Abstract
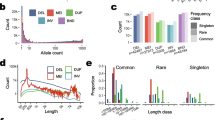
In this study, the genetic structure of indigenous Tswana and Swazi goats using the Illumina Goat50K SNP array was investigated. Two South African commercial goat breeds were included to investigate admixture with the indigenous populations in southern Africa. A total of 144 DNA samples including Boer goats (n = 24), Kalahari Red (n = 24), Swazi (n = 48), and Tswana goats (n = 48) were genotyped. Statistical analysis was performed using PLINK version 1.07. Genetic diversity, measured as expected heterozygosity, was estimated at 0.390, 0.398, 0.413, and 0.387 for Boer, Kalahari Red, Tswana, and Swazi goats, respectively. The individual inbreeding coefficient varied from 0.019 ± 0.05 to 0.011 ± 0.06 for the Tswana and Swazi goats, respectively. The Principal component analysis clustered the populations according to geographical origin and breed type. Linkage disequilibrium (LD) for shorter intervals (0–10 kb) ranged from 0.44 to 0.56 and commercial breeds had higher values. Effective population sizes decreased with generations and at the 13th generation ranged between 87 for Boer to 266 for Tswana goats. The Tswana population exhibited the highest level of genetic variation and effective population size, which holds potential for improved production in marginal regions. A national strategy is required to maintain genetic diversity in communal goat production systems through well-structured breeding and conservation programs.




Similar content being viewed by others
References
Alexander, D.H., Novembre, J., Lange, K., 2009. Fast model-based estimation of ancestry in unrelated individuals, Genome Research, 19, 1655–1664.
Barbato, M., Orozco-terWengel, P., Tapio, M., Bruford, M.W., 2015. SNeP: A tool to estimate trends in recent effective population size trajectories using genome-wide SNP data, Frontiers in Genetics, 6, 109.
Brito, L.F., McEwan, J.C., Miller, S.P., Pickering, N.K., Bain, W.E., Dodds, K.G., Schenkel, F.S., Clarke, S.M., 2017. Genetic diversity of a New Zealand multi-breed sheep population and composite breeds’ history revealed by a high-density SNP chip. BMC Genetics, 18(1), 25.
Corbin, L. J., Liu A., Bishop S., Woolliams, J., 2012. Estimation of historical effective population size using linkage disequilibria with marker data, Journal of Animal Breeding and Genetics, 129(4), 257–270.
Gama, L.T., Bressan, M.C., 2011. Biotechnology applications for the sustainable management of goat genetic resources, Small Ruminant Research, 98(1), 133–146.
Garrine, C.M.L.P., 2007. Genetic characterization of indigenous goat populations of Mozambique (Doctoral dissertation, University of Pretoria).
Goddard, M.E., Hayes, B.J., 2009. Mapping genes for complex traits in domestic animals and their use in breeding programmes, Nature Reviews Genetics, 10(6), 381–391.
Groeneveld, L.F., Lenstra, J.A., Eding, H., Toro, M.A., Scherf, B., Pilling, D., Negrini, R., Finlay, E. K., Jianlin, H., Groeneveld, E., Weigend, S., The GLOBALDIV Consortium (2010), Genetic diversity in farm animals – a review, Animal Genetics, 41, 6–31.
Gwaze, F.R., Chimonyo, M., Dzama, K., 2009. Communal goat production in Southern Africa: A review, Tropical Animal Health and Production, 41, 1157–1168.
Hanotte, O., Jianlin, H., 2006. Genetic characterization of livestock populations and its use in conservation decision-making. The Role of Biotechnology in Exploring and Protecting Agricultural Genetic Resources. Food and Agriculture Organization of the United Nations, Rome, 89–96.
Hoffmann, I., 2010. Climate change and the characterization, breeding and conservation of animal genetic resources, Animal Genetics, 41, 32–46.
Kantanen, J., Olsaker, I., Holm, L.E., Lien, S., Vilkki, J., Brusgaard, K., Eythorsdottir, E., Danell, B., Adalsteinsson, S., 2000. Genetic diversity and population structure of 20 North European cattle breeds, Journal of Heredity, 91(6),446–457.
Kim, E.S., Elbeltagy, A.R., Aboul-Naga, A.M., Rischkowsky, B., Sayre, B., Mwacharo, J.M., Rothschild, M.F., 2016. Multiple genomic signatures of selection in goats and sheep indigenous to a hot arid environment, Heredity, 116(3), 255.
Lenstra, J.A., Groeneveld, L.F., Eding, H., Kantanen, J., Williams, J.L., Taberlet, P., Nicolazzi, E.L., Sölkner, J., Simianer, H., Ciani, E., Garcia, J.F., 2012. Molecular tools and analytical approaches for the characterization of farm animal genetic diversity, Animal Genetics, 43(5), 483–502.
Makina, S.O., Muchadeyi, F.C., van Marle-Köster, E., MacNeil, M.D., Maiwashe, A., 2014. Genetic diversity and population structure among six cattle breeds in South Africa using a whole genome SNP panel, Frontiers in Genetics, 5, 333.
Mdladla, K., Dzomba, E., Huson, H., Muchadeyi, F., 2016. Population genomic structure and linkage disequilibrium analysis of South African goat breeds using genome-wide SNP data, Animal Genetics, 47(4), 471–482.
