Abstract
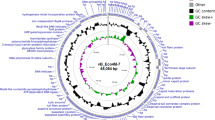
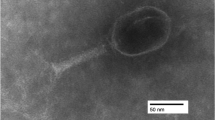
Bacteriophages represent one prospect for preventing and treating multi-drug-resistant Escherichia coli. In this study, we have isolated a novel E. coli-specific bacteriophage and characterised its biological properties. vB_EcoM-ep3 has a broad host range and was able to lyse 9 out of 15 clinical isolates of multi-drug-resistant pathogenic E. coli from chickens. The optimal multiplicity of infection for vB_EcoM-ep3 in host bacteria was 0.01. vB_EcoM-ep3 was thermostable at temperatures below 50 °C for up to 60 min. Electron microscopy demonstrated that vB_EcoM-ep3 belongs to Myoviridae. The vB_EcoM-ep3 genome contained 42,351 pairs of nucleotides with a GC content of 53.35 %. There were 52 predicted open reading frames that appeared to overlap and have a modular structure. Phylogenetic analysis indicates that the closest evolutionary relative to vB_EcoM-ep3 is the previously reported E. coli phage vB_EcoM_ECO1230-10. However, there was no homology between reported E. coli phage lysins and the vB_EcoM-ep3 lysin gene. Lysep3 was 58 % similar to the Pseudomonas phage PPpW-3 lysin despite showing no similarities at the gene sequence level. And Lysep3 has good lysis activity.












Similar content being viewed by others
References
G. Rocchi, M. Capozzi, Recenti Prog. Med. 90, 613–618 (1999)
J.L. Platell, R.N. Cobbold, J.R. Johnson, D.J. Trott, J. Antimicrob. Chemother. 65, 1936–1938 (2010)
E.R. Grela, Pol. J. Vet. Sci. 11, 405–409 (2008)
J.W. Oguttu, C.M. Veary, J.A. Picard, J. S. Afr. Vet. Assoc. 79, 161–166 (2008)
S. Matsuzaki, J. Uchiyama, I. Takemura-Uchiyama, M. Daibata, Nature 509, S9 (2014)
R.J. Payne, D. Phil, V.A. Jansen, Clin. Pharmacol. Ther. 68, 225–230 (2000)
C.R. Merril, B. Biswas, R. Carlton, N.C. Jensen, G.J. Creed, S. Zullo, S. Adhya, in Proceedings of the National Academy of Sciences USA, vol 93, pp. 3188–3192, 1996
Y. Tanji, T. Shimada, H. Fukudomi, K. Miyanaga, Y. Nakai, H. Unno, J. Biosci. Bioeng. 100, 280–287 (2005)
M.K. Waldor, D.I. Friedman, S.L. Adhya, Phages :their role in bacterial pathogenesis and biotechnology (ASM Press, Washington, DC, 2005)
D. Jabes, Curr. Opin. Microbiol. 14, 564–569 (2011)
J.N. Housby, N.H. Mann, Drug Discov. Today 14, 536–540 (2009)
N.T. Lin, P.Y. Chiou, K.C. Chang, L.K. Chen, M.J. Lai, Res. Microbiol. 161, 308–314 (2010)
D.W. Russell, J. Sambrook, Molecular cloning :a laboratory manual (Cold Spring Harbor Laboratory Press, New York, 2001)
P. Barrow, M. Lovell, A.J. Berchieri, Clin. Diagn. Lab. Immunol. 5, 294–298 (1998)
J. Gu, W. Xu, L. Lei, J. Huang, X. Feng, C. Sun, C. Du, J. Zuo, Y. Li, T. Du, L. Li, W. Han, J. Clin. Microbiol. 49, 111–117 (2011)
C.K. Mathews, Bacteriophage T4 (American Society for Microbiology, Washington, DC., 1983)
P. Hyman, S.T. Abedon, Methods Mol. Biol. 501, 175–202 (2009)
J. Klumpp, D.E. Fouts, S. Sozhamannan, Bacteriophage 2, 190–199 (2012)
C. Zhang, W. Li, W. Liu, L. Zou, C. Yan, K. Lu, H. Ren, Appl. Environ. Microbiol. 79, 5559–5565 (2013)
T.M. Santos, R.C. Bicalho, Vet. Microbiol. 148, 267–275 (2011)
J.W. Jun, S.K. Yun, H.J. Kim, J.Y. Chai, S.C. Park, Res. Microbiol. 165, 671–678 (2014)
J. Borysowski, B. Weber-Dabrowska, A. Gorski, Exp. Biol. Med. 231, 366–377 (2006)
P. Li, B. Chen, Z. Song, Y. Song, Y. Yang, P. Ma, H. Wang, J. Ying, P. Ren, L. Yang, G. Gao, S. Jin, Q. Bao, H. Yang, Gene 507, 125–134 (2012)
B. Son, J. Yun, J.A. Lim, H. Shin, S. Heu, S. Ryu, BMC Microbiol. 12, 33 (2012)
Y. Yuan, Q. Peng, M. Gao, BMC Microbiol. 12, 297 (2012)
H. Oliveira, V. Thiagarajan, M. Walmagh, S. Sillankorva, R. Lavigne, M.T. Neves-Petersen, L.D. Kluskens, J. Azeredo, PLoS One 9, e108376 (2014)
G.L. Lau, C.C. Sieo, W.S. Tan, M. Hair-Bejo, A. Jalila, Y.W. Ho, Poult. Sci. 89, 2589–2596 (2010)
G.L. Lau, C.C. Sieo, W.S. Tan, Y.W. Ho, J. Sci. Food Agric. 92, 2657–2663 (2012)
H. Li, M.L. Ma, H.J. Xie, J. Kong, World J. Microbiol. Biotechnol. 28, 1–6 (2012)
J.P. Higgins, S.E. Higgins, K.L. Guenther, W. Huff, A.M. Donoghue, D.J. Donoghue, B.M. Hargis, Poult. Sci. 84, 1141–1145 (2005)
T.H. Lim, M.S. Kim, D.H. Lee, Y.N. Lee, J.K. Park, H.N. Youn, H.J. Lee, S.Y. Yang, Y.W. Cho, J.B. Lee, S.Y. Park, I.S. Choi, C.S. Song, Res. Vet. Sci. 93, 1173–1178 (2012)
N. Janez, A. Kokosin, E. Zaletel, T. Vranac, J. Kovac, D. Vuckovic, M.S. Smole, S.V. Curin, Q. Zhang, T. Accetto, A. Podgornik, M. Peterka, FEMS Microbiol Lett (2014). doi:10.1111/1574-6968.12556
Acknowledgments
The study was supported by funds from the National Key Basic Research Programme of China (No. 2013CB127205). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by Joachim Jakob Bugert.
Rights and permissions
About this article
Cite this article
Lv, M., Wang, S., Yan, G. et al. Genome sequencing and analysis of an Escherichia coli phage vB_EcoM-ep3 with a novel lysin, Lysep3. Virus Genes 50, 487–497 (2015). https://doi.org/10.1007/s11262-015-1195-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-015-1195-8