Abstract
Graves’ disease (GD) is the commonest cause of hyperthyroidism and has a strong female preponderance. Everyday clinical practice suggests strong aggregation within families and twin studies demonstrate that genetic factors account for 60-80% of risk of developing GD. In this review, we collate numerous genetic studies and outline the discoveries over the years, starting with historic candidate gene studies and then exploring more recent genome-wide linkage and association studies, which have involved substantial cohorts of East Asian patients as well as those of European descent. Variants in genes including HLA, CTLA4, and PTPN22 have been shown to have substantial individual effects on disease susceptibility. In addition, we examine emerging evidence concerning the possibility that genetic variants may correlate with relevant clinical phenotypes including age of onset of GD, severity of thyrotoxicosis, goitre size and relapse of hyperthyroidism following antithyroid drug therapy, as well as thyroid eye disease. This review supports the inheritance of GD as a complex genetic trait, with a growing number of more than 80 susceptibility loci identified so far. Future implementation of more targeted clinical therapies requires larger studies investigating the influence of these genetic variants on the various phenotypes and different outcomes of conventional treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Epidemiology and heritability
Graves' disease (GD) is the commonest cause of hyperthyroidism affecting approximately 3% of women and 0.5% of men during their lifetime [1]. GD, in common with most other autoimmune diseases, has a clear female preponderance with a female to male ratio of 6-7:1. The incidence was 0.04% per year during a 12-year follow-up of the Nurses Health II study in North America [2]. Interestingly, a recent longitudinal study of 22 million people from the UK Clinical Practice Research Datalink showed a doubling in incidence of GD, from 0.03% per year in 2000-2002 to 0.07% per year by 2017-2019 [3]. In members of the US armed forces, Graves’ disease occurred nearly twice as commonly in Black and Asian/Pacific Islander women than in White women [4].
Twin studies in the Danish and Swedish populations suggest strong genetic aggregation, with a concordance rate for monozygotic twins of between 20% and 35%, compared to 2-3% in dizygotic twins [5, 6], fitting a model where genetic factors account for 60-80% of the risk of developing GD [5, 6]. An estimate of heritability can also be given by the sibling recurrence risk ratio, known as λs. A Hungarian study showed that 23 of 435 (5.3%) GD probands had siblings with GD (21 sisters, 2 brothers), compared with a background population frequency of GD of 0.65% [7]. A similar study performed in Newcastle, UK showed 15 of 190 (7.9%) GD probands had similarly affected siblings (9 sisters, 6 brothers), compared with a background prevalence of GD of 0.8% [8]. This allows the calculation of λs for GD as 8–10, which compares to that of 8 for rheumatoid arthritis, 15 for type 1 diabetes and 20 for multiple sclerosis [9]. This supports that GD, in common with many other autoimmune conditions is inherited as a complex genetic trait, with strong familial clustering but without a classical Mendelian pattern of inheritance [5, 6, 8, 9].
A key question given the large 60-80% genetic contribution to disease susceptibility is what constitutes the remaining non-genetic or potential ‘precipitating’ factors. Although outside the scope of this review, a longitudinal study of Dutch people who had a sibling with autoimmune thyroid disease (AITD) showed that pregnancy was a risk factor for development of GD, whereas oestrogen use was protective [10]. In addition, there are high quality studies indicating that stress, including post-traumatic stress disorder, cigarette use and iodine status also contribute as non-genetic factors that determine the manifestation of GD [11,12,13].
Although there are a few studies that have investigated the evolving field of epigenetics of GD, several of these are small with inadequate power, and no independently replicated results have been found thus far [14,15,16,17]. For these reasons, the remainder of this review will focus on germline inherited genetic variation in Graves’ disease.
2 Historical genetic studies: candidate gene studies and the trinity of organ specific autoimmunity
Early genetic studies focussed on alleles of the major histocompatibility complex (MHC), primarily because this was one of the first loci for which reliable markers could be used to infer genotype. Using serological assays for human leukocyte antigen (HLA) typing initially allowed strong association to be established between the MHC alleles (“HL-A8”) on the short arm of chromosome 6 and GD [18]. This has now been refined such that in populations of European decent, an associated haplotype HLA-DRB1*0301-DQB1*0201-DQA1*0501 has been confirmed in many different studies, with an odds ratio between 2 and 3 compared to the unaffected population [19, 20]. In Asian populations, different HLA associations have been confirmed, including DRB1*0405 and DRB1*1403 in Japanese [21, 22], DRB1*0803 and DRB1*1602 in Koreans [23], and DRB1*1602 in the Thai population [24]. The associated DRB1*0301 allele encodes for an arginine residue at position 74 of the MHC-DRβ chain, and this has been hypothesized to change the binding pocket of the MHC to facilitate binding of T lymphocyte peptide antigen [25]. Nevertheless, as only around 50% of GD patients of European ancestry carry the DRB1*0301 haplotype, it suggests that MHC-restricted presentation of a single antigenic peptide is unlikely to account for all cases of GD.
Following on from the identification of MHC as a GD locus, a candidate gene study examined a microsatellite repeat polymorphism in the then recently discovered cytotoxic T-lymphocyte antigen-4 (CTLA4) gene. CTLA4 was an excellent candidate gene for an autoimmune disorder as a negative costimulatory molecule, present as a ‘second signal’ modulating T cell receptor activation in response to MHC-antigen presentation at the immunological synapse. Even though this study was relatively small with only 133 Graves’ patients [26], it was strongly positive and has been replicated on more than 50 occasions since then [26, 27]. In addition, association with CTLA4 alleles has been confirmed in Type 1 diabetes, coeliac disease and numerous different organ-specific autoimmune conditions. Interestingly, the susceptibility allele for autoimmunity at CTLA4 occurs in more than 50% of the healthy population, and appears to be enriched in patients with thyroid eye disease [28].
Subsequent to its identification as a susceptibility allele for type 1 diabetes [29], a missense polymorphism (R620W) in the lymphoid tyrosine phosphatase (LYP, encoded by the PTPN22 gene) was found to be associated with GD [30, 31]. This functions as another negative regulator of T lymphocyte signalling, and the associated allelic variant prevents the LYP protein from interacting with its inhibitory partner Csk, leading to unchecked TCR signalling. The PTPN22*620W allele is found in increasing frequency in Northern European countries as compared to populations in Southern Europe, and is virtually absent in Asian or African populations [23], suggesting that the variant allele may have been selected for ability to combat certain infectious disease.
Thus, by the end of the last century, alleles at the MHC, CTLA4 and PTPN22 loci had been robustly confirmed as contributing to GD susceptibility largely through the candidate gene approach. These same alleles had also been associated with Hashimoto’s thyroiditis and many different autoimmune disorders in which they were presumed to have an effect on immune system function. However, they did not explain why the thyroid gland is targeted by the immune response to give GD.
3 Discovery-based genomics and early genome-wide association studies in European-ancestry populations
Although candidate gene studies were successful in identifying several GD susceptibility genes with large effects, the approach is limited in that one cannot discover an unsuspected biological basis for the disease using this approach. With this substantial limitation in mind, discovery-based approaches using genome-wide linkage analyses of families with two or more affected relatives with AITD were published by US and European investigators [32, 33]. The US study of 56 multigenerational families replicated the known linkage at MHC [34], and showed novel evidence for linkage at three other loci labelled as GD-1 (D14S81 on chromosome 14q31), GD-2 (D20S195 on chromosome 20q11.2) and GD-3 (DXS8020 on chromosome Xq21) [32]. The linkage to 14q31 and 20q11 was later replicated in a larger dataset of 102 multiplex families, although that to the X chromosome was not [35].
