Abstract
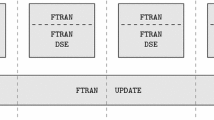
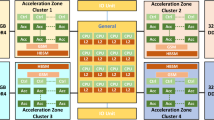
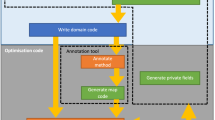
A longest common subsequence (LCS) of two strings is a common subsequence of two strings of maximal length. The LCS problem is to find an LCS of two given strings and the length of the LCS (LLCS). In this paper, we present a new linear processor array for solving the LCS problem. The array is based on parallelization of a recent LCS algorithm which consists of two phases, i.e. preprocessing and computation. The computation phase is based on bit-level dynamic programming approach. Implementations of the preprocessing and computation phases are discussed on the same processor array architecture for the LCS problem. Further, we propose a block processor array architecture which reduces the overall communication and time requirements. Finally, we develop a performance model for estimating the performance of the processor array architecture on Pentium processors.
Similar content being viewed by others
References
A. V. Aho, D. S. Hirschberg, and J. D. Ullman Bounds on the complexity of the longest common subsequence problem. Journal of Association Computing Machineny, 23:1–12, 1976.
A. Apostolico, M. Attalah, L. Larmore, and S. Mcfaddin Efficient parallel algorithms for string editing and related problems. SIAM Journal on Computing, 19:968–988, 1990.
L. Bergroth, H. Hakonen, and T. Raita A survey of longest common subsequence algorithms. In 7th International Symposium on String Processing and Information Retrieval, pp. 39–48, 2000.
M. Crochemore, C. S. Iliopoulos, Y. J. Pinzon, and J. R. Reid A fastand practical bit-vector algorithm for the longest common subsequence problem. In 11th Australasian Workshop on Combinatorial Algorithms, pp. 75–86, 2000.
M. Crochemore, C. S. Iliopoulos, Y. J. Pinzon, and J. R. Reid A fast and practical bit-vector algorithm for the longest common subsequence problem. Information Processing Letters, 80:279–285, 2001.
D. Gusfield Algorithms on Strings, Trees and Sequences: Computer Science and Computational Biology. Cambridge University Press, Gambridge, England, 1997.
D. S. Hirschberg A linear space algorithm for computing maximal common subsequences. Communications of the ACM, 18:341–343, 1975.
D. S. Hirschberg An information theoretic lower bound for the longest common subsequence problem. Information Processing Letters, 7:40–41, 1978.
R. Hughey Parallel hardware for sequence comparison and alignment. CABIOS, 12:473–479, 1996.
S. Y. Kung VLSI Array Processors. Prentice-Hall, 1988.
T. Lecroq, G. Luce, and J. F. Myoupo A faster linear systolic algorithm for recovering a longest common subsequence. Information Processing Letters, 61:129–136, 1997.
T. Lecroq, J. F. Myoupo, and D. Seme A one-phase parallel algorithm for the sequence alignment problem. Parallel Processing Letters, 8:515–526, 1998.
Y.-C. Lin New systolic arrays for the longest common subsequence problem. Parallel Computing, 20:1323–1334, 1994.
Y.-C. Lin and J.-C. Chen An efficient systolic algorithm for the longest common subsequence problem. The Journal of Supercomputing, 12:373–385, 1998.
Y.-C. Lin and J.-C. Chen Another efficient systolic algorithm for the longest common subsequence problem. Journal of the Chinese Institute of Engineers, 23:607–613, 2000.
Y.-C. Lin and J.-W. Yeh A scalable and efficient systolic algorithm for the longest common subsequence problem. Journal of Information Science and Engineering, 18:519–532, 2002.
Y.-C. Lin and J.-W. Yeh Deriving a systolic algorithm for the LCS problem. International Conference on Parallel and Distributed Processing Techniques and Applications, pp. 1890–1897, 1998.
Y.-C. Lin and J.-W. Yeh Deriving a fast systolic algorithm for the longest common subsequence problem. Parallel Algorithms and Applications, 17:1–18, 2002.
G. Luce and J. F. Myoupo Systolic-based parallel architecture for the longest common subsequences problem. Integration, the VLSI Journal, 25:53–70, 1998.
M. Lu and H. Lin Parallel algorithms for the longest common subsequence problem. IEEE Transactions on Parallel and Distributed Systems, 5:835–848, 1994.
K. G. Margaritis and D. J. Evans A VLSI processor array for flexible string matching. Parallel Algorithms and Applications, 11:45–60, 1997.
W. J. Masek and M. S. Paterson A faster algorithm computing string edit distances. Journal of Computer System Sciences, 20:18–31, 1980.
J. H. Moreno and T. Lang Matrix Computations on Systolic-Type Arrays. Kluwer Academic Publishers, 1992.
N. Petkov Systolic Parallel Processing. Elsevier Science Publishers, 1993.
N. Ranganathan and R. Sastry VLSI architectures for pattern matching. International Journal of Pattern Recognition and Artificial Intelligence, 8:815–843, 1994.
Y. Robert and M. Tchuente A systolic array for the longest common subsequence problem. Information Processing Letters, 21:191–198, 1985.
D. Sankoff and J.B. Kruskal Time Warps, String Edits and Macromolecules: The Theory and Practice of Sequence Comparison. Addison-Wesley, 1983.
J. Setubal and J. Meidanis Introduction to Computational Molecular Biology. PWS, Boston, MA, 1997.
M. Snir, S. Otto, S. Huss-Lederman, D. W. Walker and J. Dongarra MPI: The Complete Reference. The MIT Press, Cambridge, Massachusetts, 1996.
C.-B. Yang and R. C. T. Lee Systolic algorithms for the longest common subsequence problem. Journal of the Chinese Institute of Engineers, 10:691–699, 1987.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Michailidis, P.D., Margaritis, K.G. New Processor Array Architectures for the Longest Common Subsequence Problem. J Supercomput 32, 51–69 (2005). https://doi.org/10.1007/s11227-005-0158-6
Issue Date:
DOI: https://doi.org/10.1007/s11227-005-0158-6