Abstract
Aims
Key functional root traits, including mycorrhizal association and root diameter, can help project ecosystem processes like root turnover and soil carbon sequestration. It is less clear, however, how such traits relate to variations in soil biology and chemistry. Here, we examined the impact of tree species with varied root traits on soil properties and rhizoplane bacterial composition, focusing specifically on mycorrhizal association type and root diameter.
Methods
Within a long-term common garden in central Pennsylvania, USA, we selected three arbuscular mycorrhizal (AM) and three ectomycorrhizal (EM) tree species and assessed changes in (1) soil and leaf chemistry and (2) bacterial composition along fine absorptive roots, through 16S rRNA gene sequencing.
Results
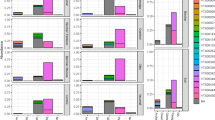
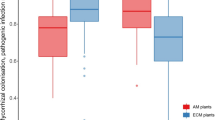
AM trees increased soil pH and soil available nitrogen relative to EM trees, and mycorrhizal association type was significantly associated with rhizoplane-associated bacterial composition. Absorptive root diameter did not clearly explain soil variability but was associated with changes in the composition of rhizoplane-associated bacteria.
Conclusions
Tree-mediated shifts to soil properties were linked to mycorrhizal association type, whereas rhizoplane recruitment of bacteria was linked to both mycorrhizal type and root diameter. This has implications for predicting changes in biogeochemical processes following shifts in tree species composition.




Similar content being viewed by others
References
Albers SV, Siebers B (2014) The prokaryotes [Internet]. Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds). Vol 9783642389, The prokaryotes: Other Major lineages of bacteria and the archaea. Springer, Berlin, Heidelberg, pp 323–346. https://doi.org/10.1007/978-3-642-38954-2
Angst G, Mueller KE, Eissenstat DM, Trumbore S, Freeman KH, Hobbie SE, Chorover J, Oleksyn J, Reich PB, Mueller CW (2018) Soil organic carbon stability in forests: distinct effects of tree species identity and traits. Glob Chang Biol 25:1529–1546. https://doi.org/10.1111/gcb.14548
Apprill A, McNally S, Parsons R, Weber L (2015) Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 75:129–137. https://doi.org/10.3354/ame01753
Bunn RA, Simpson DT, Bullington LS, Lekberg Y, Janos DP (2019) Revisiting the ‘direct mineral cycling’ hypothesis: arbuscular mycorrhizal fungi colonize leaf litter, but why? ISME J 13:1891–1898. https://doi.org/10.1038/s41396-019-0403-2
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/Nmeth.F.303
Chalot M, Brun A (1998) Physiology of organic nitrogen acquisition by ectomycorrhizal fungi and ectomycorrhizas. FEMS Microbiol Rev 22:21–44. https://doi.org/10.1111/j.1574-6976.1998.tb00359.x
Chaluvadi S, Bennetzen JL (2018) Species-associated differences in the below-ground microbiomes of wild and domesticated Setaria. Front Plant Sci 9:1183. https://doi.org/10.3389/fpls.2018.01183
Chapman SK, Langley JA, Hart SC, Koch GW (2006) Plants actively control nitrogen cycling: uncorking the microbial bottleneck. New Phytol 169:27–34. https://doi.org/10.1111/j.1469-8137.2005.01571.x
Chen W, Koide RT, Adams TS, DeForest JL, Cheng L, Eissenstat DM (2016) Root morphology and mycorrhizal symbioses together shape nutrient foraging strategies of temperate trees. Proc Natl Acad Sci USA 113:8741–8746. https://doi.org/10.1073/pnas.1601006113
Comas LH, Eissenstat DM (2009) Patterns in root trait variation among 25 co-existing North American forest species. New Phytol 182:919–928. https://doi.org/10.1111/j.1469-8137.2009.02799.x
