Abstract
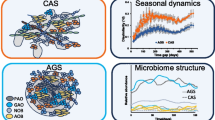
Bacteria play crucial roles in the combined system of substrate addition and C/N control, which has been demonstrated to improve aquaculture production. However, the complexity of surface-attached bacteria on substrates and suspended bacteria in the water column hamper further application of this system. This study firstly applied this combined system into the culture of grass carp, and then explored the relationship between microbial complexes from surface-attached and suspended bacteria in this system and the production of grass carp. In addition, this study investigated bacterial community structures as affected by four C/N ratios using Illumina sequencing technology. The results demonstrated that the weight gain rate and specific growth rate of grass carp in the CN20 group (C/N ratio 20:1) were the highest (P < 0.05), and dietary supplementation of the microbial complex had positive effects on the growth of grass carp (P < 0.05). Sequencing data revealed that, (1) the proportions of Verrucomicrobiae and Rhodobacter (surface-attached), sediminibacterium (suspended), and emticicia (surface-attached and suspended) were much higher in the CN20 group compared with those in the other groups (P < 0.05); (2) Rhodobacter, Flavobacterium, Acinetobacter, Pseudomonas, Planctomyces, and Cloacibacterium might be important for the microbial colonization on substrates; (3) as the C/N ratio increased, proportions of Hydrogenophaga (surface-attached and suspended), Zoogloea, and Flectobacillus (suspended) increased, but proportions of Bacillus, Clavibacter, and Cellvibro (surface-attached and suspended) decreased. In summary, a combined system of substrate addition and C/N control increased the production of grass carp, and Verrucomicrobiae and Rhodobacter in the surface-attached bacterial community were potential probiotic bacteria that contributed to the enhanced growth of grass carp.



Similar content being viewed by others
References
Anand PSS, Kohli MPS, Roy SD, Sundaray JK, Kumar S, Sinha A, Pailan GH, Sukham MK (2013a) Effect of dietary supplementation of periphyton on growth performance and digestive enzyme activities in Penaeus monodon. Aquaculture 392:59–68
Anand PSS, Kumar S, Panigrahi A, Ghoshal TK, Dayal JS, Biswas G, Sundaray JK, De D, Raja RA, Deo AD, Pillai SM, Ravichandran P (2013b) Effects of C:N ratio and substrate integration on periphyton biomass, microbial dynamics and growth of Penaeus monodon juveniles. Aquacult Int 21(2):511–524
Asaduzzaman M, Wahab MA, Verdegem MCJ, Huque S, Salam MA, Azim ME (2008) C/N ratio control and substrate addition for periphyton development jointly enhance freshwater prawn Macrobrachium rosenbergii production in ponds. Aquaculture 280:117–123
Asaduzzaman M, Wahab MA, Verdegem MCJ, Adhikary RK, Rahman SMS, Azim ME, Verreth JAJ (2010a) Effects of carbohydrate source for maintaining a high C:N ratio and fish driven re-suspension on pond ecology and production in periphyton-based freshwater prawn culture systems. Aquaculture 301:37–46
Asaduzzaman M, Rahman MM, Azim ME, Islam MA, Wahab MA, Verdegem MCJ, Verreth JAJ (2010b) Effects of C/N ratio and substrate addition on natural food communities in freshwater prawn monoculture ponds. Aquaculture 306:127–136
Besemer K, Peter H, Logue JB, Langenheder S, Lindström ES, Tranvik LJ, Battin TJ (2012) Unraveling assembly of stream biofilm communities. ISME J 6(8):1459–1468
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6(8):1621–1624
Chiu KH, Liu WS (2014) Dietary administration of the extract of Rhodobacter sphaeroides WL-APD911 enhances the growth performance and innate immune responses of seawater red tilapia (Oreochromis mossambicus × Oreochromis niloticus). Aquaculture 418:32–38
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2013) Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 41:D633–D642
Crab R, Avnimelech Y, Defoirdt T, Bossier P, Verstraete W (2007) Nitrogen removal techniques in aquaculture for a sustainable production. Aquaculture 270(1):1–14
Croese E, Jeremiasse AW, Marshall IP, Spormann AM, Euverink GJW, Geelhoed JS, Stams AJM, Plugge CM (2014) Influence of setup and carbon source on the bacterial community of biocathodes in microbial electrolysis cells. Enzyme Microb Technol 61:67–75
Cui FM, Shi JJ, Lu ZZ (1999) Production of neutral bata-mannanase by Bacillus subtilis and its properties. Acta Microbiol Sin 39(1):60–63
Dang H, Lovell CR (2002) Numerical dominance and phylotype diversity of marine Rhodobacter species during early colonization of submerged surfaces in coastal marine waters as determined by 16S ribosomal DNA sequence analysis and fluorescence in situ hybridization. Appl Environ Microbiol 68(2):496–504
