Abstract
Two tomato proteins were evaluated by over-expression in transgenic tomato for their ability to confer resistance to Clavibacter michiganensis subsp. michiganensis (Cmm). Snakin-2 (SN2) is a cysteine-rich peptide with broad-spectrum antimicrobial activity in vitro while extensin-like protein (ELP) is a major cell-wall hydroxyproline-rich glycoprotein linked with plant response to pathogen attack and wounding. Tomato plants, cultivar Mountain Fresh, were transformed via Agrobacterium tumefaciens harboring a binary vector for expression of the full-length SN2 gene or ELP cDNA under the regulation of the CaMV 35S promoter. Molecular characterization of PCR-positive putative T0 transgenic plants by Northern analysis revealed constitutive over-expression of SN2 and ELP mRNA. Junction fragment analysis by Southern blot showed that three of the four SN2 over-expressing T0 lines had single copies of complete T-DNAs while the other line had two complete T-DNA copies. All four ELP over-expressing T0 lines had a single copy T-DNA insertion. Semi-quantitative RT-PCR analysis of T1 plants revealed constitutive over-expression of SN2 and ELP. Transgenic lines that accumulated high levels of SN2 or ELP mRNA showed enhanced tolerance to Cmm resulting in a significant delay in the development of wilt symptoms and a reduction in the size of canker lesions compared to non-transformed control plants. Furthermore, in transgenic lines over-expressing SN2 or ELP bacterial populations were significantly lower (100–10,000-fold) than in non-transformed control plants. These results demonstrate that SN2 and ELP over-expression limits Cmm invasiveness suggesting potential in vivo antibacterial activity and possible biotechnological application for these two defense proteins.





Similar content being viewed by others
References
Almasia NI, Bazzini AA, Hopp HE, Vazquez-Rovere C (2008) Overexpression of snakin-1 gene enhances resistance to Rhizoctonia solani and Erwinia carotovora in transgenic potato plants. Mol Plant Pathol 9:329–338
Balaji V, Mayrose M, Sherf O, Jacob-Hirsch J, Eichenlaub R, Iraki N, Manulis-Sasson S, Rechavi G, Barash I, Sessa G (2008) Tomato transcriptional changes in response to Clavibacter michiganensis subsp. michiganensis reveal a role for ethylene in disease development. Plant Physiol 146:1797–1809
Balaji V, Sessa G, Smart CD (2011) Silencing of host basal defense response-related gene expression increases susceptibility of Nicotiana benthamiana to Clavibacter michiganensis subsp. michiganensis. Phytopathology 101:349–357
Beimen A, Bermpohl A, Meletzus D, Eichenlaub R, Barz W (1992) Accumulation of phenolic compounds in leaves of tomato plants after infection with Clavibacter michiganense subsp. michiganense strains differing in virulence. Z Naturforsch 47C:898–909
Ben-Nissan G, Weiss D (1996) The petunia homologue of tomato gast1: transcript accumulation coincides with gibberellin-induced corolla cell elongation. Plant Mol Biol 32:1067–1074
Berrocal-Lobo M, Segura A, Moreno M, Lopez G, Garcia-Olmedo F, Molina A (2002) Snakin-2, an antimicrobial peptide from potato whose gene is locally induced by wounding and responds to pathogen infection. Plant Physiol 128:951–961
Bindschedler LV, Whitelegge JP, Millar DJ, Bolwell GP (2006) A two component chitin-binding protein from French bean-association of a proline-rich protein with a cysteine-rich polypeptide. FEBS Lett 580:1541–1546
Bowles DJ (1990) Defence-related proteins in higher plants. Annu Rev Biochem 59:873–907
Cassab GI (1998) Plant cell wall proteins. Annu Rev Plant Physiol 49:281–309
Chalupowicz L, Cohen-Kandli M, Dror O, Eichenlaub R, Gartemann K-H, Sessa G, Barash I, Manulis-Sasson S (2010) Sequential expression of bacterial virulence and plant defense genes during infection of tomato with Clavibacter michiganensis subsp. michiganensis. Phytopathology 100:252–261
