Abstract
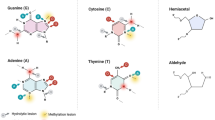
The DNA base lesions in living cells occur permanently and with high frequency as a result of the action of exogenous and endogenous factors. The main mechanism providing removal of such lesions is base excision repair.
Similar content being viewed by others
REFERENCES
Korolev, V.G., Excision Repair of Damaged DNA Bases: DNA Glycosylases, Russ. J. Genet., 2005, vol. 41, no.6, pp. 583–592.
Lindahl, T., Repair of Intrinsic DNA Lesions, Mutat. Res., 1990, vol. 238, pp. 305–311.
Doetsch, P. and Cunningham, R., Enzymology of AP Endonucleases, Mutat. Res., 1990, vol. 236, pp. 173–201.
Otterlei, M., Kavli, B., Standal, R., et al., Repair of Chromosomal Abasic Sites in Vivo Involves at Least Three Different Repair Pathways, EMBO J., 2000, vol. 19, pp. 5542–5551.
Bonura, T., Schultz, R., and Friedberg, E.C., An Enzyme Activity from Escherichia coli That Attacks Single-Stranded Deoxyribopolymers and Single-Stranded Deoxyribonucleic Acid Containing Apyrimidine Sites, Biochemistry, 1982, vol. 21, pp. 2548–2556.
Inone, T. and Kada, T., Purification and Properties of a Bacillus subtilis Endonuclease Specific for Apurinic Sites in DNA, J. Biol. Chem., 1978, vol. 253, pp. 8559–8563.
Ljungquist, S., A New Endonuclease from Escherichia coli Acting at Apurinic Sites in DNA, J. Biol. Chem., 1977, vol. 252, pp. 2808–2814.
Saporito, S.M., Smith-White, B.J., and Cunningham, R.P., Nucleotide Sequence of the xth Gene of Escherichia coli K-12, J. Bacteriol., 1988, vol. 170, pp. 4542–4547.
Yajko, D.M. and Weiss, B., Mutations Simultaneously Affecting Endonuclease II and Exonuclease III in Escherichia coli, Proc. Natl. Acad. Sci. USA, 1975, vol. 72, pp. 688–692.
Ljungquist, S., Lindahl, T., and Howard-Flanders, P., Methyl Methanesulfonate-Sensitive Mutant of Escherichia coli Deficient in an Endonuclease Specific for Apurinic Sites in Deoxyribonucleic Acid, J. Bacteriol., 1976, vol. 126, pp. 646–653.
Demple, B., Halbrook, J., and Linn, S., Escherichia coli xth Mutants Are Hypersensitive to Hydrogen Peroxide, J. Bacteriol., 1983, vol. 153, pp. 1079–1082.
Foster, P.L. and Davis, E.F., Loss of an Apurinic/Apyrimidinic Site Endonuclease Increases the Mutagenicity of N-Methyl-N′-Nitro-Nitrosoguanidine to Escherichia coli, Proc. Natl. Acad. Sci. USA, 1987, vol. 84, pp. 2891–2895.
Weiss, B., Endonuclease II of Escherichia coli Is Exonuclease III, J. Biol. Chem., 1976, vol. 251, pp. 1896–1901.
Henner, W.D., Grunberg, S.M., and Haseltine, W.A., Enzyme Action at 3′ Termini of Ionizing Radiation-Induced DNA Strand Breaks, J. Biol. Chem., 1983, vol. 258, pp. 15 198–15 205.
Demple, B., Johnson, A., and Fung, D., Exonuclease III and Endonuclease IV Remove 3′ Blocks from DNA Synthesis Primers in H2O2-Damaged Escherichia coli, Proc. Natl. Acad. Sci. USA, 1986, vol. 83, pp. 7731–7735.
Chan, E. and Weiss, B., Endonuclease IV of Escherichia coli Is Induced by Paraquat, Proc. Natl. Acad. Sci. USA, 1987, vol. 84, pp. 3189–3193.
Cunningham, R.P., Saporito, S.M., Spizer, S.G., and Weiss, B., Endonuclease IV (nfo) Mutant of Escherichia coli, J. Bacteriol., 1986, vol. 168, pp. 1120–1127.
