Abstract
The spread of polymyxin-resistant Klebsiella pneumoniae strains represents an emerging health challenge, limiting treatment options for the patients. Thus, the development of new antimicrobials is an urgent requirement. Antimicrobial peptides (AMPs) are a large class of compounds that are part of innate immune response; these peptides are promising compounds in the field of antimicrobial resistance and are present in all organisms. The present review evaluated patents on antimicrobial peptides tested against polymyxin-resistant K. pneumoniae, available on Espacenet as of September 2022. A total of 1313 patents were examined and 1197 excluded as they were out of focus for this review; 104 patents of peptides tested against K. pneumoniae were included; of which only 14 were tested against polymyxin-resistant K. pneumoniae strains. The results indicated that all AMPs evaluated were in the experimental or pre-clinical phase; the clinical phase is pending. Furthermore, a few peptides were tested effectively against polymyxin-resistant K. pneumoniae. Although, the research and patent filing alone are not enough to develop a suitable antimicrobial therapy, they can represent good starting point upon which to develop new antimicrobials. More investment is required to push these pharmaceuticals through the stages of development to introduce them into the market.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
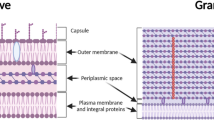
Klebsiella pneumoniae is a Gram-negative opportunistic pathogen that causes multidrug-resistant (MDR) infections (including pneumonia, urinary tract infections, and bloodstream infections) (Martin and Bachman 2018; Wang et al. 2020). In the last decade, it has emerged as a global health threat (Wyres et al. 2020; De Oliveira et al. 2020). This microorganism has an exceptional ability to develop and acquire genetic elements that encode resistance to multiple antibiotics, including carbapenems (Navon-Venezia et al. 2017; Yang et al. 2021).
Carbapenems are often used to treat MDR infections (Hansen 2021). However, carbapenem-resistant infections are difficult to treat because of the lack of effective and safe options (Soman et al. 2021). In these circumstances, polymyxins are antimicrobials of last resort for the treatment of infections caused by carbapenem-resistant K. pneumoniae (Rojas et al. 2016). Polymyxins were widely used in hospitals to treat infections, resulting in the current scenario of polymyxin-resistant strains (Zhang et al. 2021b). Resistance to polymyxin B, an antibiotic of last resort in the treatment of serious bacterial infections, leads to the failure of antibiotic treatment (Li et al. 2022).
Infections caused by polymyxin-resistant K. pneumoniae are associated with high mortality rates of around 60% (Da Silva et al. 2020). The main resistance mechanisms in polymyxin-resistant K. pneumoniae are mutations in mgrB, phoPQ, pmrAB, and crrAB and the presence of the mcr gene plasmid (De La Cadena et al. 2021). Therefore, limited therapeutic options are available to treat infections caused by polymyxin-resistant K. pneumoniae (Levin and Oliveira 2008). In this context, the introduction of conventional antimicrobial therapies often results in increased resistance (Nainu et al. 2021). In recent years, few new antibiotics have been approved, thereby compounding the scenario of antibiotic resistance (Shi et al. 2021). Therefore, initiatives for the development of new therapeutic alternatives are urgently required to control MDR K. pneumoniae (Felício et al. 2021; You et al. 2021).
Antimicrobial peptides (AMPs) are promising candidates for controlling infections caused by MDR microorganisms (Bin Hafeez et al. 2021). They are composed of 10–50 amino acids, are present in different organisms, and act against several pathogens, such as bacteria, viruses, fungi, and parasites (Rodríguez et al. 2021; Deshayes et al. 2022). AMPs have attracted interest as novel therapeutic agents because they exhibit potent and broad-spectrum antibiotic activities with a different mechanism of action from traditional antibiotics. Cationic AMPs interact and penetrate bacterial cell membranes, leading to bacterial death (Shi et al. 2021). In addition, AMPs have several mechanisms of action, which target intracellular action, acting on nucleic acids, cell wall synthesis, protein synthesis, and enzymatic activity, among others (Zhang et al. 2021a). In addition, they exhibit a broad spectrum of antibacterial, antifungal, and antiviral activities. Chemical structure optimization to create more effective synthetic peptides represents a promising strategy for the development of AMPs as a new drug class to prevent and treat systemic and topic infections (Cardoso et al. 2020). Additionally, they exhibit a reduced propensity to induce drug resistance when compared to conventional bacterial antibiotics (Wang et al. 2016). However, despite representing a promising therapy, it presents challenges for applications, which include potential toxicity in humans, poor specificity and costly manufacturing (Bahar and Ren 2013).
Several studies have described different antimicrobial peptides with distinct spectra of activity and mechanisms of action. But how many of these peptides have been patented for commercial use? How many peptides were tested against polymyxin-resistant K. pneumoniae? Patent content analysis is a vital tool to understand trends in the development of new drugs and explore the most promising target that can be produced commercially (Han 2007; Serafini et al. 2021). Therefore, the present study performed a patent review analysis to identify and explore innovation trends, therapeutic strategies, and the latest antimicrobial peptides for the treatment of polymyxin-resistant K. pneumoniae infections.