Cornelissen J, Aerts R, Cerabolini B, Werger M, van der Heijden M (2001) Carbon cycling traits of plant species are linked with mycorrhizal strategy. Oecologia 129:611–619. https://doi.org/10.1007/s004420100752
Dukunde A, Schneider D, Schmidt M, Veldkamp E, Daniel R (2019) Tree species shape soil bacterial community structure and function in temperate deciduous forests. Front Microbiol 10:1519. https://doi.org/10.3389/fmicb.2019.01519
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112:E911–E920
Eissenstat DM, Kucharski JM, Zadworny M, Adams TS, Koide RT (2015) Linking root traits to nutrient foraging in arbuscular mycorrhizal trees in a temperate forest. New Phytol 208:114–124. https://doi.org/10.1111/nph.13451
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Fujii S, Cornelissen JHC, Berg MP, Mori AS (2018) Tree leaf and root traits mediate soil faunal contribution to litter decomposition across an elevational gradient. Funct Ecol 32:840–852. https://doi.org/10.1111/1365-2435.13027
Gould IJ, Quinton JN, Weigelt A, De Deyn GB, Bardgett RD (2016) Plant diversity and root traits benefit physical properties key to soil function in grasslands. Ecol Lett 19:1140–1149. https://doi.org/10.1111/ele.12652
Guo DL, Xia MX, Wei X, Chang WJ, Liu Y, Wang ZQ (2008) Anatomical traits associated with absorption and mycorrhizal colonization are linked to root branch order in twenty-three Chinese temperate tree species. New Phytol 180:673–683. https://doi.org/10.1111/j.1469-8137.2008.02573.x
Hiiesalu I, Bahram M, Tedersoo L (2017) Plant species richness and productivity determine the diversity of soil fungal guilds in temperate coniferous forest and bog habitats. Mol Ecol 26:4846–4858. https://doi.org/10.1111/mec.14246
Hodge A, Storer K (2015) Arbuscular mycorrhiza and nitrogen: implications for individual plants through to ecosystems. Plant Soil 386:1–19. https://doi.org/10.1007/s11104-014-2162-1
Howard MM, Bell TH, Kao-Kniffin J (2017) Soil microbiome transfer method affects microbiome composition, including dominant microorganisms, in a novel environment. FEMS Microbiol Lett 364:fnx092. https://doi.org/10.1093/femsle/fnx092
Igwe AN, Vannette RL (2019) Bacterial communities differ between plant species and soil type, and differentially influence seedling establishment on serpentine soils. Plant Soil 441:423–437. https://doi.org/10.1007/s11104-019-04135-5
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453. https://doi.org/10.1038/ismej.2008.127
Keller AB, Phillips RP (2019) Leaf litter decay rates differ between mycorrhizal groups in temperate, but not tropical, forests. New Phytol 222:556–564. https://doi.org/10.1111/nph.15524
Khlifa R, Paquette A, Messier C, Reich PB, Munson AD (2017) Do temperate tree species diversity and identity influence soil microbial community function and composition? Ecol Evol 7:7965–7974. https://doi.org/10.1002/ece3.3313
Leff JW, Jones SE, Prober SM, Barberan A, Borer ET, Firn JL, Harpole WS, Hobbie SE, Hofmockel KS, Knops JMH, McCulley RL, La Pierre K, Risch AC, Seabloom EW, Schutz M, Steenbock C, Stevens CJ, Fierer N (2015) Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci USA 112:10967–10972. https://doi.org/10.1073/pnas.1508382112
Li S peng, Wang P, Chen Y, Wilson MC, Yang X, Ma C et al (2020) Island biogeography of soil bacteria and fungi: similar patterns, but different mechanisms. ISME J [Internet] 14(7):1886–1896. https://doi.org/10.1038/s41396-020-06578
Lindahl BD, Tunlid A (2015) Ectomycorrhizal fungi - potential organic matter decomposers, yet not saprotrophs. New Phytol 205:1443–1447. https://doi.org/10.1111/nph.13201
Ma Z, Guo D, Xu X, Lu M, Bardgett RD, Eissenstat DM, McCormack ML, Hedin LO (2018) Evolutionary history resolves global organization of root functional traits. Nature 555:94–97. https://doi.org/10.1038/nature25783