A disadvantage of family-based linkage studies over unrelated proband association analysis is that linkage identifies large chromosomal regions containing many potentially relevant genes and numerous allelic variants. The large gene region on 14q31 encompassing the GD-1 locus included the gene encoding the TSH receptor (TSHR), although consistent association was difficult to demonstrate in further studies. Subsequently, with an apparently un-ironic affiliation from “Target Discovery at Oxagen”, TSHR was confirmed to be a disease specific locus for GD using two independent European patient cohorts [36]. Non-coding variants within intron 1 of the TSHR gene were identified as being associated with GD by several groups [37, 38]. TSHR is expressed at low levels in the thymus presumably to allow T lymphocyte negative selection against TSHR as a “self antigen”, and people carrying the non-coding disease-associated variants were found to have lower levels of thymic TSHR mRNA expression. The hypothesis suggests that thymic T cells carrying T cell receptors specific for TSHR peptide may escape clonal deletion because of insufficient TSHR expression, leading to TSHR autoreactivity and thence GD [39].
Additional genome wide linkage studies were performed by the Oxagen investigators in 1,119 relative pairs of European origin with AITD [33] but failed to find significant linkage of any of the previously known loci including MHC, CTLA4 and PTPN22. This study instead reported linkage with 3 regions for GD that included chromosomes 2q36, 11p15 and 18p11. The strongest region of linkage for GD was identified to marker D18S53 on chromosome 18p11 that had not been previously identified [33], and in retrospect this signal probably arose from the PTPN2 gene, which has subsequently been confirmed as a GD locus. Similarly, a genome wide association scan published by the Wellcome Trust Case Control Consortium (WTCCC) included 1000 independent cases of GD, along with patients with ankylosing spondylitis, multiple sclerosis and breast cancer and compared these to 1500 randomly selected healthy British individuals from the 1958 Birth Cohort [40]. Associations were observed with SNPs in the MHC region with robust p-values of <10-20. Association was also reported and confirmed in GD patients at the TSHR locus and at a novel locus FCRL3 [40]. The FCRL3 association was rapidly replicated, not only in patients with GD [41], but also in rheumatoid arthritis and several other autoimmune conditions [41, 42]. Although the WTCCC study identified several other associations, at the time these could not be replicated [43]. That these two studies failed to replicate several previously confirmed loci was likely due to mixing cases of GD and Hashimoto thyroiditis together as well as low power, emphasizing that numerous loci contribute to complex autoimmune disease pathogenesis and that larger and more phenotypically homogeneous cohorts are needed to provide robust results.
4 Expanding the genomic findings to Asian populations
In parallel with discovery-based genomic approaches in patients of European origin, investigators working in Japan and China began performing similar studies. Independent studies from both Japanese and Chinese groups using 123 and 54 multiplex families, respectively, both pointed to a susceptibility locus at chromosome 5q31-33 [44, 45]. Although initially suspected to relate to the presence of a cluster of interleukin genes (IL3, IL4, IL5 and GM-CSF) at this location, a more substantial association study in 2,800 Chinese Graves’ patients suggested that variants in the promoter of the Secretoglobin Family 3A Member 2 (SCGB3A2, encoding uterus globulin associated protein-1:UGRP1) gene might be responsible [46]. UGRP1 is secreted in lung surfactant and bronchial epithelia, is aberrantly expressed in thyroid tissue from both Hashimoto and GD patients, as well as being a downstream target of thyroid transcription factor-1 [47]. This finding of association with alleles of SCGB3A2 was quickly replicated in a UK cohort by Simmonds et al. indicating that this is also a susceptibility locus in Caucasian patients [48].
In a staged fashion over several years, the China Consortium for the Genetics of Autoimmune Thyroid Disease published larger association studies, culminating in a genome-wide analysis containing more than 9,500 individuals with GD [49, 50]. Cumulatively, these studies robustly identified eight new susceptibility alleles including at RNASET2 (6q27), GPR174 (Xq21.1), C1QTNF6-RAC2 (22q12.3-13.1), SLAMF6 (1q23.2), ABO (9q34.2), Thyroglobulin (8q24.22) and intergenic regions at 4p14 and 14q32.2 [49, 50]. In addition, associations at MHC, CTLA4, FCRL3 and TSHR were confirmed [51]. The susceptibility variant in the PTPN22 gene (620*W) is not present in Asian populations so association could not be directly replicated, however, there was modest evidence for a different allele at PTPN22 having a role in Chinese Graves’ patients [52].
Another large three-stage genome wide association study carried out by Liu et al. in 2013 [52] showed that alleles of the BACH2 gene were associated with GD in the Chinese Han population. BACH2 had previously been identified as harbouring susceptibility variants in both Graves’ and Hashimoto’s thyroiditis patients in a smaller UK population [53]. A detailed analysis of 331-kb region in the BACH2 gene in more than 3000 patients with GD and 1468 controls showed that a SNP (rs2474619) in intron 2 was most likely to be driving the disease association, and this variant was correlated with BACH2 gene expression. BACH2 deficient mice have defective immunoglobulin class switch recombination and BACH2 is a repressor of plasma cell differentiation, being essential for normal B cell maturation [54]. Additional studies in other autoimmune disorders have confirmed association of the same BACH2 alleles with Addison’s disease, type 1 diabetes, coeliac disease, and rheumatoid arthritis [55,56,57]. Thus, the large patient datasets particularly available to Chinese researchers allowed the robust identification of many novel GD loci during 2000-2010s which have subsequently been confirmed, as well as replication of some of the earlier associations found using the candidate gene approach.
5 Large GWAS in European-ancestry populations
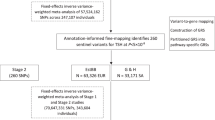
A large genome-wide association study in a mixed cohort of 30,234 cases of AITD (both Graves’ and Hashimoto thyroiditis) and 72,172 controls from Iceland and the UK Biobank found 99 genome wide significant associations at 93 loci (Fig. 1)[58]. Of the 99 associations, 84 had not been previously associated with AITD. 37 candidate genes were supported by systematic annotation. 24 of these encoded proteins were found to be directly or indirectly linked in the same functional network. Interestingly, the study found two novel, but low frequency variants that showed comparatively large effects (Odds ratio around 1.5) on the susceptibility to AITD: an intronic variant in the FLT3 gene (rs76428106-C) and a missense variant in ADCY7 (rs78534766-A). FLT3 gene variants had not previously been associated with any disease, whilst an ADCY7 variant was previously reported to be associated with ulcerative colitis. The FLT3 variant is intronic but introduces cryptic splice site and a premature stop codon predicting a truncated protein. The association was replicated in patient cohorts with systemic lupus erythematous, rheumatoid arthritis and coeliac disease [58].
Genome-wide association scan in 30,000 patients with autoimmune thyroid disease- top 10 susceptibility loci. Data taken from Saevarsdottir et al. [58], show the balance between the odds ratio of the susceptibility allele predisposing to disease vs the allele frequency in the population. Bubble size/colour represents the probability in favour of association (-log10 P value). It is notable that probability for association is less for alleles with very low (FLT3, ADCY7) and very high (IFIH1, TYK2) population frequency, owing to low power.