Takeuchi, M., Lillis, R., Demple, B., and Takeshita, M., Interactions of Escherichia coli Endonuclease IV and Exonuclease III with Abasic Sites in DNA, J. Biol. Chem., 1994, vol. 269, pp. 21 907–21 914.
Johnson, A.W. and Demple, B., Yeast DNA Diesterase for 3′-Fragments of Deoxyribose: Purification and Physical Properties of a Repair Enzyme for Oxidative DNA Damage, J. Biol. Chem., 1988, vol. 263, pp. 18 009–18 016.
Johnson, A.W. and Demple, B., Yeast DNA 3′-Repair Diesterase Is the Major Cellular Apurinic/Apyrimidinic Endonuclease: Substrate Specificity and Kinetics, J. Biol. Chem., 1988, vol. 263, pp. 18 017–18 022.
Popoff, C., Spira, A.I., Johnson, A.W., and Demple, B., Yeast Structural Gene (APN1) for the Major Apurinic Endonuclease: Homology to Escherichia coli Endonuclease IV, Proc. Natl. Acad. Sci. USA, 1990, vol. 87, pp. 4193–4197.
Ramotar, D., Popoff, S.C., Gralla, E.B., and Demple, B., Cellular Role of Yeast Apn1 Apurinic Endonuclease/3′-Diesterase: Repair of Oxidative and Alkylation DNA Damage and Control of Spontaneous Mutation, Mol. Cell. Biol., 1991, vol. 11, pp. 4537–4544.
Kunz, B.A., Henson, E.S., Roche, H., et al., Specificity of the Mutator Caused of Deletion of the Yeast Structural Gene (APN1) for the Major Apurinic Endonuclease, Proc. Natl. Acad. Sci. USA, 1994, vol. 91, pp. 8165–8169.
Ramotar, D., Popoff, S.C., and Demple, B., Complementation of DNA Repair-Deficient Escherichia coli by the Yeast Apn1 Apurinic/Apyrimidinic Endonuclease Gene, Mol. Microbiol., 1993, vol. 5, pp. 149–155.
Johnson, R.E., Torres-Ramos, C.A., Izumi, T., et al., Identification of APN2, the Saccharomyces cerevisiae Homolog of the Major Human AP Endonuclease HAP1, and Its Role in the Repair of Abasic Sites, Genes Dev., 1998, vol. 12, pp. 3137–3143.
Bennett, R.A., The Saccharomyces cerevisiae ETH1 Gene, an Inducible Homolog of Exonuclease III That Provides Resistance to DNA-Damaging Agents and Limits Spontaneous Mutagenesis, Mol. Cell. Biol., 1999, vol. 19, pp. 1800–1809.
Unk, I., Haracska, L., Johnson, R.E., et al., Apurinic Endonuclease Activity of Yeast Apn2 Protein, J. Biol. Chem., 2000, vol. 275, pp. 22 427–22 434.
Unk, I., Haracska, L., Prakash, S., and Prakash, L., 3′-Phosphodiesterase and 3′ → 5′ Exonuclease Activities of Yeast Apn2 Protein and Requirement of These Activities for Repair of Oxidative DNA Damage, Mol. Cell. Biol., 2001, vol. 21, pp. 1656–1661.
Ribar, B., Izumi, T., and Mitra, S., The Major Role of Human AP-Endonuclease Homolog Apn2 in Repair of Abasic Sites in Schizosaccharomyces pombe, Nucleic Acids Res., 2004, vol. 32, pp. 115–126.
Demple, B., Herman, T., and Chen, D.S., Cloning and Expression of APE, the cDNA Encoding the Major Human Apurinic Endonuclease: Definition of Family of DNA Repair Enzymes, Proc. Natl. Acad. Sci. USA, 1991, vol. 88, pp. 11 450–11 454.
Robson, C.N. and Hickson, I.D., Isolation of cDNA Clones Encoding a Human Apurinic/Apyrimidinic Endonuclease That Corrects DNA Repair and Mutagenesis Defects in E. coli xth (Exonuclease III) Mutants, Nucleic Acids Res., 1991, vol. 19, pp. 5519–5523.
Harrison, L., Ascione, L., Menninger, G., et al., Human Apurinic/Apyrimidinic Endonuclease Gene (APE): Structure and Genomic Mapping (Chromosome 14q11.2-12), Hum. Mol. Genet., 1992, vol. 1, pp. 677–680.
