Abstract
Aims
This study examines how inoculation with live soil influences the assembly of the endosphere microbiome of leaves and roots of chrysanthemum.
Methods
Sterilized soil was inoculated with 10% soil in which grasses had grown. Chrysanthemum was planted in these soils and control plants were grown in 100% sterilized soil. All plants were exposed to thrips, and leaves and roots were collected from inoculated and control plants that experienced high and low thrips damage. DNA was extracted and the bacterial and fungal community inside roots and leaves was determined using Illumina sequencing.
Results
Inoculation increased bacterial diversity in root but not in leaf tissues. The endosphere of both roots and leaves was dominated by Pseudomonadaceae. In leaves, the relative abundance of Pseudomonadaceae was higher in inoculated than in control plants, whereas this was opposite in roots. Leaves contained more rare bacterial families than roots. The number of fungal reads was very low and the endopshere did not differ between plants with high or low thrips damage.
Conclusions
Bacterial communities inside chrysanthemum root and leaf tissues differ considerably. Soil inoculation with entire microbiomes can be used to change root and foliar bacterial microbiomes and this is particularly effective in root tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Understanding below-aboveground interactions is a central theme in ecology, and essential for our insight into how terrestrial communities establish and function (Bardgett and Wardle 2010). An increasing number of studies is now showing that single organisms inhabiting the soil interact with plant roots to cause a cascade of physiological changes in the plant that ultimately affect organisms interacting with aboveground plant tissues (e.g. Bezemer and Van Dam 2005; Pangesti et al. 2013; Pieterse et al. 2016; Pineda et al. 2017). In parallel, there is a rapidly growing interest in understanding how entire microbiomes in soils and inside plants influence plant growth and health. A major reason is the increasing awareness that full microbiomes or consortia of several strains can provide functions to their plant hosts e.g. related to nutrition or immunity (Herrera Paredes et al. 2018; Hubbard et al. 2019; Wang et al. 2019). These functions were previously attributed to single microbial strains. However, many aspects of the role of microbiomes in influencing ecological functions of their hosts, and how microbiomes assemble inside their hosts, are still poorly understood.
The plant endosphere compartment is under strong influence of the host plant and typically contains a much lower microbial diversity than the rhizosphere (Schlaeppi and Bulgarelli 2014; Fitzpatrick et al. 2018). Endophytic microbes can be maternally transferred via the seeds but can also be acquired from the environment, and the soil is considered one of the main sources for endophytic microbes in plants (Schlaeppi and Bulgarelli 2014). To what extent the composition of the endophytic microbial community will depend on the rhizosphere community and hence on the microbiome of the soil in which the plant is rooted, is poorly understood. Within the plant, microbial assembly may also vary between compartments (e.g. Bulgarelli et al. 2013; Wagner et al. 2016). Generally, communities in the root endosphere are more diverse than within leaves (e.g. Wagner et al. 2016; Hannula et al. 2019). However, which endophytes are unique for roots and leaves, which ones are shared between compartments, and how this depends on the microbial community of the soil is not well known. Such insights are important as most of the knowledge on the contribution of beneficial microbes to the tolerance of plants to abiotic stress and their resistance to pathogens and herbivores, so far, comes from rhizosphere microbes. However, evidence is accumulating that important functions provided to the plant are also mediated by endophytic microbes (Hardoim et al. 2015; Cordovez et al. 2019; Carrión et al. 2019).
Soil inoculation is a strategy to steer the microbiome associated to plants. In previous studies our group has shown that by inoculating a background soil with a small percentage of soil that contains desired microbiomes, the soil microbiome in the background soil can be changed with cascading consequences to plant communities and even to the microbiome assembly of foliar feeding herbivore insects (Wubs et al. 2016; Hannula et al. 2019). Soil inoculation can also be applied to improve the health of agricultural crops as we have shown for chrysanthemum, an important horticultural cut flower crop (Ma et al. 2017, 2020; Hannula et al. 2020; Pineda et al. 2020). In the Netherlands this ornamental crop is grown in soil in sophisticated greenhouses, where the soil is regularly sterilized by steaming to eliminate soil borne diseases. In our previous studies we observed that inoculating the sterilized soil with soil containing microbiomes of grass species increased the biomass of chrysanthemum when grown in presence of Pythium, a soil pathogen (Ma et al. 2017, 2020; Hannula et al. 2020). Moreover, inoculation increased resistance of the plant to herbivory by thrips (Pineda et al. 2020), and it suppressed negative plant-soil feedback effects on plant growth (Ma et al. 2018). How soil inoculation affects the endosphere microbiome in chrysanthemum is, however, not yet known.
In this study, our objective is to examine how soil inoculation with a microbial community influences the assembly of the endosphere microbiome of leaves and roots of chrysanthemum. We first created a soil inoculum by growing a mixture of four wild grass species in live soil collected from a natural grassland, and inoculated a subset of this soil into sterilized soil. We then grew chrysanthemum in inoculated and uninoculated sterilized soil. All plants in this experiment were exposed to thrips. We measured plant height and aboveground biomass, and analysed the leaf and root endosphere microbiome, with the aim of answering the following research questions:
-
1
How does soil inoculation influence the diversity and composition of the endophytic bacteria and fungi in chrysanthemum leaves and roots?
-
2
What is the relationship between endophytic microbiomes of roots and leaves in chrysanthemum and which taxa are unique in roots and leaves?