Overall, only 15 of 99 variants had effect sizes with odds ratios over 1.10, and these comprised a mixture of common alleles (eg. HLA, CTLA4, PTPN22, SH2B3) that had previously been confirmed, and some rarer variants that could only be discovered once an adequately powered patient cohort had been studied (Fig. 1). Most of the genetic variants found had similar effects in both GD and Hashimoto thyroiditis (eg. CTLA4, PTPN22, RNASET2), but a small number including the FLT3 variant appeared to predispose to Hashimoto thyroiditis but confer modest protection against GD. Although this study is the largest genetic study of AITD to date, it has to be considered that there were only 2,400 patients with GD analysed, compared to more than 27,000 with hypothyroidism or Hashimoto thyroiditis meaning there is a compromise in design, sacrificing the high power gained by having data from 30,000 individuals for the potential to miss important loci, as GD becomes a subphenotype of the analysis. Overall, it suggests that there may still be much to learn concerning the specific susceptibility to GD.
6 Can genetic information inform clinical management in Graves’ disease?
While we have reviewed the genetic predisposition to GD above, important questions remain about whether there are the genotype-phenotype correlations, and if these genetic variations might have prognostic significance for patients. The phenotypic spectrum of GD is broad with patients exhibiting diverse clinical features and differing in biochemical severity. Several studies have demonstrated robust associations between specific clinical and biochemical factors and outcome in GD following withdrawal of antithyroid drugs (ATD), and have also identified prognostically relevant genetic associations.
Examining the association of genetic loci and clinical phenotype may not only provide mechanistic insight into the pathogenesis of GD, but also promote precision medicine by enabling the prediction of an individual’s risk of relapse and their likely response to different therapeutic approaches.
7 Genotype-phenotype correlation in Graves’ disease
Candidate-gene association studies have revealed that some of the genetic variants associated with the predisposition to GD have also been associated with specific clinical and biochemical phenotypes. Several of these variants have also been implicated in predicting the outcome of GD, including both thyroid-specific (TSHR and TG) and immune-system genes (e.g. CTLA4, CD40, PTNP22 and HLA) (Table 1).
7.1 Genetic variants and clinical/biochemical phenotype
Polymorphisms in both the immune-system and thyroid-specific genes have been associated with particular clinical and biochemical phenotypes. Some of the genetic variants independently associated with recurrence of GD are also associated with an earlier age of onset of GD (CTLA4, HLA, PTPN22, and TSHR) [60,61,62,63]. There are also limited data that implicate other genetic variants, such as an intronic HCP5 polymorphism (rs3094228), in younger onset GD. This polymorphism has been demonstrated to be associated in a dose-dependent manner, where the greater the number of HCP5 risk alleles, the earlier the onset of GD [64, 65]. Despite these findings, the mechanistic link between these genes and younger onset GD remains to be established.
Other clinical features have been associated with genotype including an increased risk for, and severity of GO (CTLA4, TSHR and TG) [60, 66, 67] and a larger goitre size (CTLA4, CD40, HLA, PTPN22, and TSHR) [60, 63, 68, 69]. Polymorphisms in these genes have also been associated with a more severe biochemical phenotype, reflected by higher serum triiodothyronine (21.5 ± 15.37 pmol/l) (HLA DQA1*05)[70], thyroxine (≥40 pmol/L) (PTPN22)[63] and/or thyroid stimulating hormone antibody (TRAb) levels (CTLA4, CD40, HLA, PTPN22, TG, and TSHR) [60, 67,68,69]. The risk allele (G) and genotype (GG) of the CTLA4 polymorphism, rs231775, have also been associated with the presence of persistently elevated TRAb levels after 5 years of ATD [71], suggesting that these patients may require a longer course of ATD or an alternative therapeutic approach.
8 Genetic variants and outcome in Graves’ disease
Several of the genetic variants associated with particular clinical/biochemical phenotypes have also been independently associated with an increased risk of GD reoccurrence, albeit inconsistently across different study populations [63, 72, 73]. A Taiwanese study including 262 GD patients demonstrated that the risk allele (G) and genotype (GG) of the CTLA4 polymorphism, rs231775, was associated with an increased risk of GD relapse 3 years after ATD withdrawal (relapse cases: 67% (GG) vs. 28% (AG) vs. 5% (AA)), with an odds ratio of 1.96 for the GG genotype compared with the combined group of GA plus AA genotypes [69]. This study also demonstrated association at three CD40 polymorphisms (rs745307, rs11569309, rs3765457) with relapse of GD, with odds ratios of 7.9, 8 and 2.6 for risk vs. protective genotype, respectively [69].
Both CTLA4 and PTPN22 have key roles in regulating the T cell response, with both acting as negative regulators resulting in the inhibition of T cell activation [26, 29, 74,75,76]. Therefore, polymorphisms that decrease expression of CTLA4 and PTPN22 could potentially result in T cell proliferation and subsequent relapse. Indeed, the specific CTLA4 (rs231775) and PTPN22 (rs2476601) polymorphisms associated with GD relapse, have been associated with increased T cell activation and proliferation [74], and a decreased frequency of the tolerogenic T regulatory cells [60, 77].
Conversely, CD40 which is a key co-stimulatory molecule expressed on antigen-presenting cells, drives T cell-dependent B cell activation [78]. Therefore, polymorphisms that may result in the increased expression of CD40 could potentially drive GD relapse through enhancing the humoral response. Indeed, haplotypes that include the CD40 polymorphisms associated with outcome in GD have been associated with CD40 mRNA expression levels [79]. A precariously small study of 13 GD patients suggested that response to the novel therapeutic anti-CD40 monoclonal antibody, Iscalimab, might be predicted by haplotype [79]. Additionally, certain HLA haplotypes conferring susceptibility to GD and associated with disease outcome have been the target for novel therapeutics [63, 70, 80].
Variants in two thyroid specific genes, TSHR and TG, have been associated with outcome in GD. An Indonesian study of 144 GD patients demonstrated that those with the CC genotype of the intronic TSHR polymorphism, rs2268458, had a 13.3 times higher risk of relapse than those with the TT genotype [60]. The TG gene encodes the protein thyroglobulin which has a key role in thyroid hormone production and serum levels are often elevated in GD, where they have been associated with an increased risk of relapse [81]. The genetic variants in thyroid-specific genes may interact with the non-specific immune-related genes to work synergistically and influence the outcome in GD [82].
Identifying genetic variants that are associated with both clinical phenotype and prognosis may be clinically valuable to recognise patients at higher risk of relapse, and may help elucidate mechanistic insight into the functional impacts of genetic variation in GD.
9 Predictive model
A scoring model developed for predicting relapse of GD, ‘GREAT’ (Graves’ Recurrent Events After Therapy), uses specific clinical parameters (age, serum TRAb, goitre size) to stratify GD patients into different classes of recurrence risk. Despite the inconsistencies with genotype-phenotype correlations, the addition of genetic risk alleles at HLA (DRB1-03, DQA1-05, DQB1-02) and PTPN22 (rs2476601) to form the ‘GREAT+’ score, demonstrated an improved predictive value for relapse when compared to clinical factors alone (Fig. 2) [63]. The greatest benefit of using the GREAT+ score was described in those with a moderate risk of GD relapse (GREAT score class II), in whom the addition of genotyping significantly changed the recurrence risk (a change of approximately 25% was considered clinically relevant) and hence potentially the therapeutic approach used in 37 of 98 patients (38%) [63].
10 Antithyroid drug and agranulocytosis
Antithyroid drugs are currently the mainstay of treatment in GD, but the rare side effect of agranulocytosis can be fatal. Studies have identified HLA-B*38:02 and HLA-DRB1*08:03 alleles as independent susceptibility loci for agranulocytosis with antithyroid drugs [83]. This may have clinical implications when deciding the best treatment approach for individuals at greater risk.
