Abstract
Aims
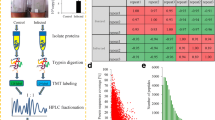
Crops often encounter a soil deficiency of nitrogen (N), the most important macronutrient for plants; however, the molecular mechanism of plant responses to N deficiency remains unclear. In this study, proteome-level changes that occur in bread wheat seedlings suffering from N deficiency were investigated to identify some N deficiency-responsive protein species in bread wheat.
Methods
We utilized isobaric tagging for relative and absolute quantification (iTRAQ) to measure changes in the proteome patterns of N-deficient wheat seedlings and validated the role of abscisic acid (ABA) using the barley stripe mosaic virus-induced gene-silencing (BSMV-VIGS) method.
Results
A total of 1515 N deficiency–responsive protein species were successfully identified in both root and leaf tissues of wheat seedlings suffering from 8-d N deficiency. Of these, abundance of wheat zeaxanthin epoxidase (TaZEP), a key ABA synthesis-related enzyme, was significantly upregulated, and the endogenous ABA contents also markedly increased. After TaZEP gene was further silenced using BSMV-VIGS method, BSMV-VIGS-TaZEP infected wheat seedlings showed enhanced sensitivity to N deficiency, suggesting silencing of TaZEP gene decreased the tolerance to N deficiency remarkably.
Conclusion
Our results identified some N deficiency-responsive protein species and revealed the role of ABA in wheat responses to N deficiency.






Similar content being viewed by others
Abbreviations
- ABA:
-
abscisic acid
- BSMV-VIGS:
-
barley stripe mosaic virus-virus induced gene-silencing
- CTK:
-
cytokinin
- DW:
-
dry weight
- ETH:
-
ethylene
- FW:
-
fresh weight
- IAA:
-
auxin
- iTRAQ:
-
isobaric tagging for relative and absolute quantification
- JA:
-
jasmonic acid
- MW:
-
molecular weights
- pI:
-
isoelectric points
- qRT-PCR:
-
quantitative real-time PCR
- SA:
-
salicylic acid
- ZEP:
-
zeaxanthin epoxidase
References
Agrawal GK, Yamazaki M, Kobayashi M, Hirochika R, Miyao A, Hirochika H (2001) Screening of the rice viviparous mutants generated by endogenous retrotransposon Tos17 insertion. Tagging of a zeaxanthin epoxidase gene and a novel OsTATC gene. Plant Physiol 125:1248–1257
Borisjuk N, Kishchenko O, Eliby S, Schramm C, Anderson P, Jatayev S, Kurishbayev A, Shavrukov Y (2019) Genetic modification for wheat improvement: from transgenesis to genome editing. Biomed Res Int 2019:6216304
Brenchley R, Spannagl M, Pfeifer M, Barker GL, D'Amore R, Allen AM et al (2012) Analysis of the bread wheat genome using whole–genome shotgun sequencing. Nature 491:705–710
Cai H, Lu Y, Xie W, Zhu T, Lian X (2012) Transcriptome response to nitrogen starvation in rice. J Biosci 37:731–747
Cao A, Xing L, Wang X, Yang X, Wang W, Sun Y, Qian C, Ni J, Chen Y, Liu D, Wang X, Chen P (2011) Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc Natl Acad Sci U S A 108:7727–7732
Chao WS, Doğramaci M, Horvath DP, Anderson JV, Foley ME (2016) Phytohormone balance and stress-related cellular responses are involved in the transition from bud to shoot growth in leafy spurge. BMC Plant Biol 16:47
Criado MV, Roberts IN, Echeverria M, Barneix AJ (2007) Plant growth regulators and induction of leaf senescence in nitrogen–deprived wheat plants. J Plant Growth Regul 26:301–307
Cuming AC, Stevenson SR (2015) From pond slime to rain forest: the evolution of ABA signalling and the acquisition of dehydration tolerance. New Phytol 206:5–7