Walker, L.J., Graig, R.B., Harris, A.L., and Hickson, I.D., A Role for the Human DNA Repair Enzyme HAP1 in Cellular Protection against DNA Damaging and Hypoxic Stress, Nucleic Acids Res., 1994, vol. 22, pp. 4884–4889.
Chen, D.S., Herman, T., and Demple, B., Two Distinct Human DNA Diesterases That Hydrolyze 3′-Blocking Deoxyribose Fragments from Oxidized DNA, Nucleic Acids Res., 1991, vol. 19, pp. 5907–5914.
Klungland, A., Hoss, M., Gunz, D., et al., Base Excision Repair of Oxidative DNA Damage Activated by XPG Protein, Mol. Cell, 1999, vol. 3, pp. 33–42.
Barzilay, G. and Hickson, I.D., Structure and Function of Apurinic/Apyrimidinic Endonucleases, BioEssays, 1995, vol. 17, pp. 713–719.
Chou, K.-M. and Cheng, Y.-C., An Exonucleolytic Activity of Human Apurinic/Apyrimidinic Endonuclease on 3′ Mispaired DNA, Nature, 2002, vol. 415, pp. 655–659.
Xu, Y.J., Kim, E.Y., and Demple, B., Excision of C-4′-Oxidized Deoxyribose Lesions from Double-Stranded DNA by Human Apurinic/Apyrimidinic Endonuclease (Ape1 Protein) and DNA Polymerase, J. Biol. Chem., 1998, vol. 273, pp. 28 837–28 844.
Hang, B., Chenna, A., Fraenkel-Conrat, H., and Singer, B., A Unusual Mechanism for the Major Human Apurinic/Apyrimidinic (AP) Endonuclease Involving 5′ Cleavage of DNA Containing a Benzene-Derived Exocyclic Adduct in the Absence of an AP Site, Proc. Natl. Acad. Sci. USA, 1996, vol. 93, pp. 13 737–13 741.
Parsons, J.L., Dianova, I.I., and Dianov, G.L., APE1 Is the Major 3′-Phosphoglycolate Activity in Human Cell Extracts, Nucleic Acids Res., 2004, vol. 32, pp. 3531–3536.
Wilson, D.M. III and Barsky, D., The Major Human Abasic Endonuclease: Formation, Consequences and Repair of Abasic Lesions in DNA, Mutat. Res., 2001, vol. 485, pp. 283–307.
Hadi, M.Z. and Wilson, D.M. III, A Second Human Protein with Homology to the Escherichia coli Abasic Endonuclease Exonuclease III, Environ. Mol. Mutagen., 2000, vol. 36, pp. 312–324.
Gorman, M.A., Morera, S., Rothwell, D.G., et al., The Crystal Structure of the Human DNA Repair Endonuclease HAP1 Suggests the Recognition of Extra-Helical Deoxyribose at DNA Abasic Sites, EMBO J., 1997, vol. 16, pp. 6548–6558.
Mol, C.D., Izumi, T., Mitra, S., and Tainer, J.A., DNA-Bound Structures and Mutants Reveal Abasic DNA Binding by APE1 DNA Repair and Coordination, Nature, 2000, vol. 403, pp. 451–455.
Yang, H., Clendenin, W.M., Wong, D., et al., Enhanced Activity of Adenine-DNA Glycosylase (Myh) by Apurinic/Apyrimidinic Endonuclease (Ape1) in Mammalian Base Excision Repair of an A/GO Mismatch, Nucleic Acids Res., 2001, vol. 29, pp. 743–752.
Vidal, A.E., Hikson, I.D., Boiteux, S., and Radicella, J.P., Mechanism of Stimulation of the DNA Glycosylase Activity of hOGG1 by the Major Human AP Endonuclease: Bypass of the AP Lyase Activity Step, Nucleic Acids Res., 2001, vol. 29, pp. 1285–1292.
Hill, J.W., Hazra, T.K., Izumi, T., and Mitra, S., Stimulation of Human 8-Oxoguanine-DNA Glycosylase by AP-Endonuclease: Potential Coordination of the Initial Steps in Base Excision Repair, Nucleic Acids Res., 2001, vol. 29, pp. 430–438.
Ischenko, A.A. and Saparbaev, M.R., Alternative Nucleotide Incision Repair Pathway for Oxidative DNA Damage, Nature, 2002, vol. 415, pp. 183–187.
