Abstract
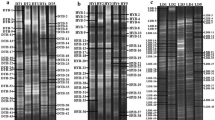
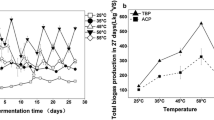
Anaerobic digestion (AD) is the most popular path of organic waste disposal. It is often used in wastewater treatment plants for excessive sludge removal. Methanogenic fermentation had usually been performed under mesophilic conditions, but in the past few years the thermophilic processes have become more popular due to economics and sludge sanitation. Methanogens, the group of microorganisms responsible for methane production, are thought to be sensitive to temperature change and it has already been proven that the communities performing methanogenesis under mesophilic and thermophilic conditions differ. But in most cases the research performed on methanogen diversity and changeability was undertaken in two separate anaerobic chambers for meso- and thermophilic conditions. It is also known that there is a group of microorganisms performing AD which are insensitive to temperature. Also the linkage between digester performance and its microbial content and community changeability is still not fully understood. That is why in this experiment we analyzed the bacterial community performing methanogenesis in a pilot scale anaerobic chamber during the shift from mesophilic to thermophilic conditions to point at the group of temperature tolerant microorganisms and their performance. The research was performed with PCR–DGGE (polymerase chain reaction–denaturing gradient gel electrophoresis). It occurred that the community biodiversity decreased together with a temperature increase. The changes were coherent for both the total bacteria community and methanogens. These bacterial shifts were also convergent with biogas production—it decreased in the beginning of the thermophilic phase with the bacterial biodiversity decrease and increased when the community seemed to be restored. DGGE results suggest that among a wide variety of microorganisms involved in AD there is a GC-rich group relatively insensitive towards temperature change, able to adapt quickly to shifts in temperature and perform AD effectively. The studies of this microbial group could be a step forward in developing more efficient anaerobic digestion technology.





Similar content being viewed by others
References
Ahring BK (1995) Methanogenesis in thermophilic biogas reactors. Antonie Van Leeuwenhoek 67(1):91–102
Ahring BK (2003) Perspectives for anaerobic digestion. Adv Biochem Eng Biotechnol 81:1–30
Ban Q, Li J, Zhang L, Zhang Y, Jha AK (2013) Phylogenetic diversity of methanogenic Archaea and kinetics of methane production at slightly acidic conditions of an anaerobic sludge. Int J Agric Biol 15:347–351
Briones A, Raskin L (2003) Diversity and dynamics of microbial communities in engineered environments and their implications for process stability. Curr Opin Biotechnol 14:270–276
Calli B, Mertoglu B, Inanc B, Yenigun O (2005) Methanogenic diversity in anaerobic reactors under extremely high ammonia levels. Enzyme Microb Technol 37:448–455
Fernández A, Huang S, Seston S, Xing J, Hickey R, Criddle C, Tiedje J (1999) How stable is stable? Function versus community composition. Appl Environ Microbiol 65:3697–3704
Goberna M, Insam H, Franke-Whittle IH (2005) Effect of biowaste sludge maturation on the diversity of thermophilic bacteria and archaea in an anaerobic reactor. Appl Environ Microbiol 75(8):2566–2572
Hernon F, Forbes C, Colleran E (2006) Identification of mesophilic and thermophilic fermentative species in anaerobic granular sludge. Water Sci Technol 54:19–24
Kim M, Speece RE (2002) Aerobic waste activated sludge (WAS) for start-up seed of mesophilic and thermophilic anaerobic digestion. Water Res 36:3860–3866
Leven L, Anders RBE, Schnurer A (2007) Effect of process temperature on bacerial and archeal communities in two methanogenic bioreactors treatind organic household waste. FEMS Microbiol Ecol 59:683–693
Ma K, Liu X, Dong X (2005) Methanobacterium beijingense sp. nov., a novel methanogen isolated from anaerobic digesters. Int J Syst Evol Microbiol 55:325–329
Muyzer G, De Waal EC, Uitierlnden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplifi ed genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Narihiro T, Sekiguchi Y (2007) Microbial communities in anaerobic digestion processes for waste and wastewater treatment: a microbiological update. Curr Opin Biotechnol 18:273–278
Ros MA, Franke-Whittle IH, Morales AB, Insam H, Ayuso M, Pascual JA (2013) Archaeal community dynamics and abiotic characteristics in a mesophilic anaerobic co-digestion process treating fruit and vegetable processing waste sludge with chopped fresh artichoke waste. Bioresour Technol 136:1–7
Sekiguchi Y, Kamagata Y, Syutsubo K, Ohashi A, Harada H, Nakamura K (1998) Phylogenetic diversity of mesophilic and thermophilic granular sludges determined by 16S rRNA gene analysis. Microbiology 144:2655–2665
Shi J, Zhongjiang W, Stiverson J, Zhongtang Y, Li Y (2013) Reactor performance and microbial community dynamics during solid-state anaerobic digestion of corn stover at mesophilic and thermophilic conditions. Bioresour Technol 136:574–581
Song BYJ, Woo JH, Kwon SJ, Koh SC (2006) Microbial activity and population structure of anaerobic sludge alternately exposed to mesophilic and thermophilic conditions. KSCE J Civ Eng 10:319–323
Zhou M, Hernandez-Sanabria E, Guan LL (2010) Characterization of variation in rumen methanogenic communities under dietary and host feed efficiency conditions, as determined by PCR-DGGE. Appl Environ Microbiol 76:3776–3786
Zumstein E, Moletta R, Godon JJ (2000) Examination of two years of community dynamics in an anaerobic bioreactor using fluorescence polymerase chain reaction (PCR) single-strand conformation polymorphism analysis. Environ Microbiol 2:69–78
Acknowledgments
This work was performed using the equipment purchased in the project “Silesian BIO-FARMA. The Center for Biotechnology, Bioengineering and Biocomputing”, EFRR PO IG, 2007–2013.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ziembińska-Buczyńska, A., Banach, A., Bacza, T. et al. Diversity and variability of methanogens during the shift from mesophilic to thermohilic conditions while biogas production. World J Microbiol Biotechnol 30, 3047–3053 (2014). https://doi.org/10.1007/s11274-014-1731-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-014-1731-z