Search strategy
The flowchart of the present review was reported in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines (Page et al. 2021). The search for patents in the specialized online database: Espacenet from the European Patent Office (EPO), as described (Serafini et al. 2021) was performed with modifications. The present review evaluated relevant patents deposited until September 2022. This review did not include a meta-analysis, risk of bias assessment, assessment of causes of heterogeneity, or assessments of certainty (or confidence) or robustness. First, keywords were selected using the MeSH (Medical Subject Headings) for indexing articles for PubMed. The title, abstract, and full text of the articles were searched using a combination of "peptide", "Klebsiella pneumoniae," and "resistance polymyxin." The PubMed literature database was searched using the same keywords to compare the number of articles with the number of patents identified from the patent databases.
Study selection
Two independent reviewers (GHAS and LR) screened the search results obtained from patent databases and PubMed for inclusion. Full text of the qualified patents and articles were screened by the same two independent reviewers. A third reviewer (SS) was called in to resolve any differences of opinion in order to form a consensus. Inclusion criteria were as follows: (1) studies with antimicrobial peptides tested against K. pneumoniae. The exclusion criteria were as follows: (1) Patents containing peptides that were not tested against Klebsiella pneumoniae; (2) duplicate and unavailable patents.
The search results (via Espacenet), containing the country, year, applicant type, mechanism of action, and International Patent Classification (IPC) code, were exported to a Microsoft Excel (Microsoft Corporation, Redmond, WA, USA).
Results
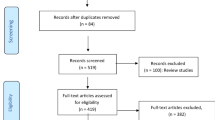
A total of 1313 patents were identified for the preliminary evaluation of the database (Fig. 1). Of these, 1197 patents were excluded because they did not fit the scope or did not contain descriptive data. Moreover, we identified 116 patents for peptides tested against K. pneumoniae, of which 12 patents were excluded due to duplication. Out of those remaining 104, 14 eligible patents were included in this review. We emphasize that using the same keywords in the "Advanced search" option returned only 3 patents, of which two were in Chinese (no translation) and the other patent contained peptides that were not tested against polymyxin-resistant K. pneumoniae. Therefore, this search option is limiting as it could not search for patents within the scope. Because of this reason, the general search method was used.
The article search identified 1055 publications from 1990 to 2022. Of these, books, documents, meta-analyses, reviews, and systematic reviews (n = 85) were excluded, leaving 970 publications. Finally, to identify trials in the clinical phase that were being conducted against MDR K. pneumoniae, a search was performed through the Clinical Trials (https://clinicaltrials.gov/) registry database (search terms: peptide | Infection, Bacterial); the search yielded no such trials.
In the last hundred years, more than 80 peptide drugs (for osteoporosis, multiple sclerosis, diabetes, cancer, human immunodeficiency virus infection, and chronic pain) have been introduced into the pharmacological market (Muttenthaler et al. 2021). Thus, patent analysis becomes essential to determine the generated innovations, predict new technological developments, and evaluate the progress of these innovations (Serafini et al. 2021). We identified an information gap regarding the development of peptides against MDR K. pneumoniae. Therefore, by analyzing patent filings, we can expose the main trends in the development of AMPs, including countries, filing years, and the action spectrum of peptides against several microorganisms (sensitive, MDR, and polymyxin-resistant).
Trends in the discovery of AMPs against polymyxin-resistant K. pneumoniae
Results indicated that patents were filed for AMPs against K. pneumoniae from 1993. In the last 30 years (1993–2022), 90 patents have been filed. The distribution of the number of patent deposits revealed both upward and downward trends over the years (Fig. 2A). However, since 2019, patent deposits have remained high, reaching a peak in 2021 with 14 patent deposits, the highest number ever registered since 1993. However, of these deposits, only 14 were peptide patents (2010–2022) tested against polymyxin-resistant K. pneumoniae (Fig. 2B). Between 2016 (n = 1) and 2017 (n = 7), the number of patent deposits increased. This may be due to the increase in the reporting of polymyxin-resistant isolates and the description of the first plasmid-mediated resistance mechanism against polymyxins (mcr-1) in China (Liu et al. 2016). Thus, as bacteria continue to develop resistance, initiatives are being established to develop new antibacterial agents (Stephens et al. 2020).
Frequency per year. A Patents of antimicrobial peptides against K. pneumoniae; B Patents of polymyxin-resistant K. pneumoniae antimicrobial peptides among the years 2010–2022; C Scientific articles published in the PubMed, using the terms “peptide”, “Klebsiella pneumoniae”, “resistance polymyxin”, database by year from 1990 to o October 10, 2022 (books, meta-analysis, review and systematic review, were not considered)
External funding for performing high-tech research leading to publications and patents is an increasingly prominent desire in the scientific community (Krauss and Kuttenkeuler 2021). However, the total number of patent publications was lower compared with the number of scientific articles published in the same period (Fig. 2C). Furthermore, since 2019, this number has decreased, possibly due to the global closing as a result of the COVID-19 pandemic.