Gros, L., Ishchenko, A.A., Ide, H., et al., The Major Human Endonuclease (Ape1) Is Involved in the Nucleotide Incision Repair Pathway, Nucleic Acids Res., 2004, vol. 32, pp. 73–81.
Wang, Z., Wu, X., and Friedberg, E.C., DNA Repair Synthesis during Base Excision Repair in Vitro Is Catalyzed by DNA Polymerase ε and Is Influenced by DNA Polymerases α and β in Saccharomyces cerevisiae, Mol. Cell. Biol., 1993, vol. 13, pp. 1051–1058.
Blank, A., Kim, B., and Loeb, L.A., DNA Polymerase δ Is Required for Base Excision Repair of DNA Methylation Damage in Saccharomyces cerevisiae, Proc. Natl. Acad. Sci. USA, 1994, vol. 91, pp. 9047–9051.
Suszek, W., Baranowska, H., Zuk, J., and Jachymczyk, W.J., DNA Polymerase III Is Required for DNA Repair in Saccharomyces cerevisiae, Curr. Genet., 1993, vol. 24, pp. 200–204.
Leem, S.-H., Ropp, P.A., and Sugino, A., The Yeast Saccharomyces cerevisiae DNA Polymerase IV: Possible Involvement in Double Strand Break DNA Repair, Nucleic Acids Res., 1994, vol. 22, pp. 3011–3017.
Prasad, R., Widen, S.G., Singhal, R.K., et al., Yeast Open Reading Frame YCR14C Encodes a DNA β-Polymerase-Like Enzyme, Nucleic Acids Res., 1994, vol. 21, pp. 5301–5307.
McInnis, M., O'Neill, G., Fossum, K., and Reagan, M.S., Epistatic Analysis of the Roles of the RAD27 and POL4 Gene Products in DNA Base Excision Repair in S. cerevisiae, DNA Repair, 2002, vol. 1, pp. 311–315.
Wu, X. and Wang, Z., Relationships between Yeast Rad27 and Apn1 in Response to Apurinic/Apyrimidinic (AP) Sites in DNA, Nucleic Acids Res., 1999, vol. 27, pp. 956–962.
Harrington, J.J. and Lieber, M.R., Functional Domains within FEN-1 and RAD2 Define a Family of Structure-Specific Endonucleases: Implications for Nucleotide Excision Repair, Genes Dev., 1994, vol. 8, pp. 1344–1355.
Zhu, F.X., Biswas, E.E., and Biswas, S.B., Purification and Characterization of the DNA Polymerase α-Associated Exonuclease: The RTH1 Gene Product, Biochemistry, 1997, vol. 36, pp. 5947–5954.
Kozhina, T.N., Kozhin, S.A., Latypov, V.F., and Korolev, V.G., RAD29 and RAD31, New Genes of Yeast Saccharomyces cerevisiae Involved in DNA Repair Control: Isolation and Genetic Characterization of Mutants, Russ. J. Genet., 2000, vol. 36, no.6, pp. 627–633.
Kozhin, S.A., Kozhina, T.N., Latypov, V.F., and Korolev, V.G., RAD29 and RAD31, New Genes of the Yeast Saccharomyces cerevisiae Involved in DNA Repair Control: Determining Possible Functions of These Genes, Russ. J. Genet., 2000, vol. 36, no.8, pp. 845–852.
Unk, I., Haracska, L., Gomes, X.V., et al., Stimulation of 3′ → 5′ Exonuclease and 3′-Phosphodiesterase Activities of Yeast Apn2 by Proliferating Cell Nuclear Antigen, Mol. Cell. Biol., 2002, vol. 22, pp. 6480–6486.
Sobol, R.W., Horton, J.K., Kuh, R., et al., Requirement of Mammalian DNA Polymerase β in Base Excision Repair, Nature, 1996, vol. 379, pp. 183–186.
Dianov, G., Price, A., and Lindahl, T., Generation of Single-Nucleotide Repair Patches Following Excision of Uracil Residues from DNA, Mol. Cell. Biol., 1992, vol. 12, pp. 1605–1612.
Klungland, A. and Lindahl, T., Second Pathway for Completion of Human DNA Base Excision Repair: Reconstitution with Purified Proteins and Requirement for DNAase IV (FEN1), EMBO J., 1997, vol. 16, pp. 3341–3348.