Curci PL, Cigliano RA, Zuluaga DL, Janni M, Sanseverino W, Sonnante G (2017) Transcriptomic response of durum wheat to nitrogen starvation. Sci Rep-UK 7:1176
Deng G, Liu L, Zhong X, Lao C, Wang H, Wang B, Zhu C, Shah F, Peng D (2014) Comparative proteome analysis of the response of ramie under N, P and K deficiency. Planta 239:1175–1186
Elberse IAM, van Damme JMM, Tienderen PHV (2003) Plasticity of growth characteristics in wild barley (Hordeum spontaneum) in response to nutrient limitation. J Ecol 91:371–382
Feldman M, Levy AA, Fahima T, Korol A (2012) Genomic asymmetry in allopolyploid plants: wheat as a model. J Exp Bot 63:5045–5059
Findlay GD, MacCoss MJ, Swanson WJ (2009) Proteomic discovery of previously unannotated, rapidly evolving seminal fluid genes in Drosophila. Genome Res 19:886–896
Gelli M, Duo Y, Konda AR, Zhang C, Holding D, Dweikat I (2014) Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genomics 15:60–66
Grobei MA, Qeli E, Brunner E, Rehrauer H, Zhang R, Roschitzki B, Basler K, Ahrens CH, Grossniklaus U (2009) Deterministic protein inference for shotgun proteomics data provides new insights into Arabidopsis pollen development and function. Genome Res 19:1786–1800
Grün A, Buchner P, Broadley MR, Hawkesford MJ (2018) Identification and expression profiling of Pht1 phosphate transporters in wheat in controlled environments and in the field. Plant Biol 20:374–389
Guo T, Xuan H, Yang Y, Wang L, Wei L, Wang Y, Kang G (2014) Transcription analysis of genes encoding the wheat root transporter NRT1 and NRT2 families during nitrogen starvation. J Plant Growth Regul 33:837–848
Hakeem KR, Ahmad A, Iqbal M, Gucel S, Ozturk M (2011) Nitrogen-efficient rice cultivars can reduce nitrate pollution. Environ Sci Pollut Res 18:1184–1193
Han YL, Liao JY, Yu Y, Song HX, Rong N, Guan CY, Lepo JE, Ismail AM, Zhang ZH (2017) Exogenous abscisic acid promotes the nitrogen use efficiency of Brassica napus by increasing nitrogen remobilization in the leaves. J Plant Nutr 40:2540–2549
He M, Zhu C, Dong K, Zhang T, Cheng Z, Li J, Yan Y (2015) Comparative proteome analysis of embryo and endosperm reveals central differential expression proteins involved in wheat seed germination. BMC Plant Biol 15:1–17
He X, Ma H, Zhao X, Nie S, Li Y, Zhang Z et al (2016a) Comparative RNA-seq analysis reveals that regulatory network of maize root development controls the expression of genes in response to N stress. PLoS One 11:3
He L, Zhang H, Zhang Y, Song X, Feng W, Kang G, Wang C, Guo T (2016b) Estimating canopy leaf nitrogen concentration in winter wheat based on multi-angular hyperspectral remote sensing. Eur J Agron 73:170–185
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Calif Agric Exp Stn Circ 347:1–32
Hu G, Jin K, Yoo MJ, Grupp K, Chen S, Wendel JF (2013) Proteomic profiling of developing cotton fibers from wild and domesticated Gossypium barbadense. New Phytol 200:570–582
Hu J, Rampitsch C, Bykova NV (2015) Advances in plant proteomics toward improvement of crop productivity and stress resistance. Front Plant Sci 6:209
International Wheat Genome Sequencing Consortium (IWGSC) (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:661
Jia J, Zhao S, Kong X, Li Y, Zhao G, He W et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496:91–95
Jin X, Li W, Hu D, Shi X, Zhang X, Zhang F, Fu Z, Ding D, Liu Z, Tang J (2015) Biological responses and proteomic changes in maize seedlings under nitrogen deficiency. Plant Mol