Universities and companies leading the discovery of AMPs against polymyxin-resistant K. pneumoniae
The requirement to develop new antimicrobial drugs that are effective against MDR pathogens has spurred the research community to invest in various drug discovery strategies. (Farha and Brown 2019). Universities and industrial companies around the world claim the majority of patents on AMPs (Fig. 3) against polymyxin-resistant K. pneumoniae. Notably, the number of scientific publications does not follow the number of patent applications (Fig. 2A–C). This may be because the results of the articles do not qualify for the patentability criteria (such as industrial application, novelty, and inventive step) or the patent applications take a long time to be processed.
China and the United States lead the patent ranking in the discovery of AMPs against polymyxin-resistant K. pneumoniae
AMPs are promising compounds to address antimicrobial resistance, thereby representing a starting point for the development of new antimicrobials (Annunziato and Costantino 2020). For this reason, some countries have focused their research on this area. The United States (US) represents the country with the highest number of inventions, followed by China (Fig. 4A). This data corroborates the Worldwide Intellectual Property Office (WIPO) https://www.wipo.int/ipstats/en/statistics/country_profile indicators, where China and the US lead the ranking of patents in 2020, with 1,441,085 and 495,883 patents, respectively. This result is likely due to the US’s stable economy and investment in technology and intellectual property (Serafini et al. 2021).
Number of patent publications by country. A Antimicrobial peptides against K. pneumoniae; B Polymyxin-resistant K. pneumoniae antimicrobial peptides. SG Singapore, NZ New Zealand, RU Russian Federation, ES Spain, CA Canada, KR Korea, AU Australia, EP European Patent Office (EPO), WO World Intellectual Property Organization (WIPO), CN China, US United States of America
Classification of AMP patents against polymyxin-resistant K. pneumoniae
The IPC code is a universal application classification system that is administered by WIPO; it classifies patents according to technical fields to establish an orderly tool for searching patent documents (World Intellectual Property 2022). Most AMP patents against K. pneumoniae were classified in category A (human needs [n = 80]; highlighting: A61K [53], A01N [16], A61P [8]) or in category C (chemistry [n = 10]; especially C07K [8]; Fig. 5A). Among AMP patents against polymyxin-resistant K. pneumoniae, 92.8% were categorized as A61K (Fig. 5B).
Patents per International Patent Classification (IPC). A Antimicrobial peptides against K. pneumoniae; B Polymyxin-resistant K. pneumoniae antimicrobial peptides. C12N: microorganisms or enzymes; compositions there of; propagating, preserving, or maintaining microorganisms; mutation or genetic engineering; culture media; C07K: peptides; A01H: new plants or processes for obtaining them; plant reproduction by tissue culture techniques; A01P: biocidal, pest repellent, pest attractant, or plant growth regulatory activity of chemical compounds or preparations; A01N: preservation of bodies of humans or animals or plants or parts thereof; A23: foods or foodstuffs; treatment thereof, not covered by other classes; A61L: methods or apparatus for sterilizing materials or objects in general; disinfection, sterilization, or deodorization of air; chemical aspects of bandages, dressings, absorbent pads, or surgical articles; materials for bandages, dressings, absorbent pads, or surgical articles; A61K: preparations for medical, dental, or toilet purposes; A61P: specific therapeutic activity of chemical compounds or medicinal preparations
Effective peptides against polymyxin-resistant K. pneumoniae
After reviewing data on AMPs against K. pneumoniae (n = 90), patents tested against polymyxin-resistant strains (n = 14) were selected for a detailed review. We hereby summarize key trends in peptide drug discovery and development by including data on sources, structures, modes of action, and the resistance profile of the tested microorganisms (Table 1).
Discussion
In this manuscript, we performed a critical patent review on advances in the development of antimicrobial peptides against polymyxin-resistant K. pneumoniae. The patent application with the registration of "WO2010130007A1" (Li et al. 2010) refers to synthetic peptide antibiotics and polymyxin analog compounds that are effective against Gram-negative bacteria, including polymyxin-sensitive and MDR-resistant Gram-negative bacteria (colistin MICs > 128 mg/L). Specifically, for polymyxin-resistant K. pneumoniae the in vitro results demonstrated that the compound caused bacterial cell death. No toxicity and no adverse effects were observed in vivo using rat and mice models. The in vivo results using the mouse lung infection model corroborate the in vitro results, confirming the antibacterial activity. The compounds were subjected to antibacterial activity measurements against MDR strains, including polymyxin-resistant Pseudomonas aeruginosa, Acinetobacter baumannii, and K. pneumoniae. The minimum bactericidal concentration for compound 1 was 10.3 mg/L. Moreover, compounds 2–5 caused bacterial cell death. In contrast, colistin was inactive (even at 128 mg/L) against the resistant strain. A neutropenic mouse lung infection model demonstrated the superior in vivo efficacy of compound 3 compared with that of colistin (p < 0.045).
