Abstract
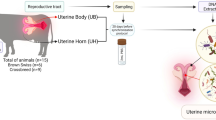
The present study, using 16 s rRNA sequencing of the V3-V4 hypervariable region, was aimed to check diversity of vaginal microbiota throughout different stages of the estrous cycle in Bos indicus, with attention to changes in progesterone hormone and microorganism diversity. Metagenomic research was conducted on vaginal swabs obtained from nine healthy Indian Gir cows at estrus (day 0), metestrus (day 04), diestrus (day 12), and proestrus (day 16) phases of the estrous cycle. The findings revealed that the diestrus phase has a different bacterial diversity than the other three estrous cycle phases, implying that progesterone affects bacterial diversity. Proteobacteria and Firmicutes were the most abundant phyla at the phylum level, accounting for 94% of bacterial diversity. Actinobacteriota, Patescibacteria, Cyanobacteria, and Bacteroidota were among the less prevalent phyla observed in all samples. After statistical analysis, Bacillaceae, Alcaligenes, Enterobacteriaceae, and Morganellaceae families were more significant. The Enterobacteriaceae family was found to be lower in the diestrus phase than in the other three phases; in contrast, all statistically significant genera were high at the diestrus phase. The luteal stage had higher levels of Micrococcus, Stenotrophomonas, UGC-010, Massilia, and Methylobacillus than the follicular stage, and statistical analysis revealed substantial difference between the luteal and follicular stages. Lactobacillus genus was present in both the estrus and diestrus phases. This study represents an important step toward the understanding of microbial diversity within different stages of the estrous cycle of Indian cows.







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Alves, L.F., Westmann, C.A., Lovate, G.L., de Siqueira, G.M.V., Borelli, T.C., and Guazzaroni, M.E., 2018. Metagenomic approaches for understanding new concepts in microbial science, International Journal of Genomics, 2312987.
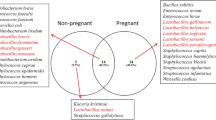
Ault, T.B., Clemmons, B.A., Reese, S.T., Dantas, F.G., Franco, G.A., Smith, T.P.L., Edwards, J.L., Myer, P.R., and Pohler, K.G., 2019. Uterine and vaginal bacterial community diversity prior to artificial insemination between pregnant and nonpregnant postpartum cows, Journal of Animal Science, 97, 4298-4304.
Baker, J.M., Chase, D.M., and Herbst-Kralovetz, M.M., 2018. Uterine Microbiota: residents, tourists, or invaders?, Frontier in Immunology, 9, 208.
Benjamini, Y., and Hochberg, Y., 1995. Controlling the False discovery rate: A practical and powerful approach to multiple testing, Journal of the Royal Statistical Society: Series B (Methodological), 57, 289-300.
Bolyen, E., Rideout, J.R., Dillon, M.R., Bokulich, N.A., Abnet, C.C., Al-Ghalith, G.A., Alexander, H., Alm, E.J., Arumugam, M., Asnicar, F., Bai, Y., Bisanz, J.E., Bittinger, K., Brejnrod, A., Brislawn, C.J., Brown, C.T., Callahan, B.J., Caraballo-Rodríguez, A.M., Chase, J., Cope, E.K., Da Silva, R., Diener, C., Dorrestein, P.C., Douglas, G.M., Durall, D.M., Duvallet, C., Edwardson, C.F., Ernst, M., Estaki, M., Fouquier, J., Gauglitz, J.M., Gibbons, S.M., Gibson, D.L., Gonzalez, A., Gorlick, K., Guo, J., Hillmann, B., Holmes, S., Holste, H., Huttenhower, C., Huttley, G.A., Janssen, S., Jarmusch, A.K., Jiang, L., Kaehler, B.D., Kang, K.B., Keefe, C.R., Keim, P., Kelley, S.T., Knights, D., Koester, I., Kosciolek, T., Kreps, J., Langille, M.G.I., Lee, J., Ley, R., Liu, Y.X., Loftfield, E., Lozupone, C., Maher, M., Marotz, C., Martin, B.D., McDonald, D., McIver, L.J., Melnik, A.V., Metcalf, J.L., Morgan, S.C., Morton, J.T., Naimey, A.T., Navas-Molina, J.A., Nothias, L.F., Orchanian, S.B., Pearson, T., Peoples, S.L., Petras, D., Preuss, M.L., Pruesse, E., Rasmussen, L.B., Rivers, A., Robeson, M.S., 2nd, Rosenthal, P., Segata, N., Shaffer, M., Shiffer, A., Sinha, R., Song, S.J., Spear, J.R., Swafford, A.D., Thompson, L.R., Torres, P.J., Trinh, P., Tripathi, A., Turnbaugh, P.J., Ul-Hasan, S., van der Hooft, J.J.J., Vargas, F., Vázquez-Baeza, Y., Vogtmann, E., von Hippel, M., Walters, W., Wan, Y., Wang, M., Warren, J., Weber, K.C., Williamson, C.H.D., Willis, A.D., Xu, Z.Z., Zaneveld, J.R., Zhang, Y., Zhu, Q., Knight, R., and Caporaso, J.G., 2019. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2, Nature Biotechnology, 37, 852-857.