Mekuriaw, G., Mwacharo, J., Tesfaye, K., Tadelle, D., Okeyo, M., Djikeng, A., Liu, B., Osama, S., Grossen, C., Zhang, W., 2016. High Density SNP Chips Array Uncovers Genetic Diversity and Population Structure of 16 Ethiopian and Chinese Goat Populations. Plant and Animal Genome Conference XXIV. In: January 9–13. SanDiego: CA; 2016. p. 2016.
Meuwissen, T., 2009. Genetic management of small populations: a review. Acta Agriculturae Scandinavica A, 59(2), 71–9.
Monau, P.I., Visser, C., Nsoso, S.J., van Marle-Koster, E. 2018. Phenotypic and genetic characterization of indigenous Tswana goats, South African Journal of Animal Science, 48, 925-934.
Mueller, J.P., Rischkowsky, B., Haile, A., Philipsson, J., Mwai, O., Besbes, B., Valle Zárate, A., Tibbo, M., Mirkena, T., Duguma, G., Sölkner, J., 2015. Community-based livestock breeding programmes: essentials and examples. Journal of Animal Breeding and Genetics, 132(2), 155–168.
Muema, E.K., Wakhungu, J.W., Hanotte, O., Jianlin, H., 2009. Genetic diversity and relationship of indigenous goats of Sub-saharan Africa using microsatellite DNA markers. Livestock Research for Rural Development, 21(2), 28.
Mwai, O., Hanotte, O., Kwon, Y-J., Cho, S., 2015. African indigenous cattle: unique genetic resources in a rapidly changing world. Asian-Australasian Journal of Animal Sciences, 28,911.
Onzima, R.B., Upadhyay, M.R., Mukiibi, R., Kanis, E., Groenen, M.A.M., Crooijmans, R.P.M.A., 2018. Genome-wide population structure and admixture analysis reveals weak differentiation among Ugandan goat breeds, Animal Genetics, 49 (1), 59–70.
Purcell, S., Neale, B., Todd-Brown, K., Thomas L., Ferreira, M.A., Bender, D., Maller, J., Sklar, P., De Bakker, P.I., Daly, M.J., 2007. PLINK: A tool set for whole-genome association and population-based linkage analyses, American Journal of Human Genetics, 81(3), 559–575.
Rahmatalla, S.A., Arends, D., Reissmann, M., Ahmed, A.S., Wimmers, K., Reyer, H. and Brockmann, G.A., 2017. Whole genome population genetics analysis of Sudanese goats identifies regions harbouring genes associated with major traits, BMC Genetics, 18(1), 92.
Rege, J.E.O., Marshall, K., Notenbaert, A., Ojango, J.M.K., Okeyo, A.M., 2011. Pro-poor animal improvement and breeding—What can science do? Livestock Science, 136(1), 15–28.
Ribeiro, M.N., Bruno-de-Sousa, C., Martinez-Martinez, A., Ginja, C., Menezes, M.P.C., Pimenta-Filho, E.C., Delgado, J.V., Gama, L.T., 2012. Drift across the Atlantic: genetic differentiation and population structure in Brazilian and Portuguese native goat breeds, Journal of Animal Breeding and Genetics, 129, 79–87.
SADC, 2013. Selected indicators for the SADC region for 2011 with charts. Accessed from http://www.sadc.int/files/6213/6267/6607/Selected_Indicators_2011_with_charts06March2013_FINAL.pdf (on 08 July 2018).
Smith, A.B., 1992. Origins and spread of pastoralism in Africa, Annual Review of Anthropology, 21, 125–141.
Statistical Analysis Systems. 2009. SAS 9.3 Qualification Tools User’s Guide. SAS Institute Inc: Cary N.C.
Tada, O., Muchenje, V., Dzama, K., 2013. Effective population size and inbreeding rate of indigenous Nguni cattle under in situ conservation in the low-input communal production system, South African Journal of Animal Science, 43(2), 137–142.
Tosser-Klopp, G., Bardou, P., Bouchez, O., Cabau, C., Crooijmans, R., Dong, Y., Donnadieu-Tonon, C., Eggen, A., Heuven, H.C., Jamli, S., Jiken, A.J., 2014. Design and characterization of a 52K SNP chip for goats, PLoSONE. 9(1), p.e86227.
Visser, C., Lashmar, S. F., Van Marle-Köster, E., Poli, M.A., Allain, D., 2016. Genetic diversity and population structure in South African, French and Argentinean Angora goats from genome-wide SNP data, PLoSONE. 11(5), p.e0154353.
Yang, J., Lee, S.H., Goddard, M.E., Visscher, P.M., 2011. GCTA: a tool for genome-wide complex trait analysis, The American Journal of Human Genetics, 88(1),76–82.
Zvinorova, P.I., 2017. A genome-wide association study on mechanisms underlying genetic resistance to gastrointestinal parasites in goats, Zimbabwe (Doctoral dissertation, Stellenbosch: Stellenbosch University).
Acknowledgments
The authors would like to acknowledge the Organization of Women in Science and Botswana University of Agriculture and Natural Science for financial support for the student. Sincere gratitude to the goat farmers who participated in this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Monau, P.I., Visser, C., Muchadeyi, F.C. et al. Population structure of indigenous southern African goats based on the Illumina Goat50K SNP panel. Trop Anim Health Prod 52, 1795–1802 (2020). https://doi.org/10.1007/s11250-019-02190-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-019-02190-9