“CN108467424A” (Kui et al. 2018) refers to the field of biotechnology, in particular, to the linear antibacterial oligopeptide, designated SLAP-S25. Short linear antibacterial peptides (SLAP) have the advantages of simple structure, low production cost, high safety, and superior application prospects. Assays evaluating the antimicrobial effect against Gram-negative bacteria demonstrated that the SLAP-S25 oligopeptide has a minimum inhibitory concentration (MIC) of 0.5–32 µg/mL (including against resistant bacteria). Researchers evaluated the synergistic effect of SLAP-S25 and antibacterial drugs (tetracycline, vancomycin, ofloxacin, ampicillin, imipenem, rifampicin, or polymyxin) by the checkerboard method against polymyxin-resistant K. pneumoniae (mcr-1, mcr-6) and demonstrated a strong synergistic effect (ΣFIC < 0.5) for rifampicin (0.065), ofloxacin (0.127), and tetracycline (0.129). The synergistic treatment effectively reduced the number of colonies in the thighs of mice compared with that of the polymyxin-treated group. In the bacteremia experiment in mice, treatment with SLAP-S25 increased the survival rate, and no hemolytic activity was detected. Therefore, SLAP-S25 demonstrated antimicrobial activity in vivo and in vitro against resistant K. pneumoniae. When evaluating the interaction between SLAP-S25 and lipopolysaccharides (LPSs), the researchers identified that SLAP-S25 acted in a similar way to LPS-targeted antibiotics (Song et al. 2020).
Similarly, "US2020323950A1" and "WO2020014642A2" (Kraus and Otvos 2020a, b) patent applications are about combinations of antibacterial peptide monomers and analogs derived from antibiotics; the applications also include methods of using the combination to increase the sensitivity of antibiotic-resistant bacteria to an antibiotic, thereby broadening the therapeutic index of the AMP.
Proline-rich peptide dimers, such as A3-APO, exhibit antibacterial properties, and their potential increases when used in combination with antibiotics. Thus, "US2020323950A1" (Kraus and Otvos 2020b) patent application comprises 8 peptides with a composition of A3-APO analogs, derivatives or oligomers there of, or a pharmaceutically acceptable salt and an antibiotic. The activity of the peptide against MDR K. pneumoniae K97/09 (resistant to ceftazidime, ceftriaxone, imipenem, meropenem, ciprofloxacin, gentamicin, and colistin) was determined in vitro. The MIC value of A3-APO alone against polymyxin-resistant K. pneumoniae was 32 mg/L; A3-APO exhibited synergism when combined with colistin (FIC = 0.08), and an additive activity for the A3-APO/imipenem combination (FIC = 0.53) was also observed. In vivo, the combination of A3-APO with colistin was tested in a mouse model with systemic K. pneumoniae infection. In the same model, the combination of A3-APO significantly reduced colistin doses and prolonged their survival. A3-APO monotherapy at 0.5 mg/kg or 1.0 mg/kg resulted in a 20%–40% prolonged survival and reduction in blood bacterial counts.
The monomer peptides described herein, when administered with other antibiotics, are highly effective against bacterial infections, either alone or in combination with other antibiotics. The patent application "WO2020014642A2"(Kraus and Otvos 2020a) comprises monomer peptides, such as Chexl-Arg20 (commercially being developed as ARV-1502) or its dimeric form, A3-APO. The activity of A3-APO against polymyxin-resistant K. pneumoniae (K97/09) was determined in vitro, and an MIC value of 32 mg/L was obtained. The mechanism of action of A3-APO or ARV-1502 is non-membrane disruptive and is mostly related to the activation of host defense mechanisms rather than direct bacterial killing (Ostorhazi et al. 2011; Otvos Jr. et al. 2018).
Likewise, "WO2019200378A1" and "WO2017172929A1"(Kao 2017; Kao et al. 2019) patent applications are about peptide compositions with bactericidal activities; they describe a method of treating bacterial infections using compositions of antimicrobial peptides or variants. Thus, "WO2019200378A1"(Kao et al. 2019) provides antimicrobial peptide compositions that are selected from the peptide group consisting of SEQ ID NOs: 1–11. The peptide named B22a was tested against 20 strains of Gram-negative bacteria by using the broth dilution assay to determine the MIC; results demonstrated that it has antimicrobial activity against members of the Enterobacteriaceae. B22a resulted in only a two-fold increase in MIC value (8 µM) when tested against polymyxin-resistant K. pneumoniae. Thus, B22a inhibits the growth of MDR clinical isolates, including polymyxin-resistant ones. Additionally, the peptide named PB22N (MIC: 8 µM) was also effective in inhibiting polymyxin-resistant clinical isolates of K. pneumoniae.