Frosina, G., Fortini, P., Rossi, O., et al., Two Pathways for Base Excision Repair in Mammalian Cells, J. Biol. Chem., 1996, vol. 271, pp. 9573–9578.
Matsumoto, Y. and Kim, K., Excision of Deoxyribose Phosphate Residues by DNA Polymerase β during DNA Repair, Science, 1995, vol. 269, pp. 699–702.
Kumar, A., Widen, S.G., Williams, K.R., et al., Studies of the Domain Structure of Mammalian DNA Polymerase: Identification of a Discrete Template Binding Domain, J. Biol. Chem., 1990, vol. 265, pp. 2124–2131.
Kumar, A., Abbotts, J., Karawya, E., and Wilson, S.H., Identification and Properties of the Catalytic Domain of Mammalian DNA Polymerase β, Biochemistry, 1990, vol. 29, pp. 7156–7159.
Sugo, N., Aratani, Y., Nagashima, Y., et al., Neonatal Lethality with Abnormal Neurogenesis in Mice Deficient in DNA Polymerase, EMBO J., 2000, vol. 19, pp. 1397–1404.
Fortini, P., Pascucci, B., Belisario, F., and Dogliotti, E., DNA Polymerase β Is Required for Efficient DNA Strand Break Repair Induced by Methyl Methanesulfonate but Not by Hydrogen Peroxide, Nucleic Acids Res., 2000, vol. 28, pp. 3040–3046.
Sobol, R.W., Watson, D.E., Nakamura, J., et al., Mutations Associated with Base Excision Repair Deficiency and Methylation-Induced Genotoxic Stress, Proc. Natl. Acad. Sci. USA, 2002, vol. 99, pp. 6860–6865.
Singhal, R.K., Prasad, R., and Wilson, S.H., DNA Polymerase β Conducts the Gap-Filling Step in Uracil-Initiated Base Excision Repair in a Bovine Testis Nuclear Extract, J. Biol. Chem., 1995, vol. 270, pp. 949–957.
Prasad, R., Singhal, R.K., Srivastava, D.K., et al., Specific Interaction of DNA Polymerase β and DNA Ligase I in a Multiprotein Base Excision Repair Complex from Bovine Testis, J. Biol. Chem., 1996, vol. 271, pp. 16 000–16 007.
Allison, S.L., Dianova, I.I., and Dianov, G.L., DNA Polymerase β Is the Major DRP Lyase Involved in Repair of Oxidative Base Lesions in DNA by Mammalian Cell Extracts, EMBO J., 2001, vol. 20, pp. 6919–6926.
Dianov, G.L., Thybo, T., Dianova, I.I., et al., Single Nucleotide Patch Base Excision Repair Is the Major Pathway for Removal of Thymine Glycol from DNA in Human Cell Extracts, J. Biol. Chem., 2000, vol. 275, pp. 11 809–11 813.
Saitoh, T., Shinmura, K., Yamaguchi, S., et al., Enhancement of OGG1 Protein AP Lyase Activity by Increase of APEX Protein, Mutat. Res., 2001, vol. 486, pp. 31–40.
Podlucsky, A.J., Dianova, I.I., Podust, V.N., et al., Human DNA Polymerase β Initiates DNA Synthesis during Long-Patch Repair of Reduced AP Sites in DNA, EMBO J., 2001, vol. 20, pp. 1477–1482.
Dianov, G.L., Prasad, R., Wilson, S.H., and Bohr, V.A., Role of DNA Polymerase β in the Excision Step of Long Patch Mammalian Base Excision Repair, J. Biol. Chem., 1999, vol. 274, pp. 13 741–13 743.
Fortini, P., Pascucci, B., Parlanti, E., et al., Different DNA Polymerases Are Involved in the Short-and Long-Patch Base Excision Repair in Mammalian Cells, Biochemistry, 1998, vol. 37, pp. 3575–3580.
Parikh, S.S., Mol, C.D., Slupphang, G., et al., Base-Excision Repair Initiation Revealed by Crystal Structures and DNA-Binding Kinetics of Human Uracil-DNA Glycosylase Bound to DNA, EMBO J., 1998, vol. 17, pp. 5414–5426.