Delille A, Quiles F, Humbert F (2007) In situ monitoring of the nascent Pseudomonas fluorescens biofilm response to variations in the dissolved organic carbon level in low-nutrient water by attenuated total reflectance-Fourier transform infrared spectroscopy. Appl Environ Microbiol 73(18):5782–5788
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 1 6S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
De-Schryver P, Verstraete W (2009) Nitrogen removal from aquaculture pond water by heterotrophic nitrogen assimilation in lab-scale sequencing batch reactors. Bioresour Technol 100(3):1162–1167
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–998
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200
Gao L, Bao WY, Zhang TW, Shan HW, Ma S (2013) Effect of water C:N ratio on the growth, antagonism and C:N ratio of Bacillus, Lactic Acid Bacteria and Vibrio. Period Ocean Univ China 43(1):34–40
Gao C, Wang A, Wu WM, Yin Y, Zhao YG (2014) Enrichment of anodic biofilm inoculated with anaerobic or aerobic sludge in single chambered air-cathode microbial fuel cells. Bioresour Technol 167:124–132
Ghanbari M, Kneifel W, Domig KJ (2015) A new view of the fish gut microbiome: advances from next-generation sequencing. Aquaculture 448:464–475
Gjermansen M, Ragas P, Sternberg C, Molin S, Tolker-Nielsen T (2005) Characterization of starvation-induced dispersion in Pseudomonas putida biofilms. Environ Microbiol 7(6):894–904
Haque MR, Islam MA, Rahman MM, Shirin MF, Wahab MA, Azim ME (2013) Effects of C/N ratio and periphyton substrates on pond ecology and production performance in giant freshwater prawn Macrobrachium rosenbergii (De Man, 1879) and tilapia Oreochromis niloticus (Linnaeus, 1758) polyculture system. Aquac Res 12270:1–17
Liu C, Wang K, Jiang JH, Liu WJ, Wang JY (2015) A novel bioflocculant produced by a salt-tolerant, alkaliphilic and biofilm-forming strain Bacillus agaradhaerens C9 and its application in harvesting Chlorella minutissima. Biochem Eng J 93:166–172
Magoc T, Salzberg S (2011) FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963
McDougald D, Rice SA, Barraud N, Steinberg PD, Kjelleberg S (2012) Should we stay or should we go: mechanisms and ecological consequences for biofilm dispersal. Nat Rev Microbiol 10(1):39–50
Reti KL, Thomas MC, Yanke LJ, Selinger LB, Inglis GD (2013) Effect of antimicrobial growth promoter administration on the intestinal microbiota of beef cattle. Gut Pathog 5(1):1–17
Wang YB (2007) Effect of probiotics on growth performance and digestive enzyme activity of the shrimp Penaeus vannamei. Aquaculture 269:259–264
Wang Y (2011) Use of probiotics Bacillus coagulans, Rhodopseudomonas palustris and Lactobacillus acidophilus as growth promoters in grass carp (Ctenopharyngodon idella) fingerlings. Aquac Nutr 17(2):e372–e378
Wang G, Yu E, Xie J, Yu D, Li Z, Luo W, Qiu L, Zheng Z (2015) Effect of C/N ratio on water quality in zero-water exchange tanks and the biofloc supplementation in feed on the growth performance of crucian carp, Carassius auratus. Aquaculture 443:98–104
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PLoS One 7(2):e30440
Wu S, Tian JY, Gatesoupe FJ, Li WX, Zou H, Yang BJ, Wang GT (2013) Intestinal microbiota of gibel carp (Carassius auratus gibelio) and its origin as revealed by 454 pyrosequencing. World J Microbiol Biotechnol 29(9):1585–1595
Zhou Z, Liu Y, He S, Shi P, Gao X, Yao B, Ringø E (2009) Effects of dietary potassium diformate (KDF) on growth performance, feed conversion and intestinal bacterial community of hybrid tilapia (Oreochromis niloticus♀ × O. aureus♂). Aquaculture 291(1):89–94
Acknowledgments
This study was funded by the Modern Agro-industry Technology Research System (No. CARS-46-17), and Project No. B201201A05 from Oceanic Fisheries Technology Promotion Special Project of Guangdong Province.
Funding
This study was funded by the Modern Agro-industry Technology Research System (No. CARS-46-17), and Project No. B201201A05 from Oceanic Fisheries Technology Promotion Special Project of Guangdong Province.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Rights and permissions
About this article
Cite this article
Yu, E., Xie, J., Wang, J. et al. Surface-attached and suspended bacterial community structure as affected by C/N ratios: relationship between bacteria and fish production. World J Microbiol Biotechnol 32, 116 (2016). https://doi.org/10.1007/s11274-016-2065-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-016-2065-9