This invention “WO2017172929A1” (Kao 2017) identifies compositions for antimicrobial activities that are selected from the group of peptides (cathelicidin family) consisting of SEQ ID NOs: 11–13 (BMAP-27A, BMAP-27B, and BMAP-27C), SEQ ID NOs: 15–17 (SMAP-29B, SMAP-29C, and SMAP-29D), and SEQ ID NOs. 18–19 (BMAP-24 and B22); a total of 8 peptides. Of these, only 2 peptides, named BMAP-27B and SMAP-29D, effectively kill colistin-resistant bacteria. Furthermore, peptides B22 and BMAP-24 exhibited MICs of 2 μΜ and 4 μΜ against carbapenem and polymyxin-resistant K. pneumoniae, respectively. The results indicated that B22 and BMAP-24 inhibited colistin and carbapenem-resistant K. pneumoniae. Probably, the differences in the mechanism of action between cathelicidins and colistin are responsible for BMAP-27B and SMAP-29D-mediated killing of colistin-resistant bacteria. BMAP-27B and SMAP-29D bind to bacterial membrane lipids through basic amino acids, whereas colistin binds to bacterial membranes through its N-terminal hydrophobic region and positive regions (Velkov et al. 2013; Kao et al. 2016).
In addition, "WO2020014501A1" and "WO2019126353A2" (Balkovec et al. 2020) are about peptide compositions, including compounds containing a cyclic heptapeptide or conjugates, that can be used in the treatment of infections caused by Gram-negative bacteria. In "WO2020014501A1"(Akers-Rodriguez et al. 2019; Balkovec et al. 2020) 46 compounds containing a cyclic heptapeptide are described. In the MIC test against K. pneumoniae (polymyxin-resistant clinical isolate having a phoQ mutation [T244N]), compounds 10, 37, 47, and 39 exhibited MICs of 16, 32, 32, and 64 ug/mL, respectively. The results revealed the antimicrobial activity of these compounds against polymyxin-resistant K. pneumoniae.
In the invention "WO2019126353A2", (Akers-Rodriguez et al. 2019) 79 conjugates, containing monomers or dimers of cyclic heptapeptides, are described. In the evaluation of in vitro activity through the MIC test against K. pneumoniae (polymyxin-resistant clinical isolate carrying a mutation in phoQ [T244N]), conjugates 23, 38, and 48 exhibited MICs of 16 µg/mL; the conjugates 25, 30, 31, 34, 35, 40, 43, 44, 48, 49, 51, 52, 53, 57, 59, 60, and 77 exhibited MICs of 32 µg/mL. The results revealed the antimicrobial activity of 20 of these conjugates against polymyxin-resistant K. pneumoniae. However, in vivo test was not performed.
The “WO2019028463A1” (Huang et al. 2019) refers to Paenipeptin analogs that are novel synthetic linear lipopeptides consisting of a lipophilic terminus, an amino terminus, and a peptide interposed between the lipophilic terminus and the amino terminus or a salt. The 17 lipopeptide analogs mentioned in the application are compositionally similar to a natural mixture of linear and cyclic lipopeptides produced by Paenibacillus sp. The antibacterial activities of these 17-synthetic linear lipopeptides analogs were determined against reference strains, carbapenem-resistant isolates, and 6 polymyxin-resistant strains. The linear lipopeptide 17 exhibited potent activity with an MIC of 0.5–2 µg/mL against carbapenem-resistant clinical isolates (including A. baumannii, Enterobacter cloacae, Escherichia coli, K. pneumoniae, and P. aeruginosa). In addition, the linear lipopeptide 17 was active against polymyxin-resistant strains of E. coli (n = 3) and K. pneumoniae (n = 3), thereby exhibiting potent in vitro activity. However, it was not evaluated in vivo. Paenipeptin analogs exhibit a high affinity for LPS on the outer membrane of Gram-negative bacteria. The results demonstrated that the bactericidal activity of paenipeptins is linked to the disruption and damage of cytoplasmic membranes, as the paenipeptin analog 17 depolarizes the membrane potential (Moon et al. 2017).
“WO2019084628A1” (Cooper et al. 2019) mentions novel cyclic peptide compounds. A total of 171 peptides were tested in vitro and demonstrated antimicrobial efficacy against different strains of MDR bacteria, including polymyxin-resistant P. aeruginosa (n = 2), A. baumannii (n = 1), and K. pneumoniae (n = 2). These compounds demonstrated advantageous properties when used in combination with other antibiotics (rifampicin and minocycline), thereby exhibiting synergy (FIC ≤ 0.5).
The patent application with the registration of “CN110582507A” (Steckback 2019) focuses on synthesized peptides consisting of a polypeptide sequence of 13 formulations, named SEQ ID NO: 1 to SEQ ID NO: 13. In vitro evaluation of peptide SEQ ID NO: 1 revealed its antimicrobial activity against MDR K. pneumoniae isolates (n = 101; 28 polymyxin-resistant ones). Peptide SEQ ID NO: 1 demonstrated an MIC of 8 µg/mL against most isolates; against polymyxin-resistant K. pneumoniae, the MIC ranged from 4 to 16 µg/mL. The MIC 50/90 was observed to be 8/16 µg/mL. The “US2020207821A1” describes numerous sequences of stabilized antimicrobial peptides (StAMP) that show antimicrobial activity against Gram-negative colistin-resistant bacteria (eg A. baumannii, E. coli, K. pneumoniae) with MIC less than 10.0 μg/mL(Walensky and Mourtada 2020).
