Abstract
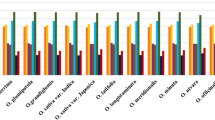
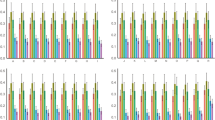
Codon usage bias (CUB) is an important evolutionary feature in a genome and has been widely documented from prokaryotes to eukaryotes. However, the significance of CUB in the Asteraceae family has not been well understood, with no Asteraceae species having been analyzed for this characteristic. Here, we use bioinformatics approaches to comparatively analyze the general patterns and influencing factors of CUB in five Asteraceae chloroplast (cp) genomes. The results indicated that the five genomes had similar codon usage patterns, showing a strong bias towards a high representation of NNA and NNT codons. Neutrality analysis showed that these cp genomes had a narrow GC distribution and no significant correlation was observed between GC12 and GC3. Parity Rule 2 (PR2) plot analysis revealed that purines were used more frequently than pyrimidines. Effective number of codons (ENc)-plot analysis showed that most genes followed the parabolic line of trajectory, but several genes with low ENc values lying below the expected curve were also observed. Furthermore, correspondence analysis of relative synonymous codon usage (RSCU) yielded a first axis that explained only a partial amount of variation of codon usage. These findings suggested that both natural selection and mutational bias contributed to codon bias, while selection was the major force to shape the codon usage in these Asteraceae cp genomes. Our study, which is the first to investigate codon usage patterns in Asteraceae plastomes, will provide helpful information about codon distribution and variation in these species, and also shed light on the genetic and evolutionary mechanisms of codon biology within this family.





Similar content being viewed by others
References
Ahmad T, Sablok G, Tatarinova TV, Xu Q, Deng XX, Guo WW (2013)Evaluation of codon biology in citrus and Poncirus trifoliata based on genomic features and frame corrected expressed sequence tag. DNA Res 20:135–150
Angellotti MC, Bhuiyan SB, Chen GR, Wan XF (2007) CodonO: codon usage bias analysis within and across genomes. Nucleic Acids Res 35:W132–W136
Batzman M, Margalit H (2011) Variation in global codon usage bias among prokaryotic organisms is associated with their lifestyles. Genome Biol 12:R109
Blake WJ, Kaern M, Cantor CR, Collins JJ (2003) Noise in eukaryotic gene expression. Nature 422:633–637
Bremer K (1994) Asteraceae: cladistics and classification. Timber, Oregon
Cosmi CC, Ragosta V, Macchiato MF (1990) Characterization of nucleotide sequences using maximum entropy techniques. J Theor Biol 147:423–432
Duret L, Mouchiroud D (1999) Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila, and Arabidopsis. Proc Natl Acad Sci USA 96:4482–4487
Fu CJ, Xiong J, Miao W (2009) Genome-wide identification and characterization of cytochrome P450 monooxygenase genes in the ciliate Tetrahymena thermophila. BMC Genomics 10:208
Grantham R, Gautier C, Gouy M (1980) Codon frequencies in 119 individual genes confirm consistent choices of degenerate bases according to genome type. Nucleic Acids Res 8:1893–1912
Greenacre MJ (1984) Theory and applications of correspondence analysis. Academic, London
Grigoriev A (1999) Strand-specific compositional asymmetries in double-stranded DNA viruses. Virus Res 60:1–19
Gu W, Zhou T, Ma J, Sun X, Lu Z (2004) The relationship between synonymous codon usage and protein structure in Escherichia coli and Homo sapiens. Biosystems 73:89–97
Gupta SK, Bhattacharyya TK, Ghosh TC (2004) Synonymous codon usage in Lactococcus lactis: mutational bias versus translational selection. J Biomol Struct Dyn 21:527–536
Hershberg R, Petrov DA (2009) General rules for optimal codon choice. PLoS Genet 5(7):e1000556
Hou ZC, Yang N (2002) Analysis of factors shaping S. pneumoniae codon usage (In Chinese with English Abstract). Acta Genet Sin 29:747–752