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Chee PP, Fober KA, Slightom JL (1989) Transformation of soybean (Glycine max) by infecting germinating seeds with Agrobacterium tumefaciens. Plant Physiol 91:1212–1218
Chilton M-D, Currier TC, Farrand SK, Bendich AJ, Gordon MP, Nester EW (1974) Agrobacterium tumefaciens DNA and PS8 bacteriophage DNA not detected in crown gall tumors. Proc Natl Acad Sci USA 71:3672–3676
Chun WCC (1982) Identification and detection of Corynebacterium michiganense in tomato seed using the enzyme-linked immunosorbent assay. MS thesis, University of Hawaii, Honolulu
Davies HA, Daniels MJ, Dow JM (1997) Induction of extracellular matrix glycoproteins in Brassica petioles by wounding and in response to Xanthomonas campestris. Mol Plant Microbe Interact 10:812–820
Davis MJ, Gillespie JA, Vidaver AK, Harris RW (1984) Clavibacter: a new genus containing some phytopathogenic coryneform bacteria, including Clavibacter xyli subsp. xyli sp. nov., subsp. nov. and Clavibacter xyli subsp. cynodontis subsp. nov., pathogens that cause ratoon stunting disease of sugarcane and bermudagrass stunting disease. Int J Syst Bacteriol 34:107–117
Epstein L, Lamport DTA (1984) An intramolecular linkage involving isodityrosine in extensin. Phytochemistry 23:1241–1246
Fry SC (1988) The growing plant cell wall: chemical and metabolic analysis. The Blackburn Press, Caldwell
Furukawa T, Sakaguchi N, Shimada H (2006) Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet Syst 81:171–180
Gartemann KH, Kirchner O, Engemann J, Grafen I, Eichenlaub R, Burger A (2003) Clavibacter michiganensis subsp. michiganensis: first step in understanding of virulence of a Gram-positive phytopathogenic bacterium. J Biotechnol 106:179–191
Gartemann K-H, Abt B, Bekel T, Burger A, Engemann J, Flügel M, Gaigalat L, Goesmann A, Grafen I, Kalinowski J, Kaup O, Kirchner O, Krause L, Linke B, McHardy A, Meyer F, Pohle S, Ruckert C, Schneiker S, Zellermann E-M, Puhler A, Eichenlaub R, Kaiser O, Bartels D (2008) The genome sequence of the tomato-pathogenic actinomycete Clavibacter michiganensis subsp. michiganensis NCPPB382 reveals a large island involved in pathogenicity. J Bacteriol 190:2138–2149
Hauck P, Thilmony R, He SY (2003) A Pseudomonas syringae type 111 effector suppresses cell wall-based extracellular defense in susceptible Arabidopsis plants. Proc Natl Acad Sci USA 100:8577–8582
Hoekema A, Hirsch PR, Hooykaas PJJ, Schilperoort RA (1983) A binary plant vector strategy based on separation of vir- and T-region of the Agrobacterium tumefaciens Ti-plasmid. Nature 303:179–180
Jahr H, Bahro R, Burger A, Ahlemeyer J, Eichenlaub R (1999) Interactions between Clavibacter michiganensis and its host plants. Environ Microbiol 1:113–118
Jahr H, Dreier J, Meletzus D, Bahro R, Eichenlaub R (2000) The endo-b-1, 4-glucanase CelA of Clavibacter michiganensis subsp. michiganensis is a pathogenicity determinant required for induction of bacterial wilt of tomato. Mol Plant Microbe Interact 13:703–714
Jakobek JL, Smith JA, Lindgren PB (1993) Suppression of bean defense responses by Pseudomonas syringae. Plant Cell 5:57–63
Jones JB, Jones JP, Stall RE, Zitter TA (1991) Compendium of tomato diseases. APS Press, St. Paul
Kotilainen M, Helariutta Y, Mehto M, Pollanen E, Albert VA, Elomaa P, Teeri TH (1999) GEG participates in the regulation of cell and organ shape during corolla and carpel development in gerbera hybrida. Plant Cell 11:1093–1104
Kovalskaya N, Hammond RW (2009) Expression and functional characterization of the plant antimicrobial snakin-1 and defensin recombinant proteins. Protein Expr Purif 63:12–17
Ling KS, Zhu HY, Alvizo H, Hu JS, Drong RF, Gonsalves D (1997) The coat protein gene of grapevine leafroll associated closterovirus-3: cloning, nucleotide sequencing and expression in transgenic plants. Arch Virol 142:1101–1116