There is an urgent need for more sustainable crop production systems. Hence, our study also tests whether soil inoculation can be used to alter endophytic microbiomes in crops with potential to promote plant health.
Materials and methods
The focal plant in our study is Dendranthema X grandiflora (Ramat.) Kitam. cv. Amadea [Chrysanthemum, syn. Chrysanthemum X morifolium (Ramat.) Hemsl., Asteraceae]. Chrysanthemum cuttings were provided by the breeding company Deliflor (Maasdijk, The Netherlands). To create the soil inoculum, we grew a mix of four wild grass species (Holcus lanatus, Lolium perenne, Alopecurus pratensis, Festuca ovina; all Poaceae) based on previous work that showed positive effects of soils of these grasses on chrysanthemum growth (Ma et al. 2017). Seeds of the four grasses were obtained from a wild plant seed supplier (Cruydt-Hoeck, Assen, The Netherlands). Seeds were surface sterilized (1 min in 1% sodium hypochlorite solution and rinsed with distilled water afterwards) and germinated on sterile glass beads in a climate chamber at 20 °C/15 °C (16 h/8 h, light/dark). The experiment was conducted in the experimental greenhouse facility at the NIOO-KNAW (Wageningen, The Netherlands), and consisted of two phases.
Phase 1: Soil conditioning
In the first phase, the conditioning phase, we grew the mixtures of the four grass species in field soil. Field soil was collected (5–20 cm below the surface) from a species-rich natural grassland (Mossel, Ede, Netherlands). This site has a sandy loam soil (83% sand, 10% silt, 4% clay, 3% organic matter). The soil was sieved (1 cm mesh size) to remove coarse fragments and macro-arthropods and homogenized. Then, four one-week-old seedlings (one of each of the four grasses species) were transplanted into each of 14 pots (11 × 11 × 12 cm) filled with 1 kg of soil. Seedlings that died within 7 days were replaced once. Pots were randomly placed in a greenhouse with controlled conditions: 70% RH and 16 h light (21 °C) /8 h dark (16 °C) photo regime. Natural daylight was supplemented with 400 W metal halide lamps (225 μmol s−1 m−2 photosynthetically active radiation, 1 lamp per 1.5 m2). Plants were checked three times per week and watered as needed. Ten weeks after transplanting, soil from the entire pot was collected by hand-shaking the full root system, keeping finer roots in the collected soil to act as a source of micro-organisms. Soil from each pot was homogenized and stored individually in closed plastic bags at 4 °C until used as soil inoculum in the test (feedback) phase. From previous studies, we know that conditioning field soil with grasses creates unique microbiomes that are beneficial for chrysanthemum growth (Ma et al. 2017; Hannula et al. 2020; Pineda et al. 2020).
Phase 2: Test phase
In the second phase, we inoculated sterilized soil and grew chrysanthemum plants in these soils. Soil for sterilization was collected from the area where the soil for conditioning was also collected. After sieving and homogenizing, the soil was sterilized by gamma irradiation (>25 K Gray gamma irradiation, Isotron, Ede, Netherlands). Fifty pots (11 cm × 11 cm × 12 cm) were filled with one kg of a homogenized mixture of 10% soil inoculum and 90% sterilized soil (based on weight). As a control we included fifty pots filled with 100% sterilized soil. Prior to planting, the soil in each pot was well watered and 100 ml half-strength Hoagland nutrient solution was added. Two chrysanthemum cuttings (without roots) were planted in each pot, and these were randomly placed on trolleys that were tightly covered with a thin transparent plastic foil for 10 days to create an environment with high humidity that favors rooting. After 10 days, one of the chrysanthemum cuttings was removed from each pot. For a few pots both cuttings did not survive; resulting in 46 pots with inoculated soil and 49 pots with 100% sterilized soil. Four days later plant height of all plants was measured. Plants were fertilized following grower practices: half-strength Hoagland nutrient solution for the first 2 weeks, and single strength Hoagland solution during the following weeks. Four weeks after planting, plant height was measured again.
Previously we observed that soil inoculation led to a reduction in thrips damage in chrysanthemum (Pineda et al. 2020). Here, our objective was to examine the interactive effects of inoculation and thrips damage. We transferred 20 adult thrips (Frankliniella occidentalis) from a culture to a subset of plants and kept all plants caged for 15 days. The culture of the thrips was established with a starting colony provided by the company Hazera Seeds (Made, The Netherlands). Thrips were reared at 16 h light and 8 h dark photo regime and 25 °C for multiple generations on pods of Romano beans (Vicia faba) purchased weekly in a local supermarket. Unfortunately, thrips were found on all plants. Therefore in this study there were no control plants. After 15 days, the cages were removed and plants were further colonized by the thrips that were present in the greenhouse compartment for two more weeks. For each plant, thrips damage was assessed as silver leaf damage, and based on visual scoring, classified as high or low damage, so that we could determine whether the endophytic microbiomes differed between the two groups. Examples of the two damage levels are shown in Fig. S1. Six to seven plants were then randomly selected from each of the four treatments (inoculated- high thrips damage, inoculated-low thrips damage, control-high thrips damage, control-low thrips damage) for molecular analysis. A leaf and root sample of each of 25 plants was collected (see below) and the remaining aboveground biomass of these plants was dried at 60 °C and dry weight was determined. Results related to high and low thrips damage are presented in the Supplementary information.