Ikemura T (1985) Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol 2:13–34
Ingvarsson PK (2007) Gene expression and protein length influence codon usage and rates of sequence evolution in Populus tremula. Mol Biol Evol 24:836–844
Kawabe A, Miyashita NT (2003) Patterns of codon usage bias in three dicot and four monocot plant species. Genes Genet Syst 78:343–352
Liu QP (2006) Analysis of codon usage pattern in the radioresistant bacterium Deinococcus radiodurans. Biosystems 85:99–106
Liu Q, Xue Q (2005) Comparative studies on codon usage pattern of chloroplasts and their host nuclear genes in four plant species. J Genet 84:55–62
Lundberg J, Bremer K (2003) A phylogenetic study of the order Asterales using one morphological and three molecular data sets. Int J Plant Sci 164:553–578
Lynn DJ, Singer GA, Hickey DA (2002) Synonymous codon usage is subject to selection in thermophilic bacteria. Nucleic Acids Res 30:4272–4277
McInerney JO (1998) Replicational and transcriptional selection on codon usage in Borrelia burgdorferi. Proc Natl Acad Sci USA 95:10698–10703
Morton BR (1999) Strand asymmetry and codon usage bias in the chloroplast genome of Euglena gracilis. Proc Natl Acad Sci USA 96:5123–5128
Morton BR (2003) The role of context-dependent mutations in generating compositional and codon usage bias in grass chloroplast DNA. J Mol Evol 56:616–629
Morton BR, Wright SI (2007) Selective constraints on codon usage of nuclear genes from Arabidopsis thaliana. Mol Biol Evol 24:122–129
Nekrutenko A, Li WH (2000) Assessment of compositional heterogeneity within and between eukaryotic genomes. Genome Res 10:1986–1995
Nie X, Lv S, Zhang Y, Du X, Wang L et al (2012) Complete chloroplast genome sequence of a major invasive species, crofton weed (Ageratina adenophora). PLoS ONE 7:e36869
Noboru S (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657
Palidwor GA, Perkins TJ, Xia X (2010) A general model of codon bias due to GC mutational bias. PloS ONE 5:e13431
Rao YS, Wu GZ, Wang ZF, Chai XW, Nie QH et al (2011) Mutation bias is the driving force of codon usage in the Gallus gallus genome. DNA Res 18:499–512
Raven JA, Allen JF (2003) Genomics and chloroplast evolution: what did cyanobacteria do for plants? Genome Biol 4:209
Romero H, Zavala A, Musto H, Bernadi G (2003) The influence of translational selection on codon usage in fishes from the family Cyprinidae. Gene 317:141–147
Rosenberg MS, Subramanian S, Kumar S (2003) Patterns of transitional mutation biases within and among mammalian genomes. Mol Biol Evol 20:988–993
Sablok G, Nayak KC, Vazquez F, Tatarinova TV (2011) Synonymous codon usage, GC3, and evolutionary patterns across plastomes of three pooid model species: emerging grass genome models for monocots. Mol Biotechnol 49:116–128
Serres-Giardi L, Belkhira K, David J, Glémina S (2012) Patterns and evolution of nucleotide landscapes in seed plants. Plant Cell 4:1379–1397
Sharp PM, Li WH (1986) An evolutionary perspective on synonymous codon usage in unicellular organisms. J Mol Evol 24:28–38
Sharp PM, Cowe E, Higgins DG, Shields DC, Wolfe KH et al (1988) Codon usage in Escherichia coli, Bacillus subtilis, Saccharomyces cerevisiae, Schizosaccharomyces pombe, Drosophila melanogaster and Homo sapiens; a review of the considerable within-species diversity. Nucleic Acids Res 16:8207–8711
Sharp PM, Emery LR, Kai Z (2010) Forces that influence the evolution of codon bias. Philos Trans R Soc B 365:1203–1212
Sueoka N (1962) On the genetic basis of variation and heterogeneity of DNA base composition. Proc Natl Acad Sci USA 48:582–592
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85:2653–2657
Sueoka N (1999) Translation-coupled violation of parity rule 2 in human genes is not the case of heterogeneity of the DNA G + Ccontent of third codon position. Gene 238:53–58
