Abstract
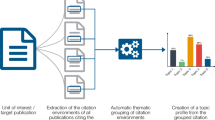
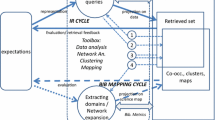
In advanced methods of delineation and mapping of scientific fields, hybrid methods open a promising path to the capitalisation of advantages of approaches based on words and citations. One way to validate the hybrid approaches is to work in cooperation with experts of the fields under scrutiny. We report here an experiment in the field of genomics, where a corpus of documents has been built by a hybrid citation-lexical method, and then clustered into research themes. Experts of the field were associated in the various stages of the process: lexical queries for building the initial set of documents, the seed; citation-based extension aiming at reducing silence; final clustering to identify noise and allow discussion on border areas. The analysis of experts’ advices show a high level of validation of the process, which combines a high-precision and low-recall seed, obtained by journal and lexical queries, and a citation-based extension enhancing the recall. This findings on the genomics field suggest that hybrid methods can efficiently retrieve a corpus of relevant literature, even in complex and emerging fields.



Similar content being viewed by others
Notes
Consortium: ADIS (Université Paris-Sud), Lereco (INRA), OST.
Probabilistic index = observed cell population c(ij)/expected cell population where expected cell population = c(i.)c(.j)/c(..).
AKM is implemented in the commercial software Neuronav by Diatopie (S. Aubin, www.diatopie.com). It is enhanced it with a basic but robust and efficient term-extracting sequence.
References
Aksnes, D. W., Olsen, T. B., & Seglen, P. O. (2000). Validation of bibliometric indicators in the field of microbiology: A Norwegian case study. Scientometrics, 49(1), 7–22.
Archambault, É., Gingras,Y., Godin, B. & Vallières F. (1999). Characterization of genomics in Canada—a bibliometric study of scientific articles and research grants 1995–1997. Prepared for Genome Canada by OST. 19 pp.
Bassecoulard, E., Lelu, A., & Zitt, M. (2007). Mapping nanosciences by citation flows: a preliminary analysis. Scientometrics, 70(3), 859–880.
Basu,A., & Lewison, G. (2006). Visualization of a scientific community of Indian origin in the US: A case study of bioinformatics and genomics. International Workshop on Webometrics, Informetrics and Scientometrics & seventh COLLNET meeting, 10–12 May 2006, Nancy.
Brin, S., & Page, L. (1998). The anatomy of a large-scale hypertextual Web search engine. Computer Networks and Isdn Systems, 30(1–7), 107–117.
Debruin, R. E., & Moed, H. F. (1993). Delimitation of scientific subfields using cognitive words from corporate addresses in scientific publications. Scientometrics, 26(1), 65–80.
Garfield, E. (1967). Primordial concepts, citation indexing and historio-bibliography. Journal Library History, 2, 235–249.
International Human Genome Sequencing Consortium (IHGSC). (2004). Finishing the euchromatic sequence of the human genome. Nature, 431(7011), 931–945.
Lelu, A. (1994). Clusters and factors: neural algorithms for a novel representation of huge and highly multidimensional data sets. In E. Diday & Y. Lechevallier (Eds.), New approaches in classification and data analysis (pp. 241–248). Berlin: Springer-Verlag.
Lelu, A., & François, C. (1992). Automatic generation of hypertext links in information retrieval systems. In D. L. al (Ed.), Proceedings of ECHT’92 (Milano) (pp. 112–121). New York: ACM Press.
McKusick, V. A., & Ruddle, F. H. (1987). A new discipline, a new name, a new journal. Genomics, 1(1), 1–2.
Rinia, E. J., Delange, C., & Moed, H. F. (1993). Measuring national output in physics—delimitation problems. Scientometrics, 28(1), 89–110.
van Leeuwen, T. N., van der Wurff, L. J., & van Raan, A. F. J. (2001). The use of combined bibliometric methods in research funding policy. Research Evaluation, 10, 195–201.
Zitt, M., & Bassecoulard, E. (2006). Delineating complex scientific fields by an hybrid lexical-citation method: An application to nanosciences. Information Processing & Management, 42(6), 1513–1531.
Zitt M., Lelu A., & Bassecoulard E. (2008). Hybrid maps of scientific fields (terms and citations): an application to nanosciences. In J. Gorraiz & Schiebel, E. (Eds.), Excellence and emergence. A new challenge for the combination of quantitative and qualitative approaches. 10th International conference on science & technology indicators. Book of abstracts. Vienna, Austria, 17–20 September, 2008 (pp. 53–56). Vienna (AUT): Austrian Research Centers GmbH.
