Abstract
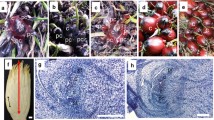
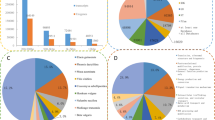
Micropropagation of oil palm by somatic embryogenesis produces a proportion of off-type individuals (approximately 5% overall) displaying a homeotically modified flower structure known as mantled. Transformation of the fertile or sterile androecium into carpel-like structures is observed in staminate and pistillate mantled flowers, respectively, resulting in lower oil yields in affected plantations. Given the epigenetic nature of the mantled condition, a gene expression-based approach was used rather than a genetic one to investigate its molecular basis. Suppression subtractive hybridisation (SSH) and macroarray hybridisation were used to compare transcriptome patterns between normal and mantled inflorescences. Two SSH libraries, enriched for complementary deoxyribonucleic acids (cDNAs) of either true-to-type or somaclonal variant material, were generated. Bioinformatic analysis of these two libraries allowed the identification of 1,350 unique sequences and their annotation by a gene ontology-based approach. Macroarray hybridisation was used to compare gene expression between normal and mantled inflorescences, and 32 genes were found to be differentially expressed. The temporal expression patterns of six genes were further investigated in more detail in relation to male and female inflorescence development. Full-length cDNAs were isolated and characterised for two of these genes, EgFB1 and EgRING1, both of which are down-regulated in the mantled inflorescences and both of which encode proteins associated with proteolytic signalling complexes. Our data shed light on gene expression changes associated with the mantled phenotype and have provided novel transcriptome markers which can help to distinguish the abnormal and wild-type inflorescences.





Similar content being viewed by others
References
Adam H, Jouannic S, Escoute J, Duval Y, Verdeil JL, Tregear JW (2005) Reproductive developmental complexity in the African oil palm (Elaeis guineensis). Am J Bot 92:1836–1852
Adam H, Jouannic S, Morcillo F, Richaud F, Duval Y, Tregear JW (2006) MADS box genes in oil palm (Elaeis guineensis): patterns in the evolution of the squamosa, deficiens, globosa, agamous, and sepallata subfamilies. J Mol Evol 62:15–31
Adam H, Jouannic S, Orieux Y, Morcillo F, Richaud F, Duval Y, Tregear JW (2007) Functional characterization of MADS box genes involved in the determination of oil palm flower structure. J Exp Bot 58:1245–1259
Aharoni A, Keizer LC, Bouwmeester HJ, Sun Z, Alvarez-Huerta M et al (2000) Identification of the SAAT gene involved in strawberry flavor biogenesis by use of DNA microarrays. Plant Cell 12:647–662
Altschul SF et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alwee SS, Van der Linden CG, Van der Schoot J, de Folter S, Angenent GC, Cheah SC, Smulders MJM (2006) Characterization of oil palm MADS box genes in relation to the mantled flower abnormality. Plant Cell Tissue Organ Cult 85:331–344
Cao Y, Yang Y, Zhang H, Li D, Zheng Z, Song F (2008) Overexpression of a rice defense-related F-box protein gene OsDRF1 in tobacco improves disease resistance through potentiation of defense gene expression. Physiol Plant 134:440–452
Chae E, Tan QK, Hill TA, Irish VF (2008) An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development. Development 135:1235–1245
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Conesa A, Gotz S (2008) Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 2008:619832
Corley RHV, Lee CH, Law IM, Wong CY (1986) Abnormal flower development in oil palm clones. Planter 62:233–240
Durand-Gasselin T, Le Guen VL, Konan E, Duval Y (1990) Oil palm (Elaeis guineensis Jacq.) plantations in Côte d’Ivoire obtained through in vitro culture-first results. Oléagineux 45:1–11
Gagne JM, Downes BP, Shiu SH, Durski AM, Vierstra RD (2002) The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc Natl Acad Sci USA 99:11519–11524
Hardtke CS, Okamoto H, Stoop-Myer C, Deng XW (2002) Biochemical evidence for ubiquitin ligase activity of the Arabidopsis COP1 interacting protein 8 (CIP8). Plant J 30:385–394
Ho MS, Tsai PI, Chien CT (2006) F-box proteins: the key to protein degradation. J Biomed Sci 13:181–191
Ho CL, Kwan YY, Choi MC, Tee SS, Ng WH et al (2007) Analysis and functional annotation of expressed sequence tags (ESTs) from multiple tissues of oil palm (Elaeis guineensis Jacq.). BMC Genomics 8:381
Hu W, Wang Y, Bowers C, Ma H (2003) Isolation, sequence analysis, and expression studies of florally expressed cDNAs in Arabidopsis. Plant Mol Biol 53:545–563