The patent “WO2022173981A1” refers to the amphiphilic peptide named SEQ ID NO:1, which showed antimicrobial activity against multidrug resistant strains of K. pneumoniae (n = 101), including polymyxin resistant (27.7%), demonstrating MIC between 2 to > 16 pg/ml and MIC50/90 of 8/ > 16 pg/ml (Steckbeck et al. 2022). The patent application with the registration number “CN111386283A” (Smith et al. 2020) are peptide inhibitors, the final molecule, named G0775. focused on peptide inhibitor G0775. The MIC of G0775 was determined against a group of 49 clinical isolates of MDR E. coli and K. pneumoniae.
In the antimicrobial evaluation, G0775 maintained effective activity against all 49 multidrug-resistant isolates, including polymyxin-resistant K. pneumoniae (n = 1). In the antimicrobial evaluation, G0775 maintained effective activity against all 49 MDR isolates, including polymyxin-resistant K. pneumoniae (n = 1). In vivo efficacy was evaluated using a bacterial lung infection model (polymyxin-resistant K. pneumoniae); results demonstrated its bacteriostatic (2 mg/kg) and bactericidal (20 mg/kg) effects. No toxicity was observed in mammalian cells. The mechanism of action of G0775 is through the inhibition of essential bacterial signal peptidase type I (Smith et al. 2018).
Our results indicated that academic sectors possess the highest number of patents published against K. pneumoniae (Fig. 3A). This may be due to improvements in research and increased knowledge of intellectual property in academia. However, regarding patent filings containing AMPs against polymyxin-resistant K. pneumoniae, companies filed more than universities, with 8 and 6 patents, respectively. The possible explanation (especially for universities) may be related to the lack of financial resources to cover patent registration and maintenance fees. Regarding the status of these peptide patent registrations, only 2 are legally active (for example, US2020207821A1; CN108467424A); while 9 were archived; 2 are pending and 1 abandoned (Table 1). Evidencing that few patents were granted, and most were archived, probably due to non-payment of an annuity. In conclusion, the number of studies aiming at the development of new molecules and products to combat MDR infections is increasing steadily. All evaluated AMPs were in the experimental or preclinical phase; therefore, the clinical phase remains pending. In conclusion, research and patent filings alone are not enough to develop adequate antimicrobial therapy, and more investment is required to push these pharmaceuticals through the development stages to introduce them into the market.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Akers-Rodriguez S, Balkovec J, Bensen D, et al (2019) Compositions and methods for the treatment of bacterial infections. WO2019126353A2
Annunziato G, Costantino G (2020) Antimicrobial peptides (AMPs): a patent review (2015–2020). Expert Opin Ther Pat 30:931–947. https://doi.org/10.1080/13543776.2020.1851679
Bahar A, Ren D (2013) Antimicrobial peptides. Pharmaceuticals 6:1543–1575. https://doi.org/10.3390/ph6121543
Balkovec JM, Blizzard T, Borchardt A, et al (2020) Compositions and methods for the treatment of bacterial infections. WO2020014501A1
Bin Hafeez A, Jiang X, Bergen PJ, Zhu Y (2021) Antimicrobial peptides: an update on classifications and databases. IJMS 22:11691. https://doi.org/10.3390/ijms222111691
Cardoso MH, Orozco RQ, Rezende SB et al (2020) Computer-aided design of antimicrobial peptides: are we generating effective drug candidates? Front Microbiol 10:3097. https://doi.org/10.3389/fmicb.2019.03097
Cooper M, Blaskovich M, Gallardo-Goday A, et al (2019) Peptide antibiotics. WO2019084628A1
Da Silva KE, Thi Nguyen TN, Boinett CJ, et al (2020) Molecular and epidemiological surveillance of polymyxin-resistant Klebsiella pneumoniae strains isolated from Brazil with multiple mgrB gene mutations. Int J Med Microbiol 310:151448. https://doi.org/10.1016/j.ijmm.2020.151448
De La Cadena E, Mojica MF, García-Betancur JC et al (2021) Molecular analysis of polymyxin resistance among carbapenemase-producing Klebsiella pneumoniae in Colombia. Antibiotics 10:284. https://doi.org/10.3390/antibiotics10030284
De Oliveira DMP, Forde BM, Kidd TJ et al (2020) Antimicrobial resistance in ESKAPE pathogens. Clin Microbiol Rev 33:e00181-e219. https://doi.org/10.1128/CMR.00181-19
Deshayes C, Arafath MdN, Apaire-Marchais V, Roger E (2022) Drug Delivery systems for the oral administration of antimicrobial peptides: promising tools to treat infectious diseases. Front Med Technol 3:778645. https://doi.org/10.3389/fmedt.2021.778645