11 Summary
Genotyping of GD patients may have significant translational potential by enabling a personalised treatment plan and the future implementation of novel immunomodulatory therapies for appropriate patients. However, the functional effects of these genetic variants in influencing prognosis remain largely unknown and the inconsistencies with genotype-phenotype correlations require larger GWAS studies to validate the candidate gene-association findings.
12 Overall conclusion
Numerous genetic variants that predispose to Graves’ disease have now been identified in robust studies in the Chinese population, as well as in smaller cohorts of people of white European ancestry. Inheritance appears to be truly polygenic with upwards of 80 loci identified, most of which contribute a small increased risk of disease typically around a 10-20% change in risk. Several studies have already correlated clinical features such as relapse following ATD cessation or age of onset of hyperthyroidism with specific genetic variants but much larger cohorts of patients followed for longer periods of time are now needed to determine more detailed phenotypic associations with known and unknown genetic variants that could prove clinically helpful for a precision medicine approach to GD management.
Data availability statement
No original or unpublished data are contained in this paper.
Abbreviations
- ADCY7 :
-
Adenylyl Cyclase Type 7
- AITD:
-
Autoimmune Thyroid Disease
- ATD :
-
Anti-Thyroid Drugs
- BACH2 :
-
Broad Complex-Tramtrack-Bric a Brac and Cap'n'collar Homology 2
- C1QTNF6 :
-
Complement C1q Tumour Necrosis Factor-Related Protein 6
- CD40 :
-
Cluster of Differentiation 40
- Csk:
-
C-terminal Src kinase
- CTLA4:
-
Cytotoxic T-lymphocyte Antigen-4
- FCRL3 :
-
Fc Receptor-Like Protein 3
- FLT3 :
-
Fms Related Receptor Tyrosine Kinase 3
- FT3:
-
Free Trioiodothyronine
- FT4:
-
Free Thyroxine
- GD:
-
Graves’ disease
- GM-CSF:
-
Granulocyte-Macrophage Colony-Stimulating Factor
- GO:
-
Graves' Ophthalmopathy
- GPR174 :
-
G Protein-Coupled Receptor 174
- GREAT:
-
Graves’ Recurrent Events After Therapy
- GWAS:
-
Genome-Wide Association Study
- HCP5 :
-
HLA Complex P5
- HLA:
-
Human Leukocyte Antigen
- IL:
-
Interleukin
- LYP:
-
Lymphoid Tyrosine Phosphatase
- MHC:
-
Major Histocompatibility Complex
- MHC-DRβ:
-
Major Histocompatibility Complex - DRβ isotype
- mRNA:
-
Messenger Ribonucleic Acid
- PTPN22:
-
Protein Tyrosine Phosphatase Non-receptor type 22
- RAC2:
-
Ras-related C3 botulinum toxin substrate 2
- RNASET2 :
-
Ribonuclease T2
- SCGB3A2:
-
Secretoglobin Family 3A Member 2
- SH2B3 :
-
SH2B adaptor protein 3
- SNP:
-
Single-Nucleotide Polymorphism
- T3:
-
Triiodothyronine
- T4:
-
Thyroxine
- TCR:
-
T cell Receptor
- TED:
-
Thyroid eye disease
- TG:
-
Thyroglobulin
- Treg:
-
Regulatory T cells
- TSH:
-
Thyroid Stimulating Hormone
- TSHR:
-
Thyroid Stimulating Hormone Receptor
- UGRP1:
-
Uterus Globulin Associated Protein-1
- UK:
-
United Kingdom
- US:
-
United States
- WTCCC:
-
Wellcome Trust Case Control Consortium
- λs:
-
sibling recurrence risk ratio
- +ve:
-
positive
- -ve:
-
negative
References
Nyström HF, Jansson S, Berg G. Incidence rate and clinical features of hyperthyroidism in a long-term iodine sufficient area of Sweden (Gothenburg) 2003–2005. Clinical Endocrinology. 2013;78(5):768–76.
Holm IA, Manson JE, Michels KB, Alexander EK, Willett WC, Utiger RD. Smoking and other lifestyle factors and the risk of Graves’ hyperthyroidism. Arch Intern Med. 2005;165(14):1606–11. https://doi.org/10.1001/archinte.165.14.1606. PMID: 16043678.
Conrad N, Misra S, Verbakel JY, Verbeke G, Molenberghs G, Taylor PN, Mason J, Sattar N, McMurray JJV, McInnes IB, Khunti K, Cambridge G. Incidence, prevalence, and co-occurrence of autoimmune disorders over time and by age, sex, and socioeconomic status: a population-based cohort study of 22 million individuals in the UK. Lancet. 2023;S0140–6736(23):00457–9. https://doi.org/10.1016/S0140-6736(23)00457-9. PMID: 37156255.
McLeod DS, Caturegli P, Cooper DS, Matos PG, Hutfless S. Variation in rates of autoimmune thyroid disease by race/ethnicity in US military personnel. JAMA. 2014;311(15):1563–5. https://doi.org/10.1001/jama.2013.285606. PMID: 24737370.
Brix TH, Kyvik KO, Christensen K, Hegedüs L. Evidence for a major role of heredity in Graves’ disease: A population-based study of two Danish twin cohorts. J Clin Endocrinol Metabol. 2001;86(2):930–4.
Skov J, Eriksson D, Kuja-Halkola R, Höijer J, Gudbjörnsdottir S, Svensson AM, Magnusson PKE, Ludvigsson JF, Kämpe O, Bensing S. Co-aggregation and heritability of organ-specific autoimmunity: a population-based twin study. Eur J Endocrinol. 2020;182(5):473–80. https://doi.org/10.1530/EJE-20-0049. 32229696.
Stenszky V, Kozma L, Balázs C, Rochlitz S, Bear JC, Farid NR. The genetics of Graves’ disease: HLA and disease susceptibility. J Clin Endocrinol Metab. 1985;61(4):735–40. https://doi.org/10.1210/jcem-61-4-735. PMID: 3861611.
Vaidya B, Kendall-Taylor P, Pearce SH. The genetics of autoimmune thyroid disease. J Clin Endocrinol Metab. 2002;87(12):5385–97. https://doi.org/10.1210/jc.2002-020492. PMID: 12466323.
Vyse TJ, Todd JA. Genetic analysis of autoimmune disease. Cell. 1996;85(3):311–8. https://doi.org/10.1016/s0092-8674(00)81110-1. PMID: 8616887.
Strieder TGA, Tijssen JGP, Wenzel BE, Endert E, Wiersinga WM. Prediction of progression to overt hypothyroidism or hyperthyroidism in female relatives of patients with autoimmune thyroid disease using the thyroid events Amsterdam (THEA) score. Arch Internal Med. 2008;168(15):1657–63.
Song H, Fang F, Tomasson G, Arnberg FK, Mataix-Cols D, Fernández de la Cruz L, Almqvist C, Fall K, Valdimarsdóttir UA. Association of Stress-Related Disorders With Subsequent Autoimmune Disease. JAMA 2018;319(23):2388-2400.
Vestergaard P. Smoking and thyroid disorders–a meta-analysis. Eur J Endocrinol. 2002;146(2):153–61. https://doi.org/10.1530/eje.0.1460153. PMID: 11834423.