Endosphere microbiome analysis
From each selected plant, one fully developed leaf was carefully cut with a scalpel (sterilized with ethanol in between usage) and the leaf was placed into a sterile petri dish. The stem of each plant was cut at soil level and roots were carefully removed from the pot. Soil was removed first by hand shaking the roots and then washing the roots with tap water. Immediately afterwards, leaf and root samples were surface sterilized as follows: The samples were first washed with 10 mM MgSO4 and this was repeated until the buffer solution was clear. Samples were then soaked in 10 mM MgSO4 + Tween 20 (0.01%), rinsed with 10 mM MgSO4, and surface sterilized in bleach solution (0.1% for leaf and 1% for root) for 1 min. Each sample was then immediately rinsed with sterilized water two to five times. To check the quality of the surface sterilization procedure, 100 μl of the final rinsed solution of two representative samples was plated in a 1/10 Tryptic Soy Agar (TSA) and Reasoner’s 2A (R2A) agar media and this resulted in zero colonies. The surface sterilized samples (4 g) were ground in 1 ml of 10 mM MgSO4 buffer, followed by filtering through double layers of miracloth paper. The filtrate was then centrifuged at maximum speed of 14,000 rpm for 30 min, and the pellet was used for microbial DNA extraction.
The microbial DNA extraction was performed using the FastDNA SPIN Kit (MP Biomedicals, Solon, OH, USA) following the manufacturer’s protocol. The DNA concentration and purity were measured using a Nanodrop spectrophotometer (Thermo Scientific, Hudson, NH, USA). The primers 515FB and 806RB targeting the V4 region of 16Sr RNA for bacteria (Caporaso et al. 2011: Apprill et al. 2015; Parada et al. 2016), and ITS4ngs and ITS3mix targeting the ITS2 region of the fungi were used (Tedersoo et al. 2015). During the amplification step PCR clamps were incorporated to block amplification of plant chloroplast and mitochondria – following the method developed by Lundberg et al. (2013). The bacterial 16Sr RNA amplification was performed with initial denaturation at 98 °C for 3 min followed by 25 cycles of 98 °C for 15 s, 55 °C for 15 s and 72 °C for 30 s, and a final extension at 72 °C for 3 min. The fungal ITS2 amplification was performed with initial denaturation at 98 °C for 3 min followed by 30 cycles of 98 °C for 15 s, 55 °C for 15 s and 72 °C for 30 s, with a final extension at 72 °C for 3 min. A mock community was included that consisted of 10 fungal strains and bacterial symbionts as a positive control, whereas sterilized water was used as a negative control. For library preparation, the PCR products were purified using Agencourt AMPure XP magnetic beads (Beckman Coulter). Adapters and barcodes were incorporated to each sample using the Nextera XT DNA library preparation kit sets A-C (Illumina, San Diego, CA, USA). The final PCR products were purified with AMPure beads, quantified with a Nanodrop spectrophotometer, and checked in gel electrophoresis before equimolar pooling. The samples were sequenced at McGill University and the Génome Québec Innovation Centre (Canada) using the MiSeq PE250 platform.
Bioinformatics and statistical analysis
Sequencing results of the bacterial endophytic community were analysed using an in-house pipeline using SILVA database with SINA classifier (De Hollander 2017), whereas the fungal endophytic community was analysed using PIPITS pipeline (Gweon et al. 2015). Taxonomy of fungi was assigned using rdp classifier against the UNITE fungal database (Abarenkov et al. 2010). All reads from mitochondria, chloroplast, archaea, and singletons and doubletons were removed from the dataset. For bacteria, samples with fewer than 800 reads were then removed from dataset. The number of reads for each sample was transformed to relative abundances by dividing the reads of each OTU by the total number of reads per sample. OTUs with abundance less than 0.00111 for leaf samples and 0.00004 for root samples were removed to normalize the data. This cut-off point was obtained using formula 2 × 1/(average reads per leaf or root compartment). Normalized data were then back-calculated by multiplying the relative value of each OTU by the total number of reads for the sample in original dataset. Back-calculated data were then transformed to relative data and used for further statistical analyses. The effect of soil inoculation and plant compartment (leaf/root) on bacterial Simpson diversity (log-transformed data) and evenness were examined using a two-way ANOVA, followed by a Tukey post-hoc test. PERMANOVA (permutation MANOVA using a distance matrix, R package ‘vegan’) with Bray-Curtis dissimilarity was used to test whether soil inoculation and thrips damage affected bacterial community composition in the different plant compartments. Separation of microbiomes among treatments was visualized using non-metric multidimensional scaling (NMDS) based on a Bray-Curtis dissimilarity matrix. To compare the bacterial composition in the two compartments (leaves and roots) and between high and low damage, a Venn diagram was constructed. Only those OTUs were included that were present in at least two samples per treatment combination. We also compared the relative abundances of OTUs that were present in at least two samples per treatment in both leaves and roots. To explore which bacterial taxa were over- or underrepresented in the treatments (leaves/roots, inoculated/control), the “metacoder” packages in R was used (Foster et al. 2017). The expression of taxa was considered to be significantly different when the Wilcoxon p value for the log-2 ratio of the median counts between treatments was smaller than 0.05 after correction for false discovery rates. Taxa showing the largest differences between treatments were further evaluated using ANOVA.