Waters, T.R., Gallinari, P., Jiricny, J., and Swann, P.F., Human Thymine DNA Glycosylase Binds to Apurinic Sites in DNA but Is Displaced by Human Apurinic Endonuclease 1, J. Biol. Chem., 1999, vol. 274, pp. 67–74.
Bennett, R.A.O., Wilson, D.M. III, Wong, D., and Demple, B., Interaction of Human Apurinic Endonuclease and DNA Polymerase β in the Base Excision Repair Pathways, Proc. Natl. Acad. Sci. USA, 1997, vol. 94, pp. 7166–7169.
Ranalli, T.A., Tom, S., and Bambara, R.A., AP Endonuclease 1 Coordinates Flap Endonuclease 1 and DNA Ligase 1 Activity in Long Patch Base Excision Repair, J. Biol. Chem., 2002, vol. 277, pp. 41 715–41 724.
Dianova, I.I., Bohr, V.A., and Dianov, G.L., Interaction of Human AP Endonuclease 1 with Flap Endonuclease 1 and Proliferating Cell Nuclear Antigen Involved in Long-Patch Base Excision Repair, Biochemistry, 2001, vol. 40, pp. 12 639–12 644.
DeMott, M.S., Zigman, S., and Bambara, R.A., Replication Protein A Stimulates Long Patch DNA Base Excision Repair, J. Biol. Chem., 1998, vol. 273, pp. 27 492–27 498.
Prasad, R., Dianov, G.L., Bohr, V.A., and Wilson, S.H., FEN1 Stimulation of DNA Polymerase β an Excision Step in Mammalian Long Patch Base Excision Repair, J. Biol. Chem., 2000, vol. 275, pp. 4460–4466.
Caldecott, K.W., Mammalian DNA Single-Strand Break Repair: An X-ra(y)ted Affair, Bioessays, 2001, vol. 23, pp. 447–455.
Marsin, S., Vidal, A.E., Sossou, M., et al., Role of XRCC1 in the Coordination and Stimulation of Oxidatis8-44 074.
Vidal, A.E., Boiteux, S., Hickson, I.D., and Radicella, J.P., XRCC1 Coordinates the Initial and Late Stages of DNA Abasic Site Repair through Protein-Protein Interactions, EMBO J., 2001, vol. 20, pp. 6530–6539.
Marintchev, A., Robertson, A., Dimitriadis, E.K., et al., Domain-Specific Interaction in the XRCC1-DNA Polymerase β Complex, Nucleic Acids Res., 2000, vol. 28, pp. 2049–2059.
Kubata, Y., Nash, R.A., Klungland, A., et al., Reconstitution of DNA Base Excision-Repair with Purified Human Proteins: Interaction between DNA Polymerase β and the XRCC1 Protein, EMBO J., 1996, vol. 15, pp. 6662–6670.
Cappelli, E., Taylor, R., Cevasco, M., et al., Involvement of XPCC1 and DNA Ligase III Gene Products in DNA Base Excision Repair, J. Biol. Chem., 1997, vol. 272, pp. 23 970–23 975.
Dantzer, F., de la Rubia, G., Menissier-de Murcia, J., et al., Base Excision Repair Is Impaired in Mammalian Cells Lacking Poly(ADP-Ribose) Polymerase-1, Biochemistry, 1999, vol. 39, pp. 7559–7569.
Caldecott, K.W., Aoufouchi, S., Johnson, P., and Shall, S., XRCC1 Polypeptide Interacts with DNA Polymerase β and Possible Poly(ADP-Ribose) Polymerase and DNA Ligase III and Is a Novel Molecular “Nick-Sensor” in Vitro, Nucleic Acids Res., 1996, vol. 24, pp. 4387–4394.
Allison, S.L., Dianova, I.I., and Dianov, G.L., Poly(ADP-Ribose) Polymerase in Base Excision Repair: Always Engaged, but Not Essential for DNA Damage Processing, Acta Biochem. Polonica, 2003, vol. 50, pp. 169–179.
Author information
Authors and Affiliations
Additional information
__________
Translated from Genetika, Vol. 41, No. 10, 2005, pp. 1301–1309.
Original Russian Text Copyright © 2005 by Korolev.
Rights and permissions
About this article
Cite this article
Korolev, V.G. Base Excision Repair: AP Endonucleases and DNA Polymerases. Russ J Genet 41, 1063–1070 (2005). https://doi.org/10.1007/s11177-005-0201-y
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11177-005-0201-y