Bülow Pedersen I, Knudsen N, Jørgensen T, Perrild H, Ovesen L, Laurberg P. Large differences in incidences of overt hyper- and hypothyroidism associated with a small difference in iodine intake: a prospective comparative register-based population survey. J Clin Endocrinol Metab. 2002;87(10):4462–9. https://doi.org/10.1210/jc.2002-020750. PMID: 12364419.
Limbach M, Saare M, Tserel L, Kisand K, Eglit T, Sauer S, et al. Epigenetic profiling in CD4+ and CD8+ T cells from Graves’ disease patients reveals changes in genes associated with T cell receptor signaling. J Autoimmun. 2016;67:46–56.
Cai TT, Muhali FS, Song RH, Qin Q, Wang X, Shi LF, et al. Genome-wide DNA methylation analysis in Graves’ disease. Genomics. 2015;105:204–10. https://doi.org/10.1016/j.ygeno.2015.01.001.
Xin Z, Hua L, Shi TT, Tuo X, Yang FY, Li Y, Cao X, Yang JK. A genome-wide DNA methylation analysis in peripheral blood from patients identifies risk loci associated with Graves' orbitopathy. J Endocrinol Invest 2018;41(6):719-727. https://doi.org/10.1007/s40618-017-0796-6. PMID: 29190000.
Shi TT, Hua L, Xin Z, Li Y, Liu W, Yang YL. Identifying and Validating Genes with DNA Methylation Data in the Context of Biological Network for Chinese Patients with Graves' Orbitopathy. Int J Endocrinol 2019;2019:6212681. https://doi.org/10.1155/2019/6212681. PMID: 31001336.
Grumet FC, Payne RO, Kinishi J, Kriss JP. HL A antigens as markers for disease susceptibility and autoimmunity in Grave’s disease. Journal of Clinical Endocrinology and Metabolism. 1974;39(6):1115–9.
Yanagawa T, Mangklabruks A, Chang YB, Okamoto Y, Fisfalen ME, Curran PG, et al. Human histocompatibility leukocyte antigen-DQA1*0501 allele associated with genetic susceptibility to graves’ disease in a caucasian population. Journal of Clinical Endocrinology and Metabolism. 1993;76(6):1569–74.
Heward JM, Allahabadia A, Daykin J, Carr-Smith J, Daly A, Armitage M, et al. Linkage disequilibrium between the human leukocyte antigen class II region of the major histocompatibility complex and Graves’ disease: Replication using a population case control and family-based study. Journal of Clinical Endocrinology and Metabolism. 1998;83(10):3394–7.
Katahira M, Ogata H, Takashima H, Ito T, Hodai Y, Miwata T, Goto M, Yamaguchi M, Mizoguchi A, Kawakubo M, Nakamura S. Critical amino acid variants in HLA-DRB1 allotypes in the development of Graves’ disease and Hashimoto’s thyroiditis in the Japanese population. Hum Immunol. 2021;82(4):226–31.
Ueda S, Oryoji D, Yamamoto K, Noh JY, Okamura K, Noda M, Kashiwase K, Kosuga Y, Sekiya K, Inoue K, Yamada H, Oyamada A, Nishimura Y, Yoshikai Y, Ito K, Sasazuki T. Identification of independent susceptible and protective HLA alleles in Japanese autoimmune thyroid disease and their epistasis. J Clin Endocrinol Metab. 2014;99(2):E379-83.
Park MH, Park YJ, Song EY, Park H, Kim TY, Park DJ, Park KS, Cho BY. Association of HLA-DR and -DQ genes with Graves disease in Koreans. Hum Immunol. 2005;66(6):741–7.
Wongsurawat T, Nakkuntod J, Charoenwongse P, Snabboon T, Sridama V, Hirankarn N. The association between HLA class II haplotype with Graves’ disease in Thai population. Tissue Antigens. 2006;67(1):79–83.
Ban Y, Davies TF, Greenberg DA, Concepcion ES, Osman R, Oashi T, Tomer Y. Arginine at position 74 of the HLA-DR beta1 chain is associated with Graves’ disease. Genes Immun. 2004;5(3):203–8.
Yanagawa T, Hidaka Y, Guimaraes V, Soliman M, DeGroot LJ. CTLA-4 gene polymorphism associated with Graves’ disease in a Caucasian population. J Clin Endocrinol Metab. 1995;80(1):41–5.
Kavvoura FK, Akamizu T, Awata T, Ban Y, Chistiakov DA, Frydecka I, Ghaderi A, Gough SC, Hiromatsu Y, Ploski R, Wang PW, Ban Y, Bednarczuk T, Chistiakova EI, Chojm M, Heward JM, Hiratani H, Juo SH, Karabon L, Katayama S, Kurihara S, Liu RT, Miyake I, Omrani GH, Pawlak E, Taniyama M, Tozaki T, Ioannidis JP. Cytotoxic T-lymphocyte associated antigen 4 gene polymorphisms and autoimmune thyroid disease: a meta-analysis. J Clin Endocrinol Metab. 2007;92(8):3162–70.
Vaidya B, Imrie H, Perros P, Dickinson J, McCarthy MI, Kendall-Taylor P, Pearce SH. Cytotoxic T lymphocyte antigen-4 (CTLA-4) gene polymorphism confers susceptibility to thyroid associated orbitopathy. Lancet. 1999;354(9180):743–4.
Bottini N, Musumeci L, Alonso A, Rahmouni S, Nika K, Rostamkhani M, MacMurray J, Meloni GF, Lucarelli P, Pellecchia M, Eisenbarth GS, Comings D, Mustelin T. A functional variant of lymphoid tyrosine phosphatase is associated with type I diabetes. Nat Genet. 2004;36(4):337–8.
Velaga MR, Wilson V, Jennings CE, Owen CJ, Herington S, Donaldson PT, Ball SG, James RA, Quinton R, Perros P, Pearce SH. The codon 620 tryptophan allele of the lymphoid tyrosine phosphatase (LYP) gene is a major determinant of Graves’ disease. J Clin Endocrinol Metab. 2004;89(11):5862–5.
Smyth D, Cooper JD, Collins JE, Heward JM, Franklyn JA, Howson JM, Vella A, Nutland S, Rance HE, Maier L, Barratt BJ, Guja C, Ionescu-Tîrgoviste C, Savage DA, Dunger DB, Widmer B, Strachan DP, Ring SM, Walker N, Clayton DG, Twells RC, Gough SC, Todd JA. Replication of an association between the lymphoid tyrosine phosphatase locus (LYP/PTPN22) with type 1 diabetes, and evidence for its role as a general autoimmunity locus. Diabetes. 2004;53(11):3020–3.
Tomer Y, Barbesino G, Greenberg DA, Concepcion ES, Davies TF. Mapping the major susceptibility loci for familial Graves’ and Hashimoto’s diseases: evidence for genetic heterogeneity and gene interactions. J Clin Endocrinol Metab. 1999;84:4656–64.
Taylor JC, Gough SC, Hunt PJ, Brix TH, Chatterjee K, Connell JM, Franklyn JA, Hegedus L, Robinson BG, Wiersinga WM, Wass JA, Zabaneh D, Mackay I, Weetman AP. A genome-wide screen in 1119 relative pairs with autoimmune thyroid disease. J Clin Endocrinol Metab. 2006;91(2):646–53. https://doi.org/10.1210/jc.2005-0686. Epub 2005 Nov 8 PMID: 16278270.