Lopez-Solanilla E, Gonzalez-Zorn B, Novella S, Vazquez-Boland JA, Rodriguez-Palenzuela P (2003) Susceptibility of Listeria monocytogenes to antimicrobial peptides. FEMS Microbiol Lett 226:101–105
Mazau D, Rumeau D, Esquerré-Tugayé MT (1987) Molecular approaches to understanding cell–surface interactions between plants and fungal pathogens. Plant Physiol Biochem 25:337–343
Meletzus D, Bermpohl A, Drier J, Eichenlaub R (1993) Evidence for plasmid-encoded virulence factors in the phytopathogenic bacterium Clavibacter michiganensis subsp. michiganensis NCPPB382. J Bacteriol 175:2131–2136
Mellon J, Helgeson J (1982) Interaction of a hydroxyproline-rich glycoprotein from tobacco callus with potential pathogens. Plant Physiol 70:401–405
Merkouropoulos G, Shirsat AH (2003) The unusual Arabidopsis extensin gene EXT1 is expressed throughout plant development and is induced by a variety of biotic and abiotic stresses. Planta 217:356–366
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Rogers SO, Bendich AJ (1988) Extraction of total cellular DNA from plants. In: Gelvin SB, Schilperoort RA, Verma DPS (eds) Plant molecular biology manual. Kluwer Academic Publishers, Netherlands, pp A6/1–A6/10
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Segura A, Moreno M, Madueno F, Molina A, Garcia-Olmedo F (1999) Snakin-1, a peptide from potato that is active against plant pathogens. Mol Plant Microbe Interact 12:16–23
Selitrennikoff CP (2001) Antifungal proteins. Appl Environ Microbiol 67:2883–2894
Shi L, Gast RT, Gopalraj M, Olszewski NE (1992) Characterization of a shoot-specific, GA3 and ABA-regulated gene from tomato. Plant J 2:153–159
Showalter AM (1993) Structure and function of plant cell wall proteins. Plant Cell 5:9–23
Showalter A, Rumeau D (1990) Molecular biology of the plant cell wall hydroxyproline-rich glycoproteins. In: Adair W, Mercham R (eds) Organisation and assembly of plant and animal extracellular matrix. Academic Press, New York, pp 247–281
Showalter AM, Varner JE (1989) Plant hydroxyproline-rich glycoproteins. In: Stumpf PK, Conn EE (eds) The biochemistry of plants: a comprehensive treatise. Academic Press, New York, pp 485–520
Sommer-Knudsen J, Bacic A, Clarke AE (1998) Hydroxyproline-rich plant glycoproteins. Phytochemitsry 47:483–497
Strider DL (1969) Bacterial canker of tomato caused by Corynebacterium michiganense. A literature review and bibliography. North Carolina agricultural experiment station technical bulletin, no. 193
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Thornburg RW, Carter C, Powell A, Mittler R, Rizhsky L, Horner HT (2003) A major function of the tobacco floral nectary is defense against microbial attack. Plant Syst Evol 238:211–218
Van den Heuvel KJ, Lipzig RHV, Barendse GW, Wullems GJ (2002) Regulation of expression of two novel flower-specific genes from tomato (Solanum lycopersicum) by gibberellin. J Exp Bot 53:51–59
Van Eck J, Kirk DD, Walmsley AM (2006) Tomato (Lycopersicum esculentum). In: Wang K (ed) Molecular biology: Agrobacterium protocols. Humana Press, Totowa, pp 459–473
Vance CP, Kirk TK, Sherwood RT (1980) Lignification as a mechanism of disease resistance. Annu Rev Phytopathol 18:259–288
Wei G, Shirsat AH (2006) Extensin over-expression in Arabidopsis limits pathogen invasiveness. Mol Plant Pathol 7:579–592
Wigoda N, Ben-Nissan G, Granot D, Schwartz A, Weiss D (2006) The gibberellin-induced, cysteine-rich protein GIP2 from Petunia hybrida exhibits in planta antioxidant activity. Plant J 48:796–805
Wu H-M, Zou J, May B, Gu Q, Cheung AY (1993) A tobacco gene family for flower cell wall proteins with a proline-rich domain and a cysteine-rich domain. Proc Natl Acad Sci USA 90:6829–6833
Acknowledgments
We thank Holly Lange and Kevin Conley for maintaining plants in the greenhouse. This research was supported by the New York State Agricultural Experiment Station and the United States-Israel Binational Agricultural Research and Development Fund (award No. IS-4047-07).