The number of obtained reads of fungal origin was very low despite high total read numbers obtained through deep sequencing of the samples (see results). Due to low coverage, fungal endophytes in leaf samples were not further analyzed. For roots, three samples had less than 90 fungal reads, these samples were omitted for further analysis. The remaining samples had on average 504 reads. For fungal reads, the number of reads was not related to the number of phylotypes per sample and hence normalization was not done for this dataset.
To determine whether plant biomass and height differed between inoculated and control plants and for plants with high and low thrips damage, data were analyzed using two-way ANOVA.
Results
Soil inoculation effects on the diversity and assembly of bacteria in roots and leaves
Soil inoculation altered bacterial diversity and evenness (Fig. 1). In roots, inoculation led to higher diversity and evenness but this was not so in leaves, resulting in a significant interaction between treatment and compartment (Fig. 1). Moreover, bacterial diversity in roots was higher than in leaves, while the evenness did not differ in both compartments. Bacterial diversity and composition in leaves and roots was similar in plants with high and low thrips damage (Fig. S2).
Mean (± SE) bacterial diversity (a) and evenness (b) in leaves and roots of chrysanthemum, in control and inoculated soils and results of a two-way ANOVA testing the effect of inoculation (treatment) and plant compartment. Bars with identical letters are not significantly different (Tukey post hoc test, P < 0.05)
The composition of the bacterial community varied strongly between roots and leaves (Permanova: F1,37 = 38.6; P < 0.001; 41.2% explained variation). However, the inoculation treatment also explained a significant part of the variation (F1,37 = 8.11; P = 0.003; 8.6% explained variation) and this effect was stronger in roots than in leaves resulting in a significant interaction between treatment and compartment (F1,37 = 10.11; P < 0.001; 10.8% explained variation; Fig. 2; Table S1). The composition of the bacterial community did not differ in plants with high and low thrips damage (F1,37 = 0.52; P = 0.705; 2.6% explained variation; Fig. S3).
Nonmetric multidimensional scaling (NMDS) of bacterial composition in the endophytic microbiome of chrysanthemum in leaves of plants growing in inoculated (blue), and control (orange) soil, and in roots of plants growing in inoculated (grey) and control (yellow) soil. Soil conditioned by grasses (10%) was added to inoculated treatments
The dominant taxa making up more than 90% of the average bacterial reads in leaf (91%) and root (95%), belonged to the families Pseudomonadaceae (71%), Xanthomonadaceae (16%) and Enterobacteriaceae (6%) (Fig. 3a). The relative abundance of the family Pseudomonadaceae was higher in leaves than in roots, while Xanthomonadaceae and Enterobacteriaceae were more abundant in roots than in leaves. In leaves, the relative abundance of Pseudomonadaceae was higher in inoculated than in control plants, whereas this was opposite in roots. For Xanthomondaceae the reverse pattern was observed. In roots, Xanthomonadaceae were relatively more, and in leaves less abundant in the inoculated than in the control treatment (Fig. 3a). Interestingly, leaves contained more rare bacterial families (with a relative abundance of less than 0.5%) than roots. In total, 15 and 17 rare bacterial families were present in leaves of inoculated and control plants respectively, whereas 13 and 7 rare bacterial families were present in roots of inoculated and control plants (Fig. 3b).
Soil inoculation effects on individual bacterial OTUs in roots and leaves
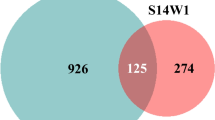
Thirteen bacterial OTUs were unique for plants grown in inoculated soil (1 in leaf and 12 in root tissues) while 12 bacterial OTUs were only found in plants grown in control soil (5 only in leaves, 6 only in roots and one in leaf and root tissues; Fig. 4). There were 44 OTUs unique for roots, and 13 OTUs unique for leaves that were found in both the control and inoculated treatments and 22 OTUs that were found in leaves and roots in both soil treatments, suggesting the existence of a core endophytic bacterial assemblage. Among the OTUs, bacteria from the genera Pseudomonas, Luteibacter, Pantoea, Ralstonia, Propinibacterium, Sphingomonas and Pelomonas significantly differed between roots and leaves and between inoculated and control soils (Table 1). Figure 5 visualizes the bacterial taxa that differ between inoculated and control plants in leaves and roots. Especially, Pseudomonas (F = 4.87, P = 0.04) and Acinetobacter (F = 4.38, P = 0.04) were significantly more abundant in leaves of inoculated plants than in leaves of control plants while Enterobacteriales (F = 4.57, P = 0.03) and Luteibacter (F = 4.61, P = 0.02) were enriched significantly in the leaves of control plants (Fig. 5). In roots, Pseudomonas (F = 9.97, P = 0.005) and Serratia (F = 3.51, P = 0.04) were more abundant when plants grew in control soil and Enterobacter (F = 4.13, P = 0.04) and Luteibacter (F = 3.49, P = 0.05) more abundant in plants grown in inoculated soil.