Vaidya B, Imrie H, Perros P, Young ET, Kelly WF, Carr D, Large DM, Toft AD, McCarthy MI, Kendall-Taylor P, Pearce SH. The cytotoxic T lymphocyte antigen-4 is a major Graves’ disease locus. Hum Mol Genet. 1999;8(7):1195–9. https://doi.org/10.1093/hmg/8.7.1195. PMID: 10369864.
Tomer Y, Ban Y, Concepcion E, Barbesino G, Villanueva R, Greenberg DA, Davies TF. Common and unique susceptibility loci in Graves and Hashimoto diseases: results of whole-genome screening in a data set of 102 multiplex families. Am J Hum Genet. 2003;73(4):736-47. https://doi.org/10.1086/378588. Epub 2003 Sep 12. PMID: 12973666; PMCID: PMC1180598.
Dechairo BM, Zabaneh D, Collins J, Brand O, Dawson GJ, Green AP, et al. Association of the TSHR gene with Graves’ disease: The first disease specific locus. European Journal of Human Genetics. 2005;13(11):1223–30.
Płoski R, Brand OJ, Jurecka-Lubieniecka B, Franaszczyk M, Kula D, Krajewski P, Karamat MA, Simmonds MJ, Franklyn JA, Gough SC, Jarząb B, Bednarczuk T. Thyroid stimulating hormone receptor (TSHR) intron 1 variants are major risk factors for Graves’ disease in three European Caucasian cohorts. PLoS One. 2010;5(11): e15512.
Colobran R, Armengol Mdel P, Faner R, Gärtner M, Tykocinski LO, Lucas A, Ruiz M, Juan M, Kyewski B, Pujol-Borrell R. Association of an SNP with intrathymic transcription of TSHR and Graves’ disease: a role for defective thymic tolerance. Hum Mol Genet. 2011;20(17):3415–23.
Pujol-Borrell R, Álvarez-Sierra D, Jaraquemada D, Marín-Sánchez A, Colobran R. Central Tolerance Mechanisms to TSHR in Graves’ Disease: Contributions to Understand the Genetic Association. Horm Metab Res. 2018;50(12):863–70.
Wellcome Trust Case Control Consortium; Australo-Anglo-American Spondylitis Consortium (TASC); Burton PR, Clayton DG, Cardon LR, Craddock N, Deloukas P, Duncanson A, Kwiatkowski DP, McCarthy MI, Ouwehand WH, Samani NJ, Todd JA, Donnelly P, Barrett JC, Davison D, Easton D, Evans DM, Leung HT, Marchini JL, Morris AP, Spencer CC, Tobin MD, Attwood AP, Boorman JP, Cant B, Everson U, Hussey JM, Jolley JD, Knight AS, Koch K, Meech E, Nutland S, Prowse CV, Stevens HE, Taylor NC, Walters GR, Walker NM, Watkins NA, Winzer T, Jones RW, McArdle WL, Ring SM, Strachan DP, Pembrey M, Breen G, St Clair D, Caesar S, Gordon-Smith K, Jones L, Fraser C, Green EK, Grozeva D, Hamshere ML, Holmans PA, Jones IR, Kirov G, Moskivina V, Nikolov I, O'Donovan MC, Owen MJ, Collier DA, Elkin A, Farmer A, Williamson R, McGuffin P, Young AH, Ferrier IN, Ball SG, Balmforth AJ, Barrett JH, Bishop TD, Iles MM, Maqbool A, Yuldasheva N, Hall AS, Braund PS, Dixon RJ, Mangino M, Stevens S, Thompson JR, Bredin F, Tremelling M, Parkes M, Drummond H, Lees CW, Nimmo ER, Satsangi J, Fisher SA, Forbes A, Lewis CM, Onnie CM, Prescott NJ, Sanderson J, Matthew CG, Barbour J, Mohiuddin MK, Todhunter CE, Mansfield JC, Ahmad T, Cummings FR, Jewell DP, Webster J, Brown MJ, Lathrop MG, Connell J, Dominiczak A, Marcano CA, Burke B, Dobson R, Gungadoo J, Lee KL, Munroe PB, Newhouse SJ, Onipinla A, Wallace C, Xue M, Caulfield M, Farrall M, Barton A; Biologics in RA Genetics and Genomics Study Syndicate (BRAGGS) Steering Committee; Bruce IN, Donovan H, Eyre S, Gilbert PD, Hilder SL, Hinks AM, John SL, Potter C, Silman AJ, Symmons DP, Thomson W, Worthington J, Dunger DB, Widmer B, Frayling TM, Freathy RM, Lango H, Perry JR, Shields BM, Weedon MN, Hattersley AT, Hitman GA, Walker M, Elliott KS, Groves CJ, Lindgren CM, Rayner NW, Timpson NJ, Zeggini E, Newport M, Sirugo G, Lyons E, Vannberg F, Hill AV, Bradbury LA, Farrar C, Pointon JJ, Wordsworth P, Brown MA, Franklyn JA, Heward JM, Simmonds MJ, Gough SC, Seal S; Breast Cancer Susceptibility Collaboration (UK); Stratton MR, Rahman N, Ban M, Goris A, Sawcer SJ, Compston A, Conway D, Jallow M, Newport M, Sirugo G, Rockett KA, Bumpstead SJ, Chaney A, Downes K, Ghori MJ, Gwilliam R, Hunt SE, Inouye M, Keniry A, King E, McGinnis R, Potter S, Ravindrarajah R, Whittaker P, Widden C, Withers D, Cardin NJ, Davison D, Ferreira T, Pereira-Gale J, Hallgrimsdo'ttir IB, Howie BN, Su Z, Teo YY, Vukcevic D, Bentley D, Brown MA, Compston A, Farrall M, Hall AS, Hattersley AT, Hill AV, Parkes M, Pembrey M, Stratton MR, Mitchell SL, Newby PR, Brand OJ, Carr-Smith J, Pearce SH, McGinnis R, Keniry A, Deloukas P, Reveille JD, Zhou X, Sims AM, Dowling A, Taylor J, Doan T, Davis JC, Savage L, Ward MM, Learch TL, Weisman MH, Brown M. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat Genet. 2007;39(11):1329-37.
Owen CJ, Kelly H, Eden JA, Merriman ME, Pearce SH, Merriman TR. Analysis of the Fc receptor-like-3 (FCRL3) locus in Caucasians with autoimmune disorders suggests a complex pattern of disease association. J Clin Endocrinol Metab. 2007;92(3):1106–11. https://doi.org/10.1210/jc.2006-2183. Epub 2007 Jan 2 PMID: 17200162.
Chistiakov DA, Chistiakov AP. Is FCRL3 a new general autoimmunity gene? Hum Immunol. 2007;68(5):375–83. https://doi.org/10.1016/j.humimm.2007.01.013.
Newby PR, Pickles OJ, Mazumdar S, Brand OJ, Carr-Smith JD, Pearce SH, Franklyn JA; Wellcome Trust Case-Control Consortium (WTCCC); Evans DM, Simmonds MJ, Gough SC. Follow-up of potential novel Graves' disease susceptibility loci, identified in the UK WTCCC genome-wide nonsynonymous SNP study. Eur J Hum Genet. 2010 ;18(9):1021-6. https://doi.org/10.1038/ejhg.2010.55.
Sakai K, Shirasawa S, Ishikawa N, Ito K, Tamai H, Kuma K, Akamizu T, Tanimura M, Furugaki K, Yamamoto K, Sasazuki T. Identification of susceptibility loci for autoimmune thyroid disease to 5q31– q33 and Hashimoto’s thyroiditis to 8q23– q24 by multipoint affected sib-pair linkage analysis in Japanese. Hum Mol Genet. 2001;10:1379–86.