Aknowledgments
This work is part of the CSTG project launched by Antoine Schoen and Bertrand Bellon, a project supported by ANR. The authors are indebted to Sylvain Aubin (DIATOPIE) for term extraction and clustering. They also thank the panel of experts for their commitment: P. Bessières (INRA), A. Lecharny (CNRS-Evry), M. Pinto (Université Paris XI), MM JP Rousset and M Dubow (IGMORS-Université Paris XI), P. Wincler (Genoscope). We also benefited from the experience in the field of Bérangère Virlon (OST). The authors are solely responsible for the views expressed in this article.
Author information
Authors and Affiliations
Corresponding author
Appendices
Annex I List of core journals
ANNUAL REVIEW OF GENOMICS AND HUMAN GENETICS | GENOME |
BIOINFORMATICS | GENOME BIOLOGY |
BMC BIOINFORMATICS | GENOME RESEARCH |
BMC GENOMICS | GENOMICS |
BRIEFINGS IN BIOINFORMATICS | JOURNAL OF PROTEOME RESEARCH |
COMPARATIVE AND FUNCTIONAL GENOMICS | MAMMALIAN GENOME |
CURRENT GENOMICS | MOLECULAR & CELLULAR PROTEOMICS |
CYTOGENETIC AND GENOME RESEARCH | MOLECULAR GENETICS AND GENOMICS |
DNA REPAIR | PHARMACOGENETICS AND GENOMICS |
DNA RESEARCH | PHARMACOGENOMICS |
DNA SEQUENCE | PHARMACOGENOMICS JOURNAL |
EXPERT REVIEW OF PROTEOMICS | PHYSIOLOGICAL GENOMICS |
GENES CHROMOSOMES & CANCER | PROTEOMICS |
Annex II List of themes
The clusters were labelled arbitrarly from M1 to M50 during the clustering process. The most central terms related to each cluster are shown to point out its thematic content
Core genomics
Extension share: high | |
M18/Population_genomics | M32/Marker/RAPD/AFLP/Polymorphism |
M20/Resistance/Resistance_genes/Plant_&_Trout_resistance | M40/QTL/Trait/Mapping/Polymorphism |
M31/LOH/Tumor_suppressor/Genome_&_Cancer | M47/Species/Phylogeny/Evolutionary_genomics |
Extension share: average | |
M 3/Plant_genomics/Transgenic_plants | M25/Patient/Disease_genomics/Biomarkers/Pharmacogenomics |
M 4/DNA_sequence/Satellite | M27/Evolution/Evolutionary_genomics |
M 5/Strain/Microbial_genomics | M28/Cancer/Genome_&_cancer |
M 6/Cell_identity_&_Gene_expression | M35/C-DNA/Transcription/C-DNA_library |
M 8/Alignment/Bioinformatics | M36/Polymorphism |
M12/Network/Biological_networks/Model | M43/Mouse/Murine_genomics |
M14/Locus/Microsatellite_locus/Polymorphism | M44/Expression/Cell_identity_&_Gene_expression |
M15/Cell_line/Tumor/Genome_&_Cancer | M45/LOD/Linkage_analysis/Polymorphism |
M16/Spectrometry/Proteomics | M46/Human/Primate/Gene_annotation/Comparative_genomics |
M22/Human/C-DNA/Gene_annotation | M48/C57BL/Congenic_strains/Murine_genomics |
M23/Exon/Genomic_organization/Gene_annotation | |
Extension share: low | |
M 1/Human_genome/Human_genome_project | M17/Map/Linkage_maps/Polymorphism |
M 9/Genome/ | M24/System/Systems_biology/Bioinformatics |
M10/Comparative_genomic_hybridization/Tumor | M38/Genome/Genome_sizes |
M11/SNPs/Polymorphism |
Border themes
Extension share: high | |
M 2/Translocation/FISH/leukemia | M21/Hybrid/Somatic_hybrids/Ferility |
Extension share: average | |
M13/Transcriptional/Saccharomyces_cerevisiae/Transcriptome | M50/Virus/Virus_replication/Virus_recombinatiio, |
M26/Virus/Nucleotide_sequence | |
Noisy, mostly Non genomics | |
Extension share: high | |
M30/Mutant/Mutagenesis | |
Extension weight: average | |
M 7/Enzyme/Escherichia_Coli | M37/Cell/DNA_damage |
M19/Repair/DNA_damage | M39/DNA/Arrays/Genomic techniques |
M29/Promoter/Transcription | M41/Signaling/Kinase/MAPK |
M33/RNA-/Virus | M42/Mutation/Missence_mutation |
M34/PCR/Methods/applications | M49/Residue/Amino_acid_sequence |
Rights and permissions
About this article
Cite this article
Laurens, P., Zitt, M. & Bassecoulard, E. Delineation of the genomics field by hybrid citation-lexical methods: interaction with experts and validation process. Scientometrics 82, 647–662 (2010). https://doi.org/10.1007/s11192-010-0177-9
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11192-010-0177-9