Abstract
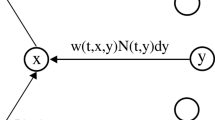
Reconstructing the functional network of a neuron cluster is a fundamental step to reveal the complex interactions among neural systems of the brain. Current approaches to reconstruct a network of neurons or neural systems focus on establishing a static network by assuming the neural network structure does not change over time. To the best of our knowledge, this is the first attempt to build a time-varying directed network of neurons by using an ordinary differential equation model, which allows us to describe the underlying dynamical mechanism of network connections. The proposed method is demonstrated by estimating a network of wide dynamic range neurons located in the dorsal horn of the rats’ spinal cord in response to pain stimuli applied to the Zusanli acupoint on the right leg. The finite sample performance of the proposed method is also investigated with a simulation study.





Similar content being viewed by others
References
Bullmore, E., Sporns, O.: Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009)
Cao, J.: Estimating generalized semiparametric additive models using parameter cascading. Stat. Comput. 22, 857–865 (2012)
Cao, J., Ramsay, J.O.: Linear mixed-effects modeling by parameter cascading. J. Am. Stat. Assoc. 105(489), 365–374 (2010)
Daley, D., Vere-Jones, D.: An Introduction to the Theory of Point Processes. Springer, New York (2003)
de Boor, C.: A Practical Guide to Splines. Springer, New York (2001)
de Haan, W., Pijnenburg, Y.A., Strijers, R.L., van der Made, Y., van der Flier, W.M., Scheltens, P., Stam, C.J.: Functional neural network analysis in frontotemporal dementia and Alzheimers disease using EEG and graph theory. BMC Neurosci. 10, 101 (2009)
de Haan, W., van der Flier, W., Wang, H., Van Mieghem, P.F., Scheltens, P., Stam, C.J.: Disruption of functional brain networks in Alzheimer’s disease: what can we learn from graph spectral analysis of resting-state magnetoencephalography? Brain Connect. 2, 45–55 (2012)
Efron, B., Tibshirani, R.J.: An Introduction to the Bootstrap. Chapman & Hall, New York (1993)
Fan, J., Li, R.: Variable selection via nonconcave penalized likelihood and its oracle properties. J. Am. Stat. Assoc. 96(456), 1348–1360 (2001)
Gerhard, F., Pipa, G., Lima, B., Neuenschwander, S., Gerstner, W.: Extraction of network topology from multi-electrode recordings: is there a small-world effect? Front. Comput. Neurosci. 5, 4 (2011)
Horvath, S.: Weighted Network Analysis: Applications in Genomics and Systems Biology. Springer, New York (2014)
Kass, R.E., Eden, U., Brown, E.: Analysis of Neural Data. Springer, New York (2014)
Lewis, P.A.W., Shedler, G.S.: Simulation of nonhomogeneous poisson processes by thinning. Nav. Res. Logist. Q. 26, 403–413 (1979)
Lin, Z., Cao, J., Wang, L., Wang, H.: Locally sparse estimator for functional linear regression models. J. Comput. Graph. Stat. 26, 306–318 (2017)
Luo, W., Cao, J., Gallagher, M., Wiles, J.: Estimating the intensity of ward admission and its effect on emergency department access block. Stat. Med. 32(15), 2681–2694 (2013)
Nie, Y., Wang, L., Cao, J.: Estimating time-varying directed gene regulation networks. Biometrics 73, 1231–1242 (2017)
Paninski, L.: Maximum likelihood estimation of cascade point-process neural encoding models. Netw. Comput. Neural Syst. 15(4), 243–262 (2004)
Picard, R., Cook, D.: Cross-validation of regression models. J. Am. Stat. Assoc. 79, 575–583 (1984)
Qin, Qing, Wang, J., Xue, M., Deng, B., Wei, X.: Charactering neural spiking activity evoked by acupuncture through state-space model. Appl. Math. Model. 39, 1400–1408 (2015)
Siggiridou, E., Kugiumtzis, D., Kimishkidis, V.K.: Correlation networks for identifying changes in brain connectivity during epileptiform discharges and transcranial magnetic stimulation. Sensors 14, 12585–12597 (2014)
Truccolo, W., Eden, U., Fellows, M.R., Donoghue, J.P., Brown, E.N.: A point process framework for relating neural spiking activity to spiking history, neural ensemble, and extrinsic covariate effects. J. Neurophysiol. 93, 1074–1089 (2005)
Wahba, G.: Spline Models for Observational Data. Society for Industrial and Applied Mathematics, Philadelphia (1990)
Wang, L., Cao, J., Ramsay, J.O., Burger, D., Laporte, C., Rockstrohk, J.: Estimating mixed-effects differential equation models. Stat. Comput. 24, 111–121 (2014)
Wood, S.N.: Generalized Additive Models: An Introduction with R, 2nd edn. Chapman and Hall/CRC, London (2017)
Xiao, L., Zhang, Y., Dai, J., Li, J., Li, W.: New Noise–Tolerant ZNN models with predefined-time convergence for time-variant Sylvester equation solving. IEEE Trans. Syst. Man Cybern. Syst. (2019). https://doi.org/10.1109/TSMC.2019.2930646
Zhang, J., Cao, J.: Finding common modules in a time-varying network with application to the drosophila melanogaster gene regulation network. J. Am. Stat. Assoc. 112(519), 994–1008 (2017)
Acknowledgements
The authors are very grateful for the constructive comments from the Editor, the Associate Editor, and two reviewers. The authors would also like to thank Prof. Jiang Wang from Tianjin University for providing us the data. This research was supported by Cao’s Discovery Grant (RGPIN-2018-06008) from the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, H., Cao, J. Estimating time-varying directed neural networks. Stat Comput 30, 1209–1220 (2020). https://doi.org/10.1007/s11222-020-09941-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11222-020-09941-x