Abstract
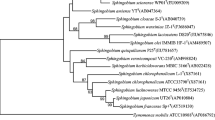
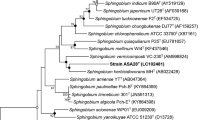
Three strains of bacteria (designated as YBL1, YBL2, YBL3 respectively) capable of degrading isoproturon, 3-(4-isopropylphenyl)-1, 1-dimethylurea, were isolated from the soils of two herbicide plants. Based on the comparative analysis of the 16S rRNA gene, and phenotypic and biochemical characterization, these strains were identified as Sphingobium sp. The optimum conditions for isoproturon degradation by these strains were pH 7.0, and temperature 30°C. Mg2+ (1 mM) enhanced the isoproturon degradation rate, while Ni2+ and Cu2+ (1 mmol l−1) inhibited isoproturon degradation significantly. These three strains also showed the ability to remove the residues of other phenylurea herbicides such as chlorotoluron, diuron and fluometuron in mineral salt culture medium. The N-demethylation was the first step of degradation of dimethylurea-substituted herbicides. Strain YBL1 was found capable of degrading both dimethylurea-substituted herbicides and methoxymethylphenyl-urea herbicides i.e. linuron (3-(3,4-dichlorophenyl)-1-methoxy-1-methylurea). Using the PCR method, partial sequences of the catechol 1,2-dioxygenase gene were obtained from these strains.








Similar content being viewed by others
References
Barles RM, Topp EE, Blackwell BA (1979) Accelerated parathion degradation in soil inoculated with acclimated bacteria under field conditions. Arch Environ Contam Toxicol 8:647–660. doi:10.1007/BF01054867
Bending GD, Lincoln SD, Sorensen SR (2003) In-field spatial variability in the degradation of the phenyl-urea herbicide isoproturon is the result of interactions between degradative Sphingomonas spp. and soil pH. Appl Environ Microbiol 69:827–834. doi:10.1128/AEM.69.2.827-834.2003
Bending GD, Lincoln SD, Edmondson RN (2006) Spatial variation in the degradation rate of the pesticides isoproturon, azoxystrobin and diflufenican in soil and its relationship with chemical and microbial properties. Environ Pollut 139:279–287. doi:10.1016/j.envpol.2005.05.011
Bending GD, Rodriguez-Cruz MS (2007) Microbial aspects of the interaction between soil depth and biodegradation of the herbicide isoproturon. Chemosphere 66:664–671. doi:10.1016/j.chemosphere.2006.07.099
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Celis E, Elefsiniotis P, Singhal N (2008) Biodegradation of agricultural herbicides in sequencing batch reactors under aerobic or anaerobic conditions. Water Res 42:3218–3224. doi:10.1016/j.watres.2008.04.008
Dai M, Copley SD (2004) Genome shuffling improves degradation of the anthropogenic pesticide pentachlorophenol by Sphingobium chlorophenolicum ATCC 39723. Appl Environ Microbiol 70:2391–2397. doi:10.1128/AEM.70.4.2391-2397.2004
Dejonghe W, Berteloot E, Goris J (2003) Synergistic degradation of linuron by a bacterial consortium and isolation of a single linuron-degrading variovorax strain. Appl Environ Microbiol 69:1532–1541. doi:10.1128/AEM.69.3.1532-1541.2003
El-Sebai TE, Lagacherie B, Soulas G (2004) Isolation and characterization of an isoproturon-mineralising Methylopela sp. TES from French agricultural soil. FEMS Microbiol Lett 239:103–110. doi:10.1016/j.femsle.2004.08.017
El-Sebai TE, Lagacherie B, Soulas G (2007) Spatial variability of isoproturon mineralizing activity within an agricultural field: geostatistical analysis of simple physicochemical and microbiological soil parameters. Environ Pollut 145:680–690. doi:10.1016/j.envpol.2006.05.034
Holt J, Krieg N, Sneath P (1994) Bergey’s manual of determinative bacteriology, 9th edn. Williams and Wilkins, Baltimore
Huang X, He J, Sun J (2007) Isolation and characterization of a metsulfuron-methyl degrading bacterium Methylopila sp. S113. Int Biodeter Biodegr 60:152–158. doi:10.1016/j.ibiod.2007.02.005
Johnson AC, Besien T, Bhardwaj C (2001) Penetration of herbicides to groundwater in an unconfined chalk aquifer following normal soil applications. J Contam Hydrol 53:101–117. doi:10.1016/S0169-7722(01)00139-5
Karpouzas DG, Walker A (2000) Factors influencing the ability of Pseudomonas putida epI to degrade ethoprophos in soil. Soil Biol Biochem 32:1753–1762. doi:10.1016/S0038-0717(00)00093-6
Kristensen KE, Jacobsen CS, Hansen LH (2006) Genetic labelling and application of the isoproturon mineralizing Sphingomonas sp. strain SRS2 in soil and rhizosphere. Lett Appl Microbiol 43:280–286. doi:10.1111/j.1472-765X.2006.01956.x
Kearney PC, Karns JS, Muldoon MT (1986) Coumaphos disposal by combined microbial and UV-ozonation reactions. J Agric Food Chem 34:702–706. doi:10.1021/jf00070a028
Mansour M, Feicht EA, Behechti A (1999) Determination photostability of selected agrochemicals in water and soil. Chemosphere 39:575–585. doi:10.1016/S0045-6535(99)00123-X
Mascolo G, Lopez A, James H (2001) By-products formation during degradation of IPU in aqueous solution II: chlorination. Water Res 35:1705–1713. doi:10.1016/S0043-1354(00)00428-0