Comparison of bacterial taxa in leaves (a) and roots (c) of plants in the control and inoculated treatment. Red branches indicate significantly higher relative abundance in the control and blue branches in the inoculated soils. In (a) and (c) black taxa indicate that there are red and blue branches within the taxa. In (b) and (d) the treatment effects for the most significant genera in the bacterial class Gammaproteobacteria are highlighted for leaves and roots respectively
Of the OTUs found both in leaves and roots in the same plant, thirteen differed significantly between leaves and roots (Fig. 6, Table 1). Five OTUs were more abundant in the roots, and seven OTUs were more abundant in the leaves. Among the five OTUs that were more abundant in roots, OTU 6, identified as Pseudomonas, was significantly more abundant in plants grown in inoculated than in control soil (Table 1). Among the seven OTUs that were more abundant in leaves, OTU 1 and 12 both also identified as Pseudomonas, were more abundant in plants grown in inoculated than in control soils (Table 1). For six OTUs identified as Pseudomonas the soil inoculation effect varied between the two plant compartments (significant interactions between treatment and compartment).
Soil inoculation effects on fungi in roots and leaves
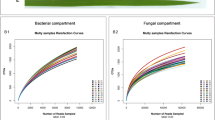
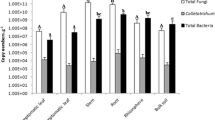
Virtually no reads from fungal origin were detected in the leaves of chrysanthemum plants (only 1 to 14 reads recovered per sample) even though around 50,000 reads were recovered in the sequenced samples. The number of fungal reads in root samples was higher than in leaf samples but still low (on average 504 fungal reads per sample). In roots, 24 fungal phylotypes were detected that occurred in at least two samples. Sordariomycetes (Ascomycota) were most frequently detected (identified as Fusarium, Chloridium and an unknown Sordariomycetes). More reads were obtained from control (705 ± 209) than from inoculated (302 ± 118) roots, but the number of fungal phylotypes was much higher (t14 = 5.10; P < 0.01) in inoculated (13.4 ± 1.4) than in in control roots (5.1 ± 0.8). Fungal patterns did not differ between plants with high and low thrips damage (Data not shown). Shoot biomass and plant height did not significantly differ between inoculated and control plants and did not vary significantly between plants with high and low thrips damage (Fig. S4).
Discussion
In this study we examined the composition of the endophytic leaf and root microbiome of chrysanthemum plants. Several studies have shown that whole soil inoculation can have cascading effects on the growth and defense responses of plants that grow in inoculated soil (e.g. Wang et al. 2019; Kos et al. 2015). However, how inoculation of the soil with new microbiomes derived from growing plants in donor soil, influences the endophytic microbiome in a plant, our main objective here, is still poorly understood. Our study now provides two important insights: first, soil inoculation changes the composition and increases the diversity of the endosphere community in a plant, but plants grown in sterilized soil also exhibit an endophytic microbiome. The source of microbes in the plants grown in sterilized soil (control treatment) could be due to incomplete sterilization, or from b) the cuttings of chrysanthemum that carried endophytic bacteria vertically transferred from the mother plant, c) the greenhouse environment where the plants were grown, d) unsterilized water used to water the plants during the experiment; or e) from the thrips. Second, the composition and diversity of the bacterial community in the roots and leaves of chrysanthemum differs considerably. Bacterial diversity is much higher in roots than in leaves, but importantly, inoculation of the soil in which the plant grows increases the diversity of bacteria in roots but not in the leaves. Hence, soil inoculation can have different effects on the microbes inhabiting above- and belowground plant parts.
Our study shows that plants, via their impact on the soil microbiome, can influence the endophytic community in other plants that grow later in the soil. Many studies have shown that plants create different soil microbiomes (e.g. Fitzpatrick et al. 2018), and that these plant -driven legacies in the soil can alter the performance of later growing plants (reviewed by Van der Putten et al. 2013). However, how these plant-mediated effects on soil microbiomes influence endophytes in later growing plants is not well-studied. Pattison et al. (2016) showed that the invasive plant Impatiens glandulifera has more fungal leaf endophyte species when grown in soil in which plants of the same species had been grown than in unconditioned soil in which no plant had been grown previously. Similarly, in our study we also showed that inoculation with plant-conditioned soil led to higher diversity of bacteria in chrysanthemum roots. However, we did not observe this effect in leaf tissues. In the study of Pattison et al. (2016) plants with more fungal species also were taller and produced more leaves, indicating a positive relationship between endophytic diversity and plant growth. In our study, we did not observe differences in growth between inoculated and uninoculated plants. It is important to mention that in our design plants received very high amounts of fertilizer and it is well possible that the bacterial endophytes provide benefits to the plant such as growth promotion, that are only visible under less optimal conditions. A limitation of the current study is that we did not measure the microbial composition in the inoculum. Hence we cannot proof wich microbes present in the inoculum entered the plant. In an earlier study (Pineda et al. 2020) we compared the bacterial and fungal community of soil inocula (conditioned by different species of grasses and forbs) and of sterilized soil. In that study, soil inocula have much higher bacterial and fungal richness than the sterilized soil as expected, and several bacterial phyla (Acidobacteria, Traumarcheota, Armatimonadetes, Atribacteria, BRC1, Candidatus_Berkelbacteria, Chlamydiae, Elusimicrobia, FBP, FCPU426, Fibrobacteres, Gemmatimonadetes, Hydrogenedentes, Nitrospirae, Parcubacteria, Verrucomicrobia, TM6) and a fungal phylum (Mucoromycota) were enriched in soil inocula.