Marini RP (1999) Are nonsignificant differences really not significant? HortScience 34:761–762. https://doi.org/10.21273/hortsci.34.5.761
Matejovic I (1996) The application of Dumas method for determination of carbon, nitrogen, and sulphur in plant samples. Rostlinna Vyroba 42:313–316
McCormack ML, Adams TS, Smithwick EA, Eissenstat DM (2012) Predicting fine root lifespan from plant functional traits in temperate trees. New Phytol 195:823–831. https://doi.org/10.1111/j.1469-8137.2012.04198.x
McCormack ML, Dickie IA, Eissenstat DM, Fahey TJ, Fernandez CW, Guo D, Helmisaari HS, Hobbie EA, Iversen CM, Jackson RB, Leppalammi-Kujansuu J, Norby RJ, Phillips RP, Pregitzer KS, Pritchard SG, Rewald B, Zadworny M (2015) Redefining fine roots improves understanding of below-ground contributions to terrestrial biosphere processes. New Phytol 207:505–518. https://doi.org/10.1111/nph.13363
McCormack ML, Guo D, Iversen CM, Chen W, Eissenstat DM, Fernandez CW, Li L, Ma C, Ma Z, Poorter H, Reich PB, Zadworny M, Zanne A (2017) Building a better foundation: improving root-trait measurements to understand and model plant and ecosystem processes. New Phytol 215:27–37. https://doi.org/10.1111/nph.14459
Mehlich A (1984) Mehlich-3 soil test extractant - a modification of Mehlich-2 extractant. Commun Soil Sci Plan 15:1409–1416. https://doi.org/10.1080/00103628409367568
Merino-Martin L, Griffiths RI, Gweon HS, Furget-Bretagnon C, Oliver A, Mao Z, Le Bissonnais Y, Stokes A (2020) Rhizosphere bacteria are more strongly related to plant root traits than fungi in temperate montane forests: insights from closed and open forest patches along an elevational gradient. Plant Soil 450:183–200. https://doi.org/10.1007/s11104-020-04479-3
Midgley MG, Brzostek E, Phillips RP (2015) Decay rates of leaf litters from arbuscular mycorrhizal trees are more sensitive to soil effects than litters from ectomycorrhizal trees. J Ecol 103:1454–1463. https://doi.org/10.1111/1365-2745.12467
Mueller KE, Eissenstat DM, Hobbie SE, Oleksyn J, Jagodzinski AM, Reich PB, Chadwick OA, Chorover J (2012) Tree species effects on coupled cycles of carbon, nitrogen, and acidity in mineral soils at a common garden experiment. Biogeochemistry 111:601–614. https://doi.org/10.1007/s10533-011-9695-7
Oksanen J, Blanchet GB, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2015) Vegan: Community ecology package. R package version 23–0. https://cran.r-project.org/web/packages/vegan/vegan.pdf
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414. https://doi.org/10.1111/1462-2920.13023
Pascale A, Proietti S, Pantelides IS, Stringlis IA (2019) Modulation of the root microbiome by plant molecules: the basis for targeted disease suppression and plant growth promotion. Front Plant Sci 10:1741. https://doi.org/10.3389/fpls.2019.01741
Pei ZQ, Eichenberg D, Bruelheide H, Krober W, Kuhn P, Li Y, von Oheimb G, Purschke O, Scholten T, Buscot F, Gutknecht JLM (2016) Soil and tree species traits both shape soil microbial communities during early growth of Chinese subtropical forests. 96:180–190. https://doi.org/10.1016/j.soilbio.2016.02.004
Perez-Jaramillo JE, Carrion VJ, de Hollander M, Raaijmakers JM (2018) The wild side of plant microbiomes. Microbiome 6:143. https://doi.org/10.1186/s40168-018-0519-z
Phillips RP, Fahey TJ (2006) Tree species and mycorrhizal associations influence the magnitude of rhizosphere effects. Ecology 87:1302–1313. https://doi.org/10.1890/0012-9658(2006)87[1302:Tsamai]2.0.Co;2