Farha MA, Brown ED (2019) Drug repurposing for antimicrobial discovery. Nat Microbiol 4:565–577. https://doi.org/10.1038/s41564-019-0357-1
Felício MR, Silveira GGOS, Oshiro KGN et al (2021) Polyalanine peptide variations may have different mechanisms of action against multidrug-resistant bacterial pathogens. J Antimicrob Chemother 76:1174–1186. https://doi.org/10.1093/jac/dkaa560
Han Y-J (2007) Measuring industrial knowledge stocks with patents and papers. J Informetr 1:269–276. https://doi.org/10.1016/j.joi.2007.06.001
Hansen GT (2021) Continuous evolution: perspective on the epidemiology of carbapenemase resistance among enterobacterales and other gram-negative bacteria. Infect Dis Ther 10:75–92. https://doi.org/10.1007/s40121-020-00395-2
Huang E, Moon S, Smeltzer M, Meeker D (2019) Linear lipopeptide paenipeptins and methods of using the same. WO2019028463A1
Kao C (2017) Bactericidal peptides and uses thereof. WO2017172929A1
Kao C, Lin X, Yi G, et al (2016) Cathelicidin antimicrobial peptides with reduced activation of toll-like receptor signaling have potent bactericidal activity against colistin-resistant bacteria. mBio 7:e01418–16. https://doi.org/10.1128/mBio.01418-16
Kao C, Prieto AC, Rowe-Magnus D (2019) Bactericidal peptides and uses thereof. WO2019200378A1
Kraus CN, Otvos L (2020a) Antibacterial peptide monomers and combinations for co-therapy. WO2020a014642A2
Kraus CN, Otvos L (2020b) Antibacterial peptides and combinations for co-therapy. US2020b323950A1
Krauss J, Kuttenkeuler D (2021) When to file for a patent? The scientist’s perspective. N Biotechnol 60:124–129. https://doi.org/10.1016/j.nbt.2020.10.006
Kui Z, Jianzhong S, Shuangyang D, et al (2018) Linear antibacterial oligopeptide SLAP-S25 and application thereof. CN108467424A
Levin AS, Oliveira MS (2008) The challenge of multidrug resistance: the treatment of Gram-negative rod infections. Shock 30:30–33. https://doi.org/10.1097/SHK.0b013e3181819cb8
Li J, Velkov T, Nation RL, Thompson PE (2010) Antimicrobial compounds. WO2010130007A1
Li J, Tang M, Xia F et al (2022) Emergence of polymyxin B-heteroresistant hypervirulent Klebsiella pneumoniae from an individual in the community with asymptomatic bacteriuria. BMC Microbiol 22:47. https://doi.org/10.1186/s12866-022-02462-9
Liu Y-Y, Wang Y, Walsh TR et al (2016) Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16:161–168. https://doi.org/10.1016/S1473-3099(15)00424-7
Martin RM, Bachman MA (2018) Colonization, infection, and the accessory genome of Klebsiella pneumoniae. Front Cell Infect Microbiol 8:4. https://doi.org/10.3389/fcimb.2018.00004
Moon SH, Zhang X, Zheng G et al (2017) Novel linear lipopeptide paenipeptins with potential for eradicating biofilms and sensitizing gram-negative bacteria to rifampicin and clarithromycin. J Med Chem 60:9630–9640. https://doi.org/10.1021/acs.jmedchem.7b01064
Muttenthaler M, King GF, Adams DJ, Alewood PF (2021) Trends in peptide drug discovery. Nat Rev Drug Discov 20:309–325. https://doi.org/10.1038/s41573-020-00135-8
Nainu F, Permana AD, Djide NJN et al (2021) Pharmaceutical approaches on antimicrobial resistance: prospects and challenges. Antibiotics 10:981. https://doi.org/10.3390/antibiotics10080981
Navon-Venezia S, Kondratyeva K, Carattoli A (2017) Klebsiella pneumoniae: a major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol Rev 41:252–275. https://doi.org/10.1093/femsre/fux013
Ostorhazi E, Holub MC, Rozgonyi F et al (2011) Broad-spectrum antimicrobial efficacy of peptide A3-APO in mouse models of multidrug-resistant wound and lung infections cannot be explained by in vitro activity against the pathogens involved. Int J Antimicrob Agents 37:480–484. https://doi.org/10.1016/j.ijantimicag.2011.01.003
Otvos L Jr, Ostorhazi E, Szabo D et al (2018) Synergy between proline-rich antimicrobial peptides and small molecule antibiotics against selected gram-negative pathogens in vitro and in vivo. Front Chem 6:309. https://doi.org/10.3389/fchem.2018.00309
Page MJ, McKenzie JE, Bossuyt PM, et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372:n71: https://doi.org/10.1136/bmj.n71
Rodríguez AA, Otero-González A, Ghattas M, Ständker L (2021) Discovery, optimization, and clinical application of natural antimicrobial peptides. Biomedicines 9:1381. https://doi.org/10.3390/biomedicines9101381
Rojas LJ, Salim M, Cober E et al (2016) Colistin resistance in carbapenem-resistant Klebsiella pneumoniae: laboratory detection and impact on mortality. Clin Infect Dis 64:711–718. https://doi.org/10.1093/cid/ciw805
