Abstract
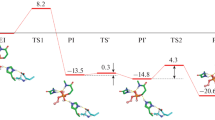
The quantum mechanics/molecular mechanics method was used to model the mechanism of hydrolysis of succinylcholine in the active site of native butyrylcholinesterase and the Asp70Gly single mutant enzyme. The energy diagram for the reaction pathway of the acylation was constructed, and the effect of the mutation on the reaction mechanism and energy characteristics of the process are discussed.
Similar content being viewed by others
References
M. Barbosa, O. Rios, M. Velásquez, J. Villalobos, J. Ehrmanns, Surg. Neurol., 2001, 55, 106.
C.-G. Zhan, F. Zheng, D. W. Landry, J. Am. Chem. Soc., 2003, 125, 2462.
L. G. Sokolovskaya, L. V. Sigolaeva, A. V. Eremenko, I. N. Kurochkin, G. F. Makhaeva, V. V. Malygin, I. E. Zykova, V. I. Kholstov, N. V. Zav’yalova, S. D. Varfolomeev, Khim. Biol. Bezopasnost’ [Chem. Biol. Safety], 2004, 13–14, 21 (in Russian).
M. Pirmohamed, B. K. Park, Trends Pharmacol. Sci., 2001, 22, 298.
M. C. Mcguire, C. P. Nogueira, C. F. Bartels, H. Light-stone, A. Hajra, A. F. Van der Spek, O. Lockridge, B. N. La Du, Proc. Natl. Acad. Sci. USA, 1989, 86, 953.
D. Suarez, N. Diaz, J. Fontecilla-Camps, M. J. Field, Bio-chemistry, 2006, 45, 7529.
C.-G. Zhan, D. Gao, Biophys. J., 2005, 89, 3863.
P. Masson, P. Legrand, C. F. Bartels, M.-T. Froment, L. M. Schopfer, O. Lockridge, Biochemistry, 1997, 36, 2266.
S. D. Varfolomeev, Khimicheskaya enzimologiya [Chemical Enzymology], Akademiya, Moscow, 2005, 480 pp. (in Russian).
K. B. Bravaya, A. V. Bochenkova, B. L. Grigorenko, I. A. Topol, S. Burt, A. V. Nemukhin, J. Chem. Theory Comput., 2006, 2, 1168.
B. L. Grigorenko, A. V. Nemukhin, M. S. Shadrina, I. A. Topol, S. K. Burt, Proteins: Structure, Function, Bioinformatics, 2007, 66, 456.
B. L. Grigorenko, A. V. Rogov, I. A. Topol, S. K. Burt, H. M. Martinez, A. V. Nemukhin, Proc. Natl. Acad. Sci., 2007, 104, 7057.
A. V. Nemukhin, S. V. Lushchekina, A. V. Bochenkova, A. A. Golubeva, S. D. Varfolomeev, J. Mol. Model., 2008, 14, 409.
Y. Zhang, J. Kua, J. A. McCammon, J. Am. Chem. Soc., 2002, 124, 10572.
H. M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T. N. Bhat, H. Weissig, I. N. Shindyalov, P. E. Bourne, Nucleic Acids Res., 2000, 28, 235.
J. M. Word, S. C. Lovell, T. H. Labean, H. C. Taylor, M. E. Zalis, B.K. Presley, J. S. Richardson, D. C. Richardson, J. Mol. Biol., 1999, 285, 1711.
D. M. Quinn, Chem. Rev., 1987, 87, 955.
M. A. Massiah, C. Viragh, P. M. Reddy, I. M. Kovach, J. Johnson, T. L. Rosenberry, A. S. Mildvan, Biochemistry, 2001, 40, 5682.
G. M. Morris, D. S. Goodsell, R. S. Halliday, R. Huey, W. E. Hart, R. K. Belew, A. J. Olson, J. Am. Chem. Soc., 1998, 19, 1639.
J. W. Ponder, TINKER: Software Tools for Molecular Design, Version 3.7, 1999, Washington University: St. Louis.
T. A. Halgren, J. Comput. Chem., 1996, 17, 490.
B. L. Grigorenko, A. V. Nemukhin, I. A. Topol, S. K. Burt, J. Phys. Chem. A, 2002, 106, 10663.
A. V. Nemukhin, B. L. Grigorenko, I. A. Topol, S. K. Burt, J. Comput. Chem., 2003, 24, 1410.
J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman, D. A. Case, J. Comput. Chem., 2004, 25, 1157.
A. A. Granovsky, PC GAMESS/Firefly version 7.1.F, http://classic.chem.msu.su/gran/gamess/index.html.
P. Masson, M.-T. Froment, P. L. Fortier, J. E. Visicchio, C. F. Bartels, O. Lockridge, Biochim. Biophys. Acta Prot. Struct. Mol. Enzym., 1998, 1387, 41.
M. Fuxreiter, A. Warshel, J. Am. Chem. Soc., 1998, 120, 183.
A. V. Nemukhin, B. L. Grigorenko, A. V. Rogov, I. A. Topol, S. K. Burt, Theor. Chem. Acc., 2004, 111, 36.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Izvestiya Akademii Nauk. Seriya Khimicheskaya, No. 1, pp. 56–61, January, 2010.
Rights and permissions
About this article
Cite this article
Lushchekina, S.V., Grigorenko, B.L., Morozov, D.I. et al. Modeling of the mechanism of hydrolysis of succinylcholine in the active site of native and modified (Asp70Gly) human butyrylcholinesterase. Russ Chem Bull 59, 55–60 (2010). https://doi.org/10.1007/s11172-010-0044-0
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11172-010-0044-0