Jin Y, Teng W, Ben S, Xiong X, Zhang J, Xu S, Shugart YY, Jin L, Chen J, Huang W. Genome-wide scan of Graves’ disease: evidence for linkage on chromosome 5q31 in Chinese Han pedigrees. J Clin Endocrinol Metab. 2003;88(4):1798–803. https://doi.org/10.1210/jc.2001-011980. PMID: 12679476.
Song HD, Liang J, Shi JY, Zhao SX, Liu Z, Zhao JJ, Peng YD, Gao GQ, Tao J, Pan CM, Shao L, Cheng F, Wang Y, Yuan GY, Xu C, Han B, Huang W, Chu X, Chen Y, Sheng Y, Li RY, Su Q, Gao L, Jia WP, Jin L, Chen MD, Chen SJ, Chen Z, Chen JL. 2009. Functional SNPs in the SCGB3A2 promoter are associated with susceptibility to Graves' disease. Hum Mol Genet. 2009;18(6):1156-70. https://doi.org/10.1093/hmg/ddn442.
Zhou Z, Zuo CL, Li XS, Ye XP, Zhang QY, Wang P, Zhang RX, Chen G, Yang JL, Chen Y, Ma QY, Song HD. Uterus globulin associated protein 1 (UGRP1) is a potential marker of progression of Graves’ disease into hypothyroidism. Mol Cell Endocrinol. 2019;20(494): 110492. https://doi.org/10.1016/j.mce.2019.110492.
Simmonds MJ, Yesmin K, Newby PR, Brand OJ, Franklyn JA, Gough SC. Confirmation of association of chromosome 5q31-33 with United Kingdom Caucasian Graves’ disease. Thyroid. 2010;20(4):413–7. https://doi.org/10.1089/thy.2009.0375. PMID: 20210668.
Chu X, Pan CM, Zhao SX, Liang J, Gao GQ, Zhang XM, Yuan GY, Li CG, Xue LQ, Shen M, Liu W, Xie F, Yang SY, Wang HF, Shi JY, Sun WW, Du WH, Zuo CL, Shi JX, Liu BL, Guo CC, Zhan M, Gu ZH, Zhang XN, Sun F, Wang ZQ, Song ZY, Zou CY, Sun WH, Guo T, Cao HM, Ma JH, Han B, Li P, Jiang H, Huang QH, Liang L, Liu LB, Chen G, Su Q, Peng YD, Zhao JJ, Ning G, Chen Z, Chen JL, Chen SJ, Huang W, Song HD; China Consortium for Genetics of Autoimmune Thyroid Disease. A genome-wide association study identifies two new risk loci for Graves' disease. Nat Genet. 2011;43(9):897-901. https://doi.org/10.1038/ng.898.
Zhao SX, Xue LQ, Liu W, Gu ZH, Pan CM, Yang SY, Zhan M, Wang HN, Liang J, Gao GQ, Zhang XM, Yuan GY, Li CG, Du WH, Liu BL, Liu LB, Chen G, Su Q, Peng YD, Zhao JJ, Ning G, Huang W, Liang L, Qi L, Chen SJ, Chen Z, Chen JL, Song HD; China Consortium for the Genetics of Autoimmune Thyroid Disease. Robust evidence for five new Graves' disease risk loci from a staged genome-wide association analysis. Hum Mol Genet. 2013;22(16):3347-62. https://doi.org/10.1093/hmg/ddt183.
Gu LQ, Zhu W, Zhao SX, Zhao L, Zhang MJ, Cui B, Song HD, Ning G, Zhao YJ. Clinical associations of the genetic variants of CTLA-4, Tg, TSHR, PTPN22, PTPN12 and FCRL3 in patients with Graves’ disease. Clin Endocrinol (Oxf). 2010;72(2):248–55. https://doi.org/10.1111/j.1365-2265.2009.03617.x.
Liu W, Wang HN, Gu ZH, Yang SY, Ye XP, Pan CM, Zhao SX, Xue LQ, Xie HJ, Yu SS, Guo CC, Du WH, Liang J, Zhang XM, Yuan GY, Li CG, Su Q, Gao GQ, Song HD; China Consortium for the Genetics of Autoimmune Thyroid Disease. Identification of BACH2 as a susceptibility gene for Graves' disease in the Chinese Han population based on a three-stage genome-wide association study. Hum Genet. 2014;133(5):661-71. https://doi.org/10.1007/s00439-013-1404-2.
Cooper JD, Simmonds MJ, Walker NM, Burren O, Brand OJ, Guo H, Wallace C, Stevens H, Coleman G; Wellcome Trust Case Control Consortium; Franklyn JA, Todd JA, Gough SC. Seven newly identified loci for autoimmune thyroid disease. Hum Mol Genet. 2012;21(23):5202-8. https://doi.org/10.1093/hmg/dds357.
Muto A, Tashiro S, Nakajima O, Hoshino H, Takahashi S, Sakoda E, Ikebe D, Yamamoto M, Igarashi K. The transcriptional programme of antibody class switching involves the repressor Bach2. Nature. 2004;429(6991):566–71. https://doi.org/10.1038/nature02596.
Pazderska A, Oftedal BE, Napier CM, Ainsworth HF, Husebye ES, Cordell HJ, Pearce SH, Mitchell AL. A Variant in the BACH2 Gene Is Associated With Susceptibility to Autoimmune Addison’s Disease in Humans. J Clin Endocrinol Metab. 2016;101(11):3865–9. https://doi.org/10.1210/jc.2016-2368.
McAllister K, Yarwood A, Bowes J, Orozco G, Viatte S, Diogo D, Hocking LJ, Steer S, Wordsworth P, Wilson AG, Morgan AW. UK Rheumatoid Arthritis Genetics Consortium; Rheumatoid Arthritis Consortium International; Identification of BACH2 and RAD51B as rheumatoid arthritis susceptibility loci in a meta-analysis of genome-wide data. Arthritis Rheum. 2013;65(12):3058–62. https://doi.org/10.1002/art.38183.
Cooper JD, Smyth DJ, Smiles AM, Plagnol V, Walker NM, Allen JE, Downes K, Barrett JC, Healy BC, Mychaleckyj JC, Warram JH, Todd JA. Meta-analysis of genome-wide association study data identifies additional type 1 diabetes risk loci. Nat Genet. 2008;40(12):1399-401. https://doi.org/10.1038/ng.249. Epub 2008 Nov 2. PMID: 18978792; PMCID: PMC2635556.
Saevarsdottir S, Olafsdottir TA, Ivarsdottir EV, et al. FLT3 stop mutation increases FLT3 ligand level and risk of autoimmune thyroid disease. Nature. 2020;584:619–23. https://doi.org/10.1038/s41586-020-2436-0.
Wang K, Zhu Q, Lu Y, Lu H, Zhang F, Wang X, Fan Y. CTLA-4 +49 G/A Polymorphism Confers Autoimmune Disease Risk: An Updated Meta-Analysis. Genet Test Mol Biomarkers. 2017;21(4):222–7.
Eliana F, Suwondo P, Asmarinah A, et al. The Role of Cytotoxic T-lymphocyte-associated Protein 4 (CTLA-4) Gene, Thyroid Stimulating Hormone Receptor (TSHR) Gene and Regulatory T-cells as Risk Factors for Relapse in Patients with Graves Disease. Acta Med Indones. 2017;49(3):195–204.