Sueoka N, Kawanishi Y (2000) DNA G + C content of the third codon position and codon usage biases of human genes. Gene 261:53–62
Sugiura M (1992) The chloroplast genome. Plant Mol Biol 19:149–168
Tatarinova TV, Alexandrov NN, Bouck JB, Feldmann KA (2010) GC3 biology in corn, rice, sorghum and other grasses. BMC Genomics 11:308
Vetrivel U, Arunkumar V, Dorairaj S (2007) ACUA: A software tool for automated codon usage analysis. Bioinformation 2:62–63
Vicario S, Moriyama EN, Powell JR (2007) Codon usage in twelve species of Drosophila. BMC Evol Biol 7:226
Wan XF, Xu D, Kleinhofs A, Zhou JZ (2004) Quantitative relationship between synonymous codon usage bias and GC composition across unicellular genomes. BMC Evol Biol 4:19
Wan XF, Xu D, Zhou J (2006) CodonO: a new informatics method measuring synonymous codon usage bias. Int J Gen Syst 35:109–125
Wang B, Yuan J, Liu J, Jin L, Chen JQ (2011) Codon usage bias and determining forces in green plant mitochondrial genomes. J Integr Plant Biol 53:324–334
Waters MT, Langdale JA (2009) The making of a chloroplast. EMBO J 28:2861–2873
Wright F (1990) The ‘effective number of codons’ used in a gene. Gene 87:23–29
Xu C, Cai XN, Chen QZ, Zhou HX, Cai Y et al (2011) Factors affecting synonymous codon usage bias in chloroplast genome of Oncidium Gower Ramsey. Evol Bioinforma 7:271–278
Yoon HS, Hackett JD, Ciniglia C, Pinto G, Bhattacharya D (2004) A molecular timeline for the origin of photosynthetic eukaryotes. Mol Biol Evol 21:809–818
Zhang WJ, Zhou J, Li ZF, Wang L, Gu X et al (2007) Comparative analysis of codon usage patterns among mitochondrion, chloroplast and nuclear genes in Triticum aestivum L. J Integr Plant Biol 49:246–254
Zhang YR, Nie XJ, Jia XO, Zhao CZ, Biradar SS et al (2012) Analysis of codon usage patterns of the chloroplast genomes in the Poaceae family. Aust J Bot 60:461–470
Zhou M, Li X (2009) Analysis of synonymous codon usage patterns in different plant mitochondrial genomes. Mol Biol Rep 36:2039–2046
Zhou M, Wei L, Li X (2008) Patterns of synonymous codon usage bias in chloroplast genomes of seed plants. For Study China 10:235–242
Acknowledgments
The authors are grateful to the anonymous reviewers for their valuable comments and suggestions, and also thank Yuerong Zhang and Licao Cui for their help with statistical analysis. This research was funded mainly by the National Basic Research and Development Program of China (Grant No. 2009CB119200) and 948 Program (2010-S1), Chinese Ministry of Agriculture.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Correlation between synonymous codon usage order (SCUO) and GC composition (GC, GC1, GC2, GC3) of the five chloroplast (cp) genomes. (JPEG 99 kb)
Fig. S2
GC content and CG skew across the five plastomes. a Local GC content across plastomes. Solid lines below each horizontal axis represent local GC content. b Local CG skew across plastomes. Histograms below each horizontal axis show local CG skew. Genomic regions in which the curve is colored as yellow-green are skewed toward higher G content, whereas genomic regions in which the curve is colored as purple are skewed toward higher C content. The height of the black/red curves represents the degree of CG skew. A A. adenophora, B G. abyssinica, C H. annuus, D J. vulgari, E L. sativa (JPEG 36 kb)
Table S1
SCUO value of the orthologous genes in five Asteraceae cp genomes (DOC 61 kb)
Table S2
Summary of the correlation between SCUO and other codon indices in five Asteraceae cp genomes. (DOC 33 kb)
Rights and permissions
About this article
Cite this article
Nie, X., Deng, P., Feng, K. et al. Comparative analysis of codon usage patterns in chloroplast genomes of the Asteraceae family. Plant Mol Biol Rep 32, 828–840 (2014). https://doi.org/10.1007/s11105-013-0691-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-013-0691-z