Phillips RP, Brzostek E, Midgley MG (2013) The mycorrhizal-associated nutrient economy: a new framework for predicting carbon-nutrient couplings in temperate forests. New Phytol 199:41–51. https://doi.org/10.1111/nph.12221
Prescott CE, Grayston SJ (2013) Tree species influence on microbial communities in litter and soil: Current knowledge and research needs. Forest Ecol Manag 309:19–27. https://doi.org/10.1016/j.foreco.2013.02.034
Pylro VS, Roesch LFW, Morais DK, Clark IM, Hirsch PR, Tótola MR (2014) Data analysis for 16S microbial profiling from different benchtop sequencing platforms. J Microbiol Meth 107:30–37. https://doi.org/10.1016/J.Mimet.2014.08.018
Ramirez KS, Craine JM, Fierer N (2012) Consistent effects of nitrogen amendments on soil microbial communities and processes across biomes. Glob Change Biol 18:1918–1927
Read DJ, Perez-Moreno J (2003) Mycorrhizas and nutrient cycling in ecosystems - a journey towards relevance? New Phytol 157:475–492. https://doi.org/10.1046/j.1469-8137.2003.00704.x
Reich PB, Oleksyn J, Modrzynski J, Mrozinski P, Hobbie SE, Eissenstat DM, Chorover J, Chadwick OA, Hale CM, Tjoelker MG (2005) Linking litter calcium, earthworms and soil properties: a common garden test with 14 tree species. Ecol Lett 8:811–818. https://doi.org/10.1111/j.1461-0248.2005.00779.x
Rognes T, Flouri T, Nichols B, Quince C, Mahe F (2016) VSEARCH: a versatile open source tool for metagenomics. Peerj 4:e2584. https://doi.org/10.7717/peerj.2584
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/Aem.01541-09
Tkacz A, Bestion E, Bo Z, Hortala M, Poole PS (2020) Influence of plant fraction, soil, and plant species on microbiota: a multikingdom comparison. mBio 11:e02785–e02719. https://doi.org/10.1128/mBio.02785-19
Trexler RV, Bell TH (2019) Testing sustained soil-to-soil contact as an approach for limiting the abiotic influence of source soils during experimental microbiome transfer. FEMS Microbiol Lett 366:fnz228. https://doi.org/10.1093/femsle/fnz228
van der Heijden MG, Martin FM, Selosse MA, Sanders IR (2015) Mycorrhizal ecology and evolution: the past, the present, and the future. New Phytol 205:1406–1423. https://doi.org/10.1111/nph.13288
White CM, DuPont ST, Hautau M, Hartman D, Finney DM, Bradley B, LaChance JC, Kaye JP (2017) Managing the trade off between nitrogen supply and retention with cover crop mixtures. Agric Ecosyst Environ 237:121–133. https://doi.org/10.1016/j.agee.2016.12.016
Yin HJ, Wheeler E, Phillips RP (2014) Root-induced changes in nutrient cycling in forests depend on exudation rates. Soil Biol Biochem 78:213–221. https://doi.org/10.1016/j.soilbio.2014.07.022
Acknowledgements
The authors thank Timothy Peoples, Franco Acevedo Luco, Jeremy Harper, Amanda Seow, and Wenqi Luo for their help with root sampling, Kevin Hockett for the use of a sonic water bath and technical advice, and Ephraim Muchada Govere and Brosi Bradley for training on soil testing. This work was supported by the USDA National Institute of Food and Agriculture (NIFA) Foundational Program (Accession number 1014758) and by the USDA NIFA Federal Appropriation under Projects PEN0 4628, 4651, and 4744 (Accession numbers 1014131, 1016233, and 1023222, respectively). Partial support was also provided by the China Scholarship Council.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Amandine Erktan.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 386 KB)
Rights and permissions
About this article
Cite this article
Yates, C.F., Guo, J., Bell, T.H. et al. Tree‐induced alterations to soil properties and rhizoplane‐associated bacteria following 23 years in a common garden. Plant Soil 461, 591–602 (2021). https://doi.org/10.1007/s11104-021-04846-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-04846-8