Tanrikulu S, Erbil Y, Ademoglu E, et al. The predictive value of CTLA-4 and Tg polymorphisms in the recurrence of Graves’ disease after antithyroid withdrawal. Endocrine. 2006;30(3):377–81.
Sahin M, Erdogan MF, Erdogan G. Cytotoxic T lymphocyte-associated molecule-4 polymorphisms in Turkish Graves’ disease patients and association with probability of remission after antithyroid therapy. Eur J Intern Med. 2005;16(5):352–5.
Vos XG, Wiersinga WM, Endert E, Zwinderman AH, Tijssen JGP. Predicting the Risk of Recurrence Before the Start of Antithyroid Drug Therapy in Patients With Graves’ Hyperthyroidism. The Journal of Clinical Endocrinology & Metabolism. 2016;101(4):1381–9.
Kuś A, Radziszewski M, Glina A, Szymański K, Jurecka-Lubieniecka B, Pawlak-Adamska E, Kula D, Wawrusiewicz-Kurylonek N, Kuś J, Miśkiewicz P, Płoski R, Bolanowski M, Daroszewski J, Jarząb B, Bossowski A, Bednarczuk T. Paediatric-onset and adult-onset Graves’ disease share multiple genetic risk factors. Clin Endocrinol (Oxf). 2019;90(2):320–7.
Lane LC, Kuś A, Bednarczuk T, Bossowski A, Daroszewski J, Jurecka-Lubieniecka B, Cordell HJ, Pearce SHS, Cheetham T, Mitchell AL. An Intronic HCP5 Variant Is Associated With Age of Onset and Susceptibility to Graves Disease in UK and Polish Cohorts. J Clin Endocrinol Metab. 2020;105(9):e3277-84.
García-Mayor RV, Álvarez-Vázquez P, Fluiters E, Valverde D, Andrade A. Long-term remission following antithyroid drug withdrawal in patients with Graves’ hyperthyroidism: parameters with prognostic value. Endocrine. 2019;63(2):316–22.
Hsiao JY, Hsieh MC, Tien KJ, Hsu SC, Shin SJ, Lin SR. Association between a C/T polymorphism in exon 33 of the thyroglobulin gene is associated with relapse of Graves’ hyperthyroidism after antithyroid withdrawal in Taiwanese. J Clin Endocrinol Metab. 2007;92(8):3197–201.
Wang PW, Chen IY, Liu RT, Hsieh CJ, Hsi E, Juo SH. CTLA-4 gene polymorphism and hyperthyroid Graves’ disease relapse after antithyroid drug withdrawal: a follow-up study. J Clin Endocrinol Metab. 2007;92:2513–8.
Wang PW, Chen IY, Juo SH, Hsi E, Liu RT, Hsieh CJ. Genotype and phenotype predictors of relapse of graves’ disease after antithyroid drug withdrawal. Eur Thyroid J. 2013;1(4):251–8.
Vejrazkova D, Vcelak J, Vaclavikova E, Vankova M, Zajickova K, Vrbikova J, Duskova M, Pacesova P, Novak Z, Bendlova B. Recurrence of Graves’ Disease: What Genetics of HLA and PTPN22 Can Tell Us. Front Endocrinol (Lausanne). 2021;23(12): 761077.
Kinjo Y, Takasu N, Komiya I, et al. Remission of Graves’ hyperthyroidism and A/G polymorphism at position 49 in exon 1 of cytotoxic T lymphocyte-associated molecule-4 gene. J Clin Endocrinol Metab. 2002;87(6):2593–6.
Kim KW, Park YJ, Kim TY, Park DJ, Park KS, Cho BY. Susceptible alleles of the CD40 and CTLA-4 genes are not associated with the relapse after antithyroid withdrawal in Graves’ disease. Thyroid. 2007;17(12):1229–34.
Vejrazkova D, Vcelak J, Vaclavikova E, Vankova M, Zajickova K, Vrbikova J, Duskova M, Pacesova P, Novak Z, Bendlova B. Recurrence of Graves’ Disease: What Genetics of HLA and PTPN22 Can Tell Us. Front Endocrinol (Lausanne). 2021;23(12): 761077.
Allison JP, Krummel MF. The Yin and Yang of T cell costimulation. Science. 1995;270(5238):932–3.
Vang T, Liu WH, Delacroix L, Wu S, Vasile S, Dahl R, Yang L, Musumeci L, Francis D, Landskron J, Tasken K, Tremblay ML, Lie BA, Page R, Mustelin T, Rahmouni S, Rickert RC, Tautz L. LYP inhibits T-cell activation when dissociated from CSK. Nat Chem Biol. 2012;8(5):437–46.
Ban Y, Davies TF, Greenberg DA, Kissin A, Marder B, Murphy B, Concepcion ES, Villanueva RB, Barbesino G, Ling V, Tomer Y. Analysis of the CTLA-4, CD28, and inducible costimulator (ICOS) genes in autoimmune thyroid disease. Genes Immun. 2003;4(8):586–93.
Valta M, Gazali AM, Viisanen T, et al. Type 1 diabetes linked PTPN22 gene polymorphism is associated with the frequency of circulating regulatory T cells. Eur J Immunol. 2020;50(4):581–8.
Elgueta R, Benson MJ, de Vries VC, Wasiuk A, Guo Y, Noelle RJ. Molecular mechanism and function of CD40/CD40L engagement in the immune system. Immunol Rev. 2009;229(1):152–72.
Faustino LC, Kahaly GJ, Frommer L, Concepcion E, Stefan-Lifshitz M, Tomer Y. Precision Medicine in Graves’ Disease: CD40 Gene Variants Predict Clinical Response to an Anti-CD40 Monoclonal Antibody. Front Endocrinol (Lausanne). 2021;4(12): 691781.
Li CW, Osman R, Menconi F, Concepcion E, Tomer Y. Cepharanthine blocks TSH receptor peptide presentation by HLA-DR3: Therapeutic implications to Graves’ disease. J Autoimmun. 2020;108: 102402.
Gong ST, Chao IM. Changes in serum thyroglobulin and thyroid autoantibodies in patients with Graves’ disease treated with antithyroid drug and their relationship to relapse. J Formos Med Assoc. 1991;90(12):1155–62.
Simmonds MJ, Howson JM, Heward JM, et al. Regression mapping of association between the human leukocyte antigen region and Graves disease. Am J Hum Genet. 2005;76(1):157–63.
Chen PL, Shih SR, Wang PW, Lin YC, Chu CC, Lin JH, Chen SC, Chang CC, Huang TS, Tsai KS, Tseng FY, Wang CY, Lu JY, Chiu WY, Chang CC, Chen YH, Chen YT, Fann CS, Yang WS, Chang TC. Genetic determinants of antithyroid drug-induced agranulocytosis by human leukocyte antigen genotyping and genome-wide association study. Nat Commun. 2015;7(6):7633.
Funding
The work of LCL and SHP has been funded by UKRI-MRC grants MR/S001611/1 and MR/V005898/1. Medical Research Council, MR/S001611/1, MR/V005898/1.
Author information
Authors and Affiliations
Contributions
SHP conceived the idea for the review; each author drafted sections of the review and revised the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
N/A.
Informed consent
N/A.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Grixti, L., Lane, L.C. & Pearce, S.H. The genetics of Graves’ disease. Rev Endocr Metab Disord 25, 203–214 (2024). https://doi.org/10.1007/s11154-023-09848-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11154-023-09848-8