Ingram GC, Doyle S, Carpenter R, Schultz EA, Simon R, Coen ES (1997) Dual role for fimbriata in regulating floral homeotic genes and cell division in Antirrhinum. EMBO J 16:6521–6534
Jain M, Nijhawan A, Arora R, Agarwal P, Ray S et al (2007) F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 143:1467–1483
Jaligot E, Rival A, Beulé T, Dussert S, Verdeil JL (2000) Somaclonal variation in oil palm (Elaeis guineensis Jacq.): the DNA methylation hypothesis. Plant Cell Rep 19:684–690
Jaligot E, Beulé T, Rival A (2002) Methylation-sensitive RFLPs: characterisation of two oil palm markers showing somaclonal variation-associated polymorphism. Theor Appl Genet 104:1263–1269
Jaligot E, Beulé T, Baurens FC, Billotte N, Rival A (2004) Search for methylation-sensitive amplification polymorphisms associated with the mantled variant phenotype in oil palm (Elaeis guineensis Jacq). Genome 47:224–228
Joazeiro CA, Weissman AM (2000) Ring finger proteins: mediators of ubiquitin ligase activity. Cell 102:549–552
Jouannic S, Argout X, Lechauve F, Fizames C, Borgel A et al (2005) Analysis of expressed sequence tags from oil palm (Elaeis guineensis). FEBS Lett 579:2709–2714
Jurado S, Diaz-Trivino S, Abraham Z, Manzano C, Gutierrez C, del Pozo C (2008) SKP2A, an F-box protein that regulates cell division, is degraded via the ubiquitin pathway. Plant J 53:828–841
Kim WY, Fujiwara S, Suh SS, Kim J, Kim Y et al (2007) Zeitlupe is a circadian photoreceptor stabilized by gigantea in blue light. Nature 449:356–360
Kim C, Lemke C, Paterson AH (2009) Functional dissection of drought-responsive gene expression patterns in Cynodon dactylon L. Plant Mol Biol 70:1–16
Kipreos ET, Pagano M (2000) The F-box protein family. Genome Biol 1: REVIEWS3002
Koiwai H, Tagiri A, Katoh S, Katoh E, Ichikawa H et al (2007) RING-H2 type ubiquitin ligase EL5 is involved in root development through the maintenance of cell viability in rice. Plant J 51:92–104
Kraft E, Bostick M, Jacobsen SE, Callis J (2008) ORTH/VIM proteins that regulate DNA methylation are functional ubiquitin E3 ligases. Plant J 56:704–715
Kubis SE, Castilho AM, Vershinin AV, Heslop-Harrison JS (2003) Retroelements, transposons and methylation status in the genome of oil palm (Elaeis guineensis) and the relationship to somaclonal variation. Plant Mol Biol 52:69–79
Laitinen RA, Pollanen E, Teeri TH, Elomaa P, Kotilainen M (2007) Transcriptional analysis of petal organogenesis in Gerbera hybrida. Planta 226:347–360
Levin JZ, Meyerowitz EM (1995) UFO: an Arabidopsis gene involved in both floral meristem and floral organ development. Plant Cell 7:529–548
Lin HC, Morcillo F, Dussert S, Tranchant-Dubreuil C, Tregear JW, Tranbarger TJ (2009) Transcriptome analysis during somatic embryogenesis of the tropical monocot Elaeis guineensis: evidence for conserved gene functions in early development. Plant Mol Biol 70:173–192
Low ET, Alias H, Boon SH, Shariff EM, Tan CY et al (2008) Oil palm (Elaeis guineensis Jacq.) tissue culture ESTs: identifying genes associated with callogenesis and embryogenesis. BMC Plant Biol 8:62
Matthes M, Singh R, Cheah SC, Karp A (2001) Variation in oil palm (Elaeis guineensis Jacq.) tissue culture-derived regenerants revealed by AFLPs with methylation-sensitive enzymes. Theor Appl Genet 102:971–979
Mazzucotelli E, Belloni S, Marone D, De Leonardis A, Guerra D et al (2006) The e3 ubiquitin ligase gene family in plants: regulation by degradation. Curr Genomics 7:509–522
Morcillo F, Gagneur C, Adam H, Richaud F, Singh R et al (2006) Somaclonal variation in micropropagated oil palm. Characterization of two novel genes with enhanced expression in epigenetically abnormal cell lines and in response to auxin. Tree Physiol 26:585–594
Ni W, Xie D, Hobbie L, Feng B, Zhao D et al (2004) Regulation of flower development in Arabidopsis by SCF complexes. Plant Physiol 134:1574–1585
Nogueira FT, De Rosa VE, Jr MM, Ulian EC, Arruda P (2003) RNA expression profiles and data mining of sugarcane response to low temperature. Plant Physiol 132:1811–1824
Osterlund MT, Hardtke CS, Wei N, Deng XW (2000) Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405:462–466
Pannetier C, Arthuis P, Lievoux D (1981) Neoformation of young Elaeis guineensis plants from primary calluses obtained on leaf fragments in vitro. Oleagineux 36:119–122
Pertea G, Huang X, Liang F, Antonescu V, Sultana R, Karamycheva S, Lee Y, White J, Cheung F, Parvizi B et al (2003) TIGR gene indices clustering tools (TGICL): a software system for fast clustering of large EST datasets. Bioinformatics 19:651–652
Peschke VM, Phillips RL, Bengenbach BG (1987) Discovery of transposable element activity among progeny of tissue culture-derived maize plants. Science 228:804–807