Carmina, E., Stanczyk, F.Z., and Lobo, R.A., 2014. Chapter 34 - Laboratory Assessment. In: J.F. Strauss and R.L. Barbieri (eds), Yen & Jaffe’s Reproductive Endocrinology (Seventh Edition), 2014, (W.B. Saunders, Philadelphia), 822–850.e823.
Coleman, D.A., Thayne, W.V., and Dailey, R.A., 1985. Factors affecting reproductive performance of dairy cows, Journal of Dairy Science, 68, 1793-1803.
Cribby, S., Taylor, M., and Reid, G., 2008. Vaginal microbiota and the use of probiotics, Interdisciplinary Perspectives on Infectious Diseases, 2008, 256490.
Giannattasio-Ferraz, S., Maskeri, L., Oliveira André, P., Barbosa-Stancioli Edel, F., Putonti, C., and Newton Irene, L.G., 2020. Three draft genome sequences of staphylococcus species isolated from the urine of healthy bovine heifers (Gyr Breed), Microbiology Resource Announcements, 9, e00389-00320.
Laguardia-Nascimento, M., Branco, K.M.G.R., Gasparini, M.R., Giannattasio-Ferraz, S., Leite, L.R., Araujo, F.M.G., Salim, A.C.d.M., Nicoli, J.R., de Oliveira, G.C., and Barbosa-Stancioli, E.F., 2015. Vaginal microbiome characterization of Nellore cattle using metagenomic analysis, PLOS ONE, 10, e0143294-e0143294.
Machado, V.S., Oikonomou, G., Bicalho, M.L., Knauer, W.A., Gilbert, R., and Bicalho, R.C., 2012. Investigation of postpartum dairy cows’ uterine microbial diversity using metagenomic pyrosequencing of the 16S rRNA gene, Veterinary Microbiology, 159, 460-469.
Mahesh, P., Suthar, V., Patil, D., Joshi, M., Bagatharia, S., and Joshi, C., 2020. Vaginal microbiota during estrous cycle and its plausible association with certain hematological parameters in Bubalus bubalis, The Indian Journal of Veterinary Sciences and Biotechnology, 15, 54-58.
Mahesh, P., Suthar, V.S., Patil, D.B., Bagatharia, S.B., Joshi, M., and Joshi, C.G., 2021. Dynamics of vaginal microbiota during estrous cycle and its association with reproductive hormones in Bubalus bubalis, The Indian Journal of Veterinary Sciences and Biotechnology, 17, 11-18.
Mehta, O., Ghosh, T.S., Kothidar, A., Gowtham, M.R., Mitra, R., Kshetrapal, P., Wadhwa, N., Thiruvengadam, R., Nair, G.B., Bhatnagar, S., and Das, B., 2020. Vaginal microbiome of pregnant Indian women: Insights into the genome of dominant lactobacillus Species, Microbial Ecology, 80, 487-499.
Miller, E.A., Livermore, J.A., Alberts, S.C., Tung, J., and Archie, E.A., 2017. Ovarian cycling and reproductive state shape the vaginal microbiota in wild baboons, Microbiome, 5, 8.
Moreno, I., Codoñer, F.M., Vilella, F., Valbuena, D., Martinez-Blanch, J.F., Jimenez-Almazán, J., Alonso, R., Alamá, P., Remohí, J., Pellicer, A., Ramon, D., and Simon, C., 2016. Evidence that the endometrial microbiota has an effect on implantation success or failure, American Journal of Obstetrics and Gynecology, 215, 684-703.