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence of the tomato snakin-2 gene reported in this paper is deposited in GenBank under the accession number HQ008860.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Kanamycin sensitivity assay to determine the lowest effective concentration that can inhibit callusing and shoot bud formation in cotyledon explants of tomato. A, Cotyledon explants were excised from aseptically grown 8-day-old seedlings and placed on 2Z medium (Van Eck et al. 2006) supplemented with 25, 50, 75, 100 and 125 mg/l kanamycin. Cotyledon explants placed on 2Z medium without kanamycin were included as a positive control. All treatments contained timentin at a concentration of 300 mg/l and 60 explants were analyzed for each treatment. B, Histogram showing the percentage of cotyledon explants (in a group of 60) that callused and formed shoot buds in the presence of increasing concentration of kanamycin. Data was scored after 18 days of culture (JPG 2.65 MB)
Fig. S2
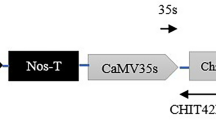
The T-DNA region of binary vectors pGA482GSN2 (A) and pGA482GELP (B) used for tomato transformation. Diagrams are not to scale. The Cauliflower mosaic virus (CaMV) 35S promoter (P35S)-driven tomato snakin-2 (SN2) gene or extensin-like protein (ELP) encoding cDNA was placed between the right border (RB) and left border (LB) of the T-DNA. The binary vector contains the neomycin phosphotransferase (nptII) gene as the plant selection marker. Pnos, nopaline synthase promoter; Tnos, nopaline synthase terminator; T35S, CaMV 35S polyadenylation signal. Solid arrows below the nptII gene indicate annealing sites of the primer pair used in the polymerase chain reaction. Bold solid lines below the map represent the fragments used as probes in Southern blot analysis. The dotted lines indicate the sizes of internal T-DNA fragments generated by HindIII digestion. The XhoI and BamHI restriction sites used for Southern blot analysis to determine the LB T-DNA junction fragments are marked with an asterisk. The dotted arrows indicate the sizes of junction fragments generated by XhoI (A) or BamHI (B) digestion (JPG 469 KB)
Fig. S3
Screening putative transgenic plants by PCR. The PCR reaction was performed with primer sets specific for the nptII gene (see Table 1). Genomic DNA from putative kanamycin-resistant plants transformed with snakin-2 (SN2) gene (A) or extensin-like protein (ELP) cDNA (B) and non-transgenic plants (NT) were used as template. The 1 kb ladder was used as the size marker (M). S and E denote plants transformed with the SN2 gene or ELP cDNA, respectively. The size of the amplicon was 625 bp (JPG 516 KB)
Fig. S4
Scoring for kanamycin resistance in T1 seeds of transgenic tomato lines over-expressing SN2 and ELP genes. A, Two replicates of a kanamycin sensitivity assay to determine the effective lethal dose that would arrest rooting and growth of tomato seedlings. Seeds of non-transformed tomato plants were surface-sterilized and placed on half-strength MS medium containing 0, 50, 75 or 100 mg/l kanamycin. A total of 30 seedlings were assayed for each treatment. Photographs were taken 10 days after culture. B, Scoring of kanamycin-resistant T1 seedlings in the segregation population of SN2 over-expressing T0 tomato lines S6, S9 and S14. C, Screening for kanamycin resistance in the T1 progeny of ELP over-expressing lines E12, E22 and E24. Kanamycin at a concentration of 100 mg/l was used in the selective screen. Seeds from non-transformed (NT) plant were placed on half-strength MS medium with and without kanamycin and treated as negative and positive controls, respectively (panels B and C, left). The kanamycin based scoring assay was validated by PCR (panels B and C, bottom) with primers specific for the nptII gene (Table S1). Genomic DNA template isolated from representative T1 seedlings sensitive (S) and resistant (R) to kanamycin in the segregating population of SN2 over-expressing T0 lines S6, S9 and S14 (panel B, bottom) and ELP lines E12, E22 and E24 (panel C, bottom). Lane NT indicates non-transformed control. A 1 kb ladder was used the DNA size marker (M) (JPG 3.78 MB)
Rights and permissions
About this article
Cite this article
Balaji, V., Smart, C.D. Over-expression of snakin-2 and extensin-like protein genes restricts pathogen invasiveness and enhances tolerance to Clavibacter michiganensis subsp. michiganensis in transgenic tomato (Solanum lycopersicum). Transgenic Res 21, 23–37 (2012). https://doi.org/10.1007/s11248-011-9506-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-011-9506-x