Serafini MR, Santos VV, Torres BGS et al (2021) A patent review of antibiofilm fungal drugs (2002-present). Crit Rev Biotechnol 41:229–248. https://doi.org/10.1080/07388551.2021.1874283
Shi J, Chen C, Wang D et al (2021) Amphipathic peptide antibiotics with potent activity against multidrug-resistant pathogens. Pharmaceutics 13:438. https://doi.org/10.3390/pharmaceutics13040438
Smith PA, Koehler MFT, Girgis HS et al (2018) Optimized arylomycins are a new class of Gram-negative antibiotics. Nature 561:189–194. https://doi.org/10.1038/s41586-018-0483-6
Smith PA, Murray JM, Koehler MFT, Heise CE (2020) Peptide antibiotic complexes and methods of use thereof. CN111386283A
Soman R, Bakthavatchalam YD, Nadarajan A et al (2021) Is it time to move away from polymyxins?: evidence and alternatives. Eur J Clin Microbiol Infect Dis 40:461–475. https://doi.org/10.1007/s10096-020-04053-w
Song M, Liu Y, Huang X et al (2020) A broad-spectrum antibiotic adjuvant reverses multidrug-resistant Gram-negative pathogens. Nat Microbiol 5:1040–1050. https://doi.org/10.1038/s41564-020-0723-z
Steckback JD (2019) Engineered antimicrobial amphiphilic peptides and methods of use. CN110582507A
Steckbeck JD, Dobbins D, Huang D (2022) Intravenous administration of engineered antimicrobial amphiphilic peptides. WO2022173981A1
Stephens LJ, Werrett MV, Sedgwick AC et al (2020) Antimicrobial innovation: a current update and perspective on the antibiotic drug development pipeline. Future Med Chem 12:2035–2065. https://doi.org/10.4155/fmc-2020-0225
Velkov T, Roberts KD, Nation RL et al (2013) Pharmacology of polymyxins: new insights into an ‘old’ class of antibiotics. Future Microbiol 8:711–724. https://doi.org/10.2217/fmb.13.39
Walensky LD, Mourtada R (2020) Stabilized anti-microbial peptides for the treatment of antibiotic-resistant bacterial infections. US2020207821A1
Wang S, Zeng X, Yang Q, Qiao S (2016) Antimicrobial peptides as potential alternatives to antibiotics in food animal industry. IJMS 17:603. https://doi.org/10.3390/ijms17050603
Wang G, Zhao G, Chao X et al (2020) The characteristic of virulence, biofilm and antibiotic resistance of Klebsiella pneumoniae. IJERPH 17:6278. https://doi.org/10.3390/ijerph17176278
World Intellectual Property (2022) WIP publishing international patent classification
Wyres KL, Lam MMC, Holt KE (2020) Population genomics of Klebsiella pneumoniae. Nat Rev Microbiol 18:344–359. https://doi.org/10.1038/s41579-019-0315-1
Yang X, Dong N, Chan EW-C et al (2021) Carbapenem resistance-encoding and virulence-encoding conjugative plasmids in Klebsiella pneumoniae. Trends Microbiol 29:65–83. https://doi.org/10.1016/j.tim.2020.04.012
You D-G, Lee H-R, Kim H-K et al (2021) A novel peptide derived from the transmembrane domain of Romo1 is a promising candidate for sepsis treatment and multidrug-resistant bacteria. IJMS 22:8243. https://doi.org/10.3390/ijms22158243
Zhang Q-Y, Yan Z-B, Meng Y-M et al (2021a) Antimicrobial peptides: mechanism of action, activity and clinical potential. Military Med Res 8:48. https://doi.org/10.1186/s40779-021-00343-2
Zhang Y, Lin Y, Zhang X et al (2021b) Combining colistin with furanone C-30 rescues colistin resistance of gram-negative bacteria in vitro and in vivo. Microbiol Spectr 9:e01231-e1321. https://doi.org/10.1128/Spectrum.01231-21
Acknowledgements
The authors are grateful for financial support from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Apoio ao Desenvolvimento do Ensino, Ciência e Tecnologia do Estado de Mato Grosso do Sul (FUNDECT) and Universidade Federal da Grande Dourados. GHAS received a research grant from CAPES, ARO and SS from CNPq.
Funding
Funding was provided by Conselho Nacional de Desenvolvimento Científico e Tecnológico (408778/2022-9), Fundação de Apoio ao Desenvolvimento do Ensino,Ciência e Tecnologia do Estado de Mato Grosso do Sul (71/031.898/2022) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (001).
Author information
Authors and Affiliations
Contributions
GHAS and LR conducted the formal analysis. ARO contributed the methodology and data review. GHAS, LR and SS wrote the manuscript. SS and LR conceived, designed and supervised the research. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest, financial or otherwise.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
de Souza, G.H.d., Rossato, L., de Oliveira, A.R. et al. Antimicrobial peptides against polymyxin-resistant Klebsiella pneumoniae: a patent review. World J Microbiol Biotechnol 39, 86 (2023). https://doi.org/10.1007/s11274-023-03530-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03530-6