A somewhat surprising outcome of our study is that we detected very few fungal OTUs in chrysanthemum roots, and no fungal community inside chrysanthemum leaves. In earlier studies, we analyzed the microbiome of bulk soil after chrysanthemum had grown in the soil, and the root associated (rhizosphere) microbiome of chrysanthemum. There we observed that this plant species exerts a negative effect on fungal species, and suppresses arbuscular mycorrhizal fungi (Ma et al. 2020; Pineda et al. 2020). A possible explanation for this could be that chrysanthemum roots and leaves contain high concentrations of chlorogenic acid. This compound has been described as a potent antifungal compound (Martínez et al. 2017). Further studies are needed that examine the relationship between soil microbes, plant endophytes, and fungi in chrysanthemum in more detail.
Among the bacterial endophytes, the family Pseudomonadaceae is the most abundant one, and OTU’s of the genus Pseudomonas show different patterns in leaves and roots and in control and in inoculated plants. Pseudomonas is a genus that consists of common bacterial endophytes that colonize different crops species. In a study with wheat cultivars, it even was the only endophytic genus colonizing two cultivars throughout a number of different plant stages - from the endosperm stage to the leaf formation stage (Kuźniar et al. 2020). Several strains of Pseudomonas are known to cause induced systemic resistance in plants which protects the plant from insect herbivores, and induce growth promotion in both shoots and roots (e.g. Pangesti et al. 2017; van de Mortel et al. 2012). Other species of the genus Pseudomonas are known as pathogens of chrysanthemum. For instance, it is known that Pseudomonas cichorii causes stem necrosis (Jones et al. 1983) and that Pseudomonas syringae causes necrotic leaf spots in chrysanthemum (Carta 1993). How the full microbiome network inside the plant supports or inhibits the functions of specific strains is a fascinating topic that needs further research.
In the conditions here evaluated, the influence of soil inoculation on the endophytic microbiome did not explain whether a plant displayed low or high thrips damage. Particular endophytic microbes such as Trichoderma can enhance plant resistance against thrips (Muvea et al. 2014). In a previous study with a different chrysanthemum cultivar we observed that soil inoculation with soils previously conditioned by several wild plant species reduced the performance of thrips (Pineda et al. 2020). Hence, our study highlights that microbe-plant-insect interactions are highly context-dependent, and that soil, combination of microbial strains present, plant genotype, and insect behaviour, all could determine whether microbial inoculation will enhance or suppress plant resistance to aboveground herbivory (Gadhave et al. 2018).
Interestingly, soil inoculation effects were more visible in the root than in the leaf compartment. Roots are in direct contact with the soil and the results suggest that the microbes specific for the inoculum did not enter the leaves. However, a recent study with the grass Deschamptia flexuosa showed that manipulation in the soil (mycorrhizal addition) lead to altered endophytic communities in the leaves (Poosakkannu et al. 2017). Alternatively, in our study, it is possible that the lesser effect of the soil inoculation on leaf endophytes, is due an overriding effect of thrips on the microbiome composition in leaves. Some thrips species are known to be vectors of bacterial and fungal diseases, for instance those belonging to the genus Pantoea (Wells et al. 2002). Unfortunately, our study does not provide a definite answer as to why the root and shoot endophytic communities responded differently to soil inoculation and a different type of design would be necessary to further explore the mechanisms involved.
In conclusion, our study shows that soil inoculation with entire microbiomes changes the root and foliar endophytic bacterial microbiome of chrysanthemum. The effects are particularly visible in root tissues. The endophytic microbiome of chrysanthemum was dominated by Pseudomonadacea and Xanthomonadacea. In leaves, the relative abundance of Pseudomonadaceae was higher in inoculated than in uninoculated plants, while Xanthomonadaceae showed the opposite pattern. In roots this was reversed, Xanthomonadaceae were relatively more abundant in the inoculated than in control plants, and Pseudomonadaceae were more abundant in control plants. Future studies should focus on the functions and ecological role of these endophytic bacteria in this important ornamental crop.