Otero, C., Saavedra, L., Silva de Ruiz, C., Wilde, O., Holgado, A.R., and Nader-Macías, M.E., 2000. Vaginal bacterial microflora modifications during the growth of healthy cows, Letters in Applied Microbiology, 31, 251-254.
Parks, D.H., Tyson, G.W., Hugenholtz, P., and Beiko, R.G., 2014. STAMP: statistical analysis of taxonomic and functional profiles, Bioinformatics, 30, 3123-3124.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F.O., 2013. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools, Nucleic Acids Research, 41, D590-596.
Quereda, J.J., Barba, M., Mocé, M.L., Gomis, J., Jiménez-Trigos, E., García-Muñoz, Á., Gómez-Martín, Á., González-Torres, P., Carbonetto, B., and García-Roselló, E., 2020. Vaginal microbiota changes during estrous cycle in dairy heifers, Frontier in Veterinary Science, 7, 371.
Ravel, J., Gajer, P., Abdo, Z., Schneider, G.M., Koenig, S.S.K., McCulle, S.L., Karlebach, S., Gorle, R., Russell, J., Tacket, C.O., Brotman, R.M., Davis, C.C., Ault, K., Peralta, L., and Forney, L.J., 2011. Vaginal microbiome of reproductive-age women, Proceedings of the National Academy of Sciences, 108, 4680-4687.
Reid, G., Chan, R.C., Bruce, A.W., and Costerton, J.W., 1985. Prevention of urinary tract infection in rats with an indigenous Lactobacillus casei strain, Infection and Immunity, 49, 320-324.
Santos, T.M., and Bicalho, R.C., 2012. Diversity and succession of bacterial communities in the uterine fluid of postpartum metritic, endometritic and healthy dairy cows, PLOS ONE, 7, e53048.
Swartz, J.D., Lachman, M., Westveer, K., O’Neill, T., Geary, T., Kott, R.W., Berardinelli, J.G., Hatfield, P.G., Thomson, J.M., Roberts, A., and Yeoman, C.J., 2014. Characterization of the vaginal microbiota of ewes and cows reveals a unique microbiota with low levels of lactobacilli and near-neutral pH, Frontier in Veterinary Science, 1, 19.
Wang, Y., Wang, J., Li, H., Fu, K., Pang, B., Yang, Y., Liu, Y., Tian, W., and Cao, R., 2018. Characterization of the cervical bacterial community in dairy cows with metritis and during different physiological phases, Theriogenology, 108, 306-313.
White, D.G., Harmon, R.J., Matos, J.E.S., and Langlois, B.E., 1989. Isolation and identification of coagulase-negative staphylococcus species from bovine body sites and streak canals of nulliparous heifers, Journal of Dairy Science, 72, 1886-1892.
Willis, A.D., 2019. Rarefaction, alpha Diversity, and statistics, Frontiers in Microbiology, 10, 2407.
Acknowledgements
The authors thank Gujarat State Biotechnology Mission, Gandhinagar, Gujarat, India, for providing funding support and Kamdhenu University for providing technical support and logistics for conducting the experiment. Furthermore, we also extend our deep gratitude to the farmers for allowing us to their farms to conduct this experiment.
Author information
Authors and Affiliations
Contributions
The experimental scheme was designed by VS, MJ, and CJ. VS and KP participated in the experiment process and assisted in sampling. The analysis of experimental data and the making of charts were completed by PG, KP, Nitin, and AP who also completed the initial draft; PG rewrote the draft. VS and MJ completed the overall modification of the manuscript. CJ and DP improved and polished the language of the article. DP, MJ, and CJ provided the necessary experimental equipment and key guidance during the experiment process. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
All experimental designs and protocols were approved by the Animal Ethics Committee of the Kamdhenu University and were under the recommendations of the academy’s guidelines for animal research.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gohil, P., Patel, K., Purohit, K. et al. Metagenomic analysis of bacterial communities during estrous cycle in Bos indicus. Trop Anim Health Prod 54, 118 (2022). https://doi.org/10.1007/s11250-022-03119-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11250-022-03119-5