S, Lascoux M (2004) Extensive sharing of chloroplast haplotypes among European birches indicates hybridization among Betula pendula, B. pubescens and B. nana. Mol Ecol 13:167–178
Petit RJ, Hampe A, Cheddadi R (2005) Climate changes and tree phylogeography in the Mediterranean. Taxon 54:877–885
Pilkington MM, Wilder JA, Mendez FL et al (2008) Contrasting signatures of population growth for mitochondrial DNA and Y chromosomes among human populations in Africa. Mol Biol Evol 25:517–525
Pons O, Petit RJ (1996) Measuring and testing genetic differentiation with ordered versus unordered alleles. Genetics 144:1237–1245
Qian H, Ricklefs RE (2000) Large-scale processes and the Asian bias in species diversity of temperate plants. Nature 407:180–182
Qian H, Ricklefs RE (2001) Diversity of temperate plants in East Asia (Qian and Ricklefs reply). Nature 413:130
Qiu Y-X, Guan B-C, Fu C-X, Comes HP (2009a) Did glacials and/or interglacials promote allopatric incipient speciation in East Asian temperate plants? Phylogeographic and coalescent analyses on refugial isolation and divergence in Dysosma versipellis. Mol Phylogenet Evol 51:281–293
Qiu Y-X, Sun Y, Zhang X-P, Lee J, Fu C-X, Comes HP (2009b) Molecular phylogeography of East Asian Kirengeshoma (Hydrangeaceae) in relation to Pleistocene climate change and landbridge configurations. New Phytol 183:480–495
Qiu Y-X, Fu C-X, Comes HP (2011) Plant molecular phylogeography in China and adjacent regions: tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol Phylogenet Evol 59:225–244
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288
Shen CF (1992) A monograph of the genus Fagus Tourn. ex L. (Fagaceae). The City University of New York, New York
Soltis DE, Morris AB, McLachlan JS, Manos PS, Soltis PS (2006) Comparative phylogeography of unglaciated eastern North America. Mol Ecol 15:4261–4293
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Tian S, López-Pujol J, Wang H-W, Ge S, Zhang Z-Y (2010) Molecular evidence for glacial expansion and interglacial retreat during Pleistocene climatic changes in a montane temperate pine (Pinus kwangtungensis Chun ex Tsiang) in southern China. Plant Syst Evol 284:219–229
Wang H-W, Ge S (2006) Phylogeography of the endangered Cathaya argyrophylla (Pinaceae) inferred from sequence variation of mitochondrial and nuclear DNA. Mol Ecol 15:4109–4122
Wang J, Gao P, Kang M, Lowe AJ, Huang H (2009) Refugia within refugia: the case study of a canopy tree (Eurycorymbus cavaleriei) in subtropical China. J Biogeogr 36:2156–2164
Wu ZY, Wu SG (1996) A proposal for a new floristic kingdom (realm)—the E. Asiatic kingdom, its delimitation and characteristics. In: Zhang AL, Wu SG (eds) Proceedings of the first international symposium on floristic characteristics and diversity of east asian plants. China Higher Education Press, Beijing, pp 3–42
Yu G, Chen X, Ni J, Cheddadi R et al (2000) Palaeovegetation of China: a pollen data-based synthesis for the mid-Holocene and Last Glacial Maximum. J Biogeogr 27:635–664
Yu G, Xue B, Liu J, Chen X (2003) LGM lake records from China and an analysis of climate dynamics using a modelling approach. Glob Planet Chang 38:223–256
Zhao Y, Yu Z, Chen F, Zhang J, Yang B (2009) Vegetation response to Holocene climate change in monsoon-influenced region of China. Earth-Sci Rev 97:242–256
Yuan QJ, Zhang ZY, Peng H, Ge S (2008) Chloroplast phylogeography of Dipentodon (Dipentodontaceae) in southwest China and northern Vietnam. Mol Ecol 17:1054–1065
Zheng Z (2000) Vegetation and climate since the late Pleistocene in southern China. J Geosci China 2:7–20
Zheng YQ, Yu G, Wang SM, Xue B, Zhuo DQ, Zeng XM, Liu HQ (2004) Simulation of paleoclimate over East Asia at 6 Ka BP and 21 Ka BP by a regional climate model. Clim Dynam 23:513–529
Zhou T-H, Li S, Qian Z-Q, Su H-L, Huang Z-H, Guo Z-G, Dai P-F, Liu Z-L, Zhao G-F (2010) Strong phylogeographic pattern of cpDNA variation reveals multiple glacial refugia for Saruma henryi Oliv. (Aristolochiaceae), an endangered herb endemic to China. Mol Phylogenet Evol 57:176–188
Acknowledgements
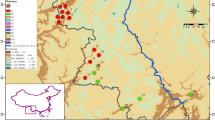
The authors thank Dr. Deng-Mei Fan for her help in drawing the haplotype distribution map. This study was funded by a project of National Science Foundation of China (30760016) and the Cultivation Program for Young Scientists of Jiangxi Province (grants to Zhi-Yong Zhang, 2008DQ01500).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by A. Kremer
Ming Lei and Qun Wang are co-first authors.
Rights and permissions
About this article
Cite this article
Lei, M., Wang, Q., Wu, ZJ. et al. Molecular phylogeography of Fagus engleriana (Fagaceae) in subtropical China: limited admixture among multiple refugia. Tree Genetics & Genomes 8, 1203–1212 (2012). https://doi.org/10.1007/s11295-012-0507-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-012-0507-6