Poncet V, Rondeau M, Tranchant C, Cayrel A, Hamon S et al (2006) SSR mining in coffee tree EST databases: potential use of EST-SSRs as markers for the Coffea genus. Mol Genet Genomics 276:436–449
Rao V, Donough CR (1990) Preliminary evidence of genetic cause for the floral abnormalities in some oil palm ramets. Elaeis 2:199–207
Rival A (2007) The oil palm. In: Pua EC, Davey MR (eds) Biotechnology in agriculture and forestry, vol 61. Transgenic crops VI. Springer-Verlag, Berlin, pp 59–80
Rival A, Beulé T, Barre P, Hamon S, Duval Y, Noirot M (1997) Comparative flow cytometric estimation of nuclear DNA content in oil palm (Elaeis guineensis Jacq) tissue cultures and seed-derived plants. Plant Cell Rep 16:884–887
Rival A, Bertrand L, Beulé T, Combes MC, Trouslot P, Lashermes P (1998) Suitability of RAPD analysis for the detection of somaclonal variants in oil palm (Elaeis guineensis Jacq). Plant Breed 117:73–76
Rival A, Jaligot E, Beulé T, Finnegan EJ (2008) Isolation and expression analysis of genes encoding MET, CMT, and DRM methyltransferases in oil palm (Elaeis guineensis Jacq.) in relation to the ‘mantled’ somaclonal variation. J Exp Bot 59:3271–3281
Samach A, Klenz JE, Kohalmi SE, Risseeuw E, Haughn GW, Crosby WL (1999) The unusual floral organs gene of Arabidopsis thaliana is an F-box protein required for normal patterning and growth in the floral meristem. Plant J 20:433–445
Sawa M, Nusinow DA, Kay SA, Imaizumi T (2007) FKF1 and gigantea complex formation is required for day-length measurement in Arabidopsis. Science 318:261–265
Sonoda Y, Yao SG, Sako K, Sato T, Kato W et al (2007) SHA1, a novel RING finger protein, functions in shoot apical meristem maintenance in Arabidopsis. Plant J 50:586–596
Stirnberg P, Furner IJ, Ottoline Leyser HM (2007) MAX2 participates in an SCF complex which acts locally at the node to suppress shoot branching. Plant J 50:80–94
Stone SL, Hauksdottir H, Troy A, Herschleb J, Kraft E, Callis J (2005) Functional analysis of the ring-type ubiquitin ligase family of Arabidopsis. Plant Physiol 137:13–30
Tamaoki M, Matsuyama T, Nakajima N, Aono M, Kubo A, Saji H (2004) A method for diagnosis of plant environmental stresses by gene expression profiling using a cDNA macroarray. Environ Pollut 131:137–145
Tregear JW, Morcillo F, Richaud F, Berger A, Singh R et al (2002) Characterization of a defensin gene expressed in oil palm inflorescences: induction during tissue culture and possible association with epigenetic somaclonal variation events. J Exp Bot 53:1387–1396
Vierstra RD (2009) The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 10:385–397
Wang XJ, Cao XL, Hong Y (2005) Isolation and characterization of flower-specific transcripts in Acacia mangium. Tree Physiol 25:167–178
Wilkinson MD, Haughn GW (1995) Unusual floral organs controls meristem identity and organ primordia fate in Arabidopsis. Plant Cell 7:1485–1499
Yu H, Wu J, Xu N, Peng M (2007) Roles of F-box proteins in plant hormone responses. Acta Biochim Biophys Sin (Shanghai) 39:915–922
Zhang X, Garreton V, Chua NH (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev 19:1532–1543
Zhang Y, Yang C, Li Y, Zheng N, Chen H et al (2007) SDIR1 is a RING finger E3 ligase that positively regulates stress-responsive abscisic acid signaling in Arabidopsis. Plant Cell 19:1912–1929
Zhang JZ, Li ZM, Yao JL, Hu CG (2009) Identification of flowering-related genes between early flowering trifoliate orange mutant and wild-type trifoliate orange (Poncirus trifoliata L. Raf.) by suppression subtraction hybridization (SSH) and macroarray. Gene 430:95–104
Acknowledgements
The authors are grateful to colleagues at Felda Agricultural Services (FASSB) for their valuable help, to Dr Helene Adam for the supply of inflorescence RNA and expert advice on inflorescence development, to Ivanna Fuentes for skilled technical support and to Myriam Collin for excellent assistance in histology and microscopy studies. We thank Drs. Alain Rival and Fabienne Morcillo for the critical reading of the manuscript. This research was supported by institutional funding from IRD and CIRAD.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Dean
Rights and permissions
About this article
Cite this article
Beulé, T., Camps, C., Debiesse, S. et al. Transcriptome analysis reveals differentially expressed genes associated with the mantled homeotic flowering abnormality in oil palm (Elaeis guineensis). Tree Genetics & Genomes 7, 169–182 (2011). https://doi.org/10.1007/s11295-010-0323-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-010-0323-9