References
Abarenkov K, Henrik Nilsson R, Larsson KH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T, Sen R, Taylor AF, Tedersoo L, Ursing BM, Vrålstad T, Liimatainen K, Peintner U, Kõljalg U (2010) The UNITE database for molecular identification of fungi – recent updates and future perspectives. New Phytol 186:281–285. https://doi.org/10.1111/j.1469-8137.2009.03160.x
Apprill A, McNally S, Parsons R, Weber L (2015) Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 75:129–137. https://doi.org/10.3354/ame01753
Bardgett RD, Wardle DA (2010) Aboveground-belowground linkages: biotic interactions, ecosystem processes, and global change. Oxford University Press, Oxford
Bezemer TM, Van Dam NM (2005) Linking aboveground and belowground interactions via induced plant defenses. Trends Ecol Evol 20:617–624. https://doi.org/10.1016/j.tree.2005.08.006
Bulgarelli D, Schlaeppi K, Spaepen S, Van Themaat EVL, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838. https://doi.org/10.1146/annurev-arplant-050312-120106
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Carrión VJ, Perez-Jaramillo J, Cordovez V, Tracanna V, De Hollander M, Ruiz-Buck D, Mendes LW, Van Ijcken WFJ, Gomez-Exposito R, Elsayed SS, Mohanraju P, Arifah A, Van der Oost J, Paulson JN, Mendes R, van Wezel GP, Medema MH, Raaijmakers JM (2019) Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366:606–612. https://doi.org/10.1126/science.aaw9285
Carta C (1993) Chrysanthemum morifolium Ramat: a new host for Pseudomonas syringae. Phytopathol Mediterr 32:159–161
Cordovez V, Dini-Andreote F, Carrion VJ, Raaijmakers JM (2019) Ecology and evolution of plant microbiomes. Annu Rev Microbiol 73:69–88. https://doi.org/10.1146/annurev-micro-090817-062524
De Hollander M (2017) Nioo-knaw/hydra:1.3. https://doi.org/10.5281/zenodo.884028
Fitzpatrick CR, Copeland J, Wang PW, Guttman DS, Kotanen PM, Johnson MTJ (2018) Assembly and ecological function of the root microbiome across angiosperm plant species. Proc Natl Acad Sci U S A 115:E1157–E1165. https://doi.org/10.1073/pnas.1717617115
Foster ZSL, Sharpton TJ, Grünwald NJ (2017) Metacoder: an R package for visualization and manipulation of community taxonomic diversity data. PLoS Comput Biol 13:e1005404. https://doi.org/10.1371/journal.pcbi.1005404
Gadhave KR, Devlin PF, Ebertz A, Ross A, Gange AC (2018) Soil inoculation with Bacillus spp. modifies root endophytic bacterial diversity, evenness, and community composition in a context-specific manner. Microb Ecol 76:741–750. https://doi.org/10.1007/s00248-018-1160-x
Gweon HS, Oliver A, Taylor J, Booth T, Gibbs M, Read DS, Griffiths RI, Schonrogge K (2015) PIPITS: an automated pipeline for analyses of fungal internal transcribed spacer sequences from the Illumina sequencing platform. Methods Ecol Evol 6:973–980. https://doi.org/10.1111/2041-210X.12399
Hannula SE, Zhu F, Heinen R, Bezemer TM (2019) Foliar-feeding insects acquire microbiomes from the soil rather than the host plant. Nat Commun 10:1254. https://doi.org/10.1038/s41467-019-09284-w
Hannula SE, Ma H, Pérez-Jaramillo JE, Pineda A, Bezemer TM (2020) Structure and ecological function of the soil microbiome affecting plant–soil feedbacks in the presence of a soil-borne pathogen. Environ Microbiol 22:660–676. https://doi.org/10.1111/1462-2920.14882
Hardoim PR, Van Overbeek LS, Berg G, Pirttilä AM, Compant S, Campisano A, Döring M, Sessitsch A (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79:293–320. https://doi.org/10.1128/MMBR.00050-14
Herrera Paredes S, Gao T, Law TF, Finkel OM, Mucyn T, Teixeira PJPL, Salas González I, Feltcher ME, Powers MJ, Shank EA, Jones CD, Jojic V, Dangl JL, Castrillo G (2018) Design of synthetic bacterial communities for predictable plant phenotypes. PLoS Biol 16:e2003962. https://doi.org/10.1371/journal.pbio.2003962
Hubbard CJ, Li B, McMinn R, Brock MT, Maignien L, Ewers BE, Kliebenstein D, Weinig C (2019) The effect of rhizosphere microbes outweighs host plant genetics in reducing insect herbivory. Mol Ecol 28: 1801-1811. 1. https://doi.org/10.1111/mec.14989
Jones JB, Engelhard AW, Raju BC (1983) Outbreak of a stem necrosis on chrysanthemum incited by Pseudomonas cichorii in Florida. Plant Dis 67:431–433. https://doi.org/10.1094/PD-67-431
Kos M, Tuijl MAB, De Roo J, Mulder PPJ, Bezemer TM (2015) Species-specific plant-soil feedback effects on aboveground plant-insect interactions. J Ecol 103:904–914. https://doi.org/10.1111/1365-2745.12402
Kuźniar A, Włodarczyk K, Grzadziel J, Goraj W, Gałazka A, Woli’nska A (2020) Culture-independent analysis of an endophytic core microbiome in two species of wheat: Triticum aestivum L. (cv. ‘Hondia’) and the first report of microbiota in Triticum spelta L. (cv. ‘Rokosz’). Syst Appl Microbiol 43:126025. https://doi.org/10.1016/j.syapm.2019.126025
Lundberg DS, Yourstone S, Mieczkowski P, Jones CD, Dangl JL (2013) Practical innovations for high-throughput amplicon sequencing. Nat Methods 10:999–1002. https://doi.org/10.1038/nmeth.2634
Ma H, Pineda A, van der Wurff AWG, Raaijmakers C, Bezemer TM (2017) Plant–soil feedback effects on growth, defense and susceptibility to a soil-borne disease in a cut flower crop: species and functional group effects. Front Plant Sci 8:2127. https://doi.org/10.3389/fpls.2017.02127