Morvan X, Mouvet C, Baran N (2006) Pesticides in the groundwater of a spring draining a sandy aquifer: temporal variability of concentrations and fluxes. J Contam Hydrol 87:176–190. doi:10.1016/j.jconhyd.2006.05.003
Ngai K-L, Neidle EL, Ornston CN (1990) Catechol and chlorocatechol 1,2-dioxygenase. Methods Enzymol 188:122–126. doi:10.1016/0076-6879(90)88022-3
Ostrofsky E, Traina S, Tuovinen O (1997) Variation in atrazine mineralization rates in relation to agricultural management practice. J Environ Qual 26:647–657
Peres F, Florin D, Grollier T (1996) Effect of the phenylurea herbicide isoproturon on periphytic diatom communities in freshwater indoor microcosms. Environ Pollut 94:141–152. doi:10.1016/S0269-7491(96)00080-2
Prakash O, Lal R (2006) Description of Sphingobium fuliginis sp. nov., a phenanthrene-degrading bacterium from a fly ash dumping site, and reclassification of Sphingomonas cloacae as Sphingobium cloacae comb.nov. Int J Syst Evol Microbiol 56:2147–2152. doi:10.1099/ijs.0.64080-0
Pussemier L, Goux S, Vanderheyden V (1997) Rapid dissipation of atrazine in soils taken from various maize fields. Weed Res 37:171–179. doi:10.1046/j.1365-3180.1997.d01-18.x
Radosevich M, Traina SJ, Hao YL (1995) Degradation and mineralization of atrazine by a soil bacterial isolate. Appl Environ Microbiol 61:297–302
Ronhede S, Jensen B, Rosendahl S (2005) Hydroxylation of the herbicide isoproturon by fungi isolated from agricultural soil. Appl Environ Microbiol 71:7927–7932. doi:10.1128/AEM.71.12.7927-7932.2005
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sharma P, Raina V, Kumari R (2006) Haloalkane dehalogenase linB is responsible for beta- and delta-hexachlorocyclohexane transformation in Sphingobium indicum B90A. Appl Environ Microbiol 72:5720–5727. doi:10.1128/AEM.00192-06
Sorensen SR, Ronen Z, Aamand J (2001) Isolation from agricultural soil and characterization of a Sphingomonas sp. able to mineralize the phenylurea herbicide isoproturon. Appl Environ Microbiol 67:5403–5409. doi:10.1128/AEM.67.12.5403-5409.2001
Sorensen SR, Albers CN, Aamand J (2008) Rapid mineralization of the phenylurea herbicide diuron by Variovorax sp. strain SRS16 in pure culture and within a two-member consortium. Appl Environ Microbiol 74:2332–2340. doi:10.1128/AEM.02687-07
Sorensen SR, Bending GD, Jacobsen CS (2003) Microbial degradation of isoproturon and related phenylurea herbicides in and below agricultural fields. FEMS Microbiol Ecol 45:1–11. doi:10.1016/S0168-6496(03)00127-2
Sun J, Huang X, He J (2006) Isolation identification of isoproturon degradation bacterium Y57 and its degradation characteristic. China Environ Sci 26:315–319 in Chinese
Tamura K, Dudley J, Nei M (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Topp E, Zhu H, Nour SM (2000) Characterization of an atrazine-degrading Pseudaminobacter sp. isolated from Canadian and French agricultural soils. Appl Environ Microbiol 66:2773–2782. doi:10.1128/AEM.66.7.2773-2782.2000
Turnbull GA, Ousley M, Walker A (2001) Degradation of substituted phenylurea herbicides by Arthrobacter globiformis strain D47 and characterization of a plasmid-associated hydrolase gene, puhA. Appl Environ Microbiol 67:2270–2275. doi:10.1128/AEM.67.5.2270-2275.2001
Walker A, Jurado-Exposito M, Bending G (2001) Spatial variability in the degradation rate of isoproturon in soil. Environ Pollut 111:407–427. doi:10.1016/S0269-7491(00)00092-0
Walker A, Bromilow RH, Nicholls PH (2008) Spatial variability in the degradation rates of isoproturon and chlorotoluron in a clay soil. Weed Res 42:39–44. doi:10.1046/j.1365-3180.2002.00260.x
Widehem P, Ait-Aissa S, Tixier C (2002) Isolation, characterization and diuron transformation capacities of a bacterial strain Arthrobacter sp. N2. Chemosphere 46:527–534. doi:10.1016/S0045-6535(01)00192-8
Widenfalk A, Bertilsson S, Sundh I (2008) Effects of pesticides on community composition and activity of sediment microbes—responses at various levels of microbial community organization. Environ Pollut 152:576–584. doi:10.1016/j.envpol.2007.07.003
Widenfalk A, Goedkoop W, Svensson JM (2004) Effects of the pesticides captan, deltamethrin, isoproturon, and pirimicarb on the microbial community of a freshwater sediment. Environ Toxicol Chem 23:1920–1927. doi:10.1897/03-345
Zhang XH, Zhang GH, Li SP (2006) Isolation and characterization of a dichlorvos-degrading strain DDV-1 of Ochrobactrum sp. Pedosphere 16:64–71. doi:10.1016/S1002-0160(06)60027-1
Acknowledgments
This work was supported by Agricultural Technology Transfer Program (grant 2007.100) and National Foundation Program for Nature Resources of Science and Technology (2005DKA21201-11).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, JQ., Huang, X., Chen, QL. et al. Isolation and characterization of three Sphingobium sp. strains capable of degrading isoproturon and cloning of the catechol 1,2-dioxygenase gene from these strains. World J Microbiol Biotechnol 25, 259–268 (2009). https://doi.org/10.1007/s11274-008-9888-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9888-y