Ma H, Pineda A, van der Wurff AWG, Bezemer TM (2018) Carry-over effects of soil inoculation on plant growth and health under sequential exposure to soil-borne diseases. Plant Soil 433:257–270. https://doi.org/10.1007/s11104-018-3837-9
Ma H, Pineda A, Hannula ES, Kielak AM, Setyarinia SN, Bezemer TM (2020) Steering root microbiomes of a commercial horticultural crop with plant-soil feedbacks. Appl Soil Ecol 150:103468. https://doi.org/10.1016/j.apsoil.2019.103468
Martínez G, Regente M, Jacobi S, Del Rio M, Pinedo M, De la Canal L (2017) Chlorogenic acid is a fungicide active against phytopathogenic fungi. Pestic Biochem Physiol 140:30–35. https://doi.org/10.1016/j.pestbp.2017.05.012
Muvea AM, Meyhofer R, Subramanian S, Poehling HM, Ekesi S, Maniania NK (2014) Colonization of onions by endophytic fungi and their impacts on the biology of Thrips tabaci. PLoS One 9:e108242. https://doi.org/10.1371/journal.pone.0108242
Pangesti N, Pineda A, Pieterse CMJ, Dicke M, van Loon JJA (2013) Two-way plant mediated interactions between root-associated microbes and insects: from ecology to mechanisms. Front Plant Sci 4:414. https://doi.org/10.3389/fpls.2013.00414
Pangesti N, Vandenbrande S, Pineda A, Dicke M, Raaijmakers JM, Van Loon JJA (2017) Antagonism between two root-associated beneficial Pseudomonas affect plant growth promotion and induced resistance against a leaf-chewing herbivore. FEMS Microbiol Ecol 93:1–8. https://doi.org/10.1093/femsec/fix038
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414. https://doi.org/10.1111/1462-2920.13023
Pattison Z, Rumble H, Tanner RA, Jin L, Gange AC (2016) Positive plant–soil feedbacks of the invasive Impatiens glandulifera and their effects on above-ground microbial communities. Weed Res 56:198–207. https://doi.org/10.1111/wre.12200
Pieterse CMJ, de Jonge R, Berendsen RL (2016) The soil-borne supremacy. Trends Plant Sci 21:171–173. https://doi.org/10.1016/j.tplants.2016.01.018
Pineda A, Kaplan I, Bezemer TM (2017) Steering soil microbiomes to suppress aboveground insect pests. Trends Plant Sci 22:770–778. https://doi.org/10.1016/j.tplants.2017.07.002
Pineda A, Kaplan I, Hannula SE, Ghanem W, Bezemer TM (2020) Conditioning the soil microbiome through plant-soil feedbacks to suppress aboveground pests. New Phytol 226:595–608. https://doi.org/10.1111/nph.16385
Poosakkannu A, Nissinen R, Kytöviita MM (2017) Native arbuscular mycorrhizal symbiosis alters foliar bacterial community composition. Mycorrhiza 27:801–810. https://doi.org/10.1007/s00572-017-0796-6
Schlaeppi K, Bulgarelli D (2014) The plant microbiome at work. Mol Plant Microbe In 28:212–217. https://doi.org/10.1094/MPMI-10-14-0334-FI
Tedersoo L, Anslan S, Bahram M, Põlme S, Riit T, Liiv I, Kõljalg U, Kisand V, Nilsson RH, Hildebrand F, Bork P, Abarenkov K (2015) Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. MycoKeys 10:1–43. https://doi.org/10.3897/mycokeys.10.4852
Van de Mortel JE, de Vos RC, Dekkers E, Pineda A, Guillod L, Bouwmeester K, van Loon JJ, DIcke M, Raaijmakers JM (2012) Metabolic and transcriptomic changes induced in Arabidopsis by the rhizobacterium Pseudomonas fluorescens SS101. Plant Physiol 160:2173–2188. https://doi.org/10.1104/pp.112.207324
Van der Putten WH, Bardgett RD, Bever JD, Bezemer TM, Casper BB, Fukami T, Kardol P, Klironomos JN, Kulmatiski A, Schweitzer JA, Suding KN, Van de Voorde TFJ, Wardle DA (2013) Plant-soil feedbacks: the past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
Wagner MR, Lundberg DS, Del Rio TG, Tringe SG, Dangl JL, Mitchell-Olds T (2016) Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat Commun 7:12151. https://doi.org/10.1038/ncomms12151
Wang M, Ruan W, Kostenko O, Carvalho S, Hannula SE, Mulder PPJ, Bu F, Putten WH, Bezemer TM (2019) Removal of soil biota alters soil feedback effects on plant growth and defense chemistry. New Phytol 221:1478–1491. https://doi.org/10.1111/nph.15485
Wells ML, Gitaitis RD, Sanders FH (2002) Association of tobacco thrips, Frankliniella fusca (Thysanoptera: Thripidae) with two species of bacteria of the genus Pantoea. Ann Entomol Soc Am 95:719–723. https://doi.org/10.1603/0013-8746(2002)095[0719:AOTTFF]2.0.CO;2
Wubs ERJ, Van der Putten WH, Bosch M, Bezemer TM (2016) Soil inoculation steers restoration of terrestrial ecosystems. Nat Plants 2:16107. https://doi.org/10.1038/nplants.2016.107
Acknowledgements
This work was funded by the Netherlands Organization for Scientific Research (NWO Groen grant no. 870.15.080 and NWO VICI grant no. 865.14.006). We thank A. Post from Deliflor for providing chrysanthemum cuttings, T. Snoeren from Hazera for providing thrips to start our own culture, and two anonymous reviewers for constructive comments on a earlier version. Sequencing of the samples was performed in collaboration with McGill University and Génome Québec Innovation Centre, Canada. This is publication 7014 of NIOO-KNAW.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Birgit Mitter.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1293 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pangesti, N., Pineda, A., Hannula, S.E. et al. Soil inoculation alters the endosphere microbiome of chrysanthemum roots and leaves. Plant Soil 455, 107–119 (2020). https://doi.org/10.1007/s11104-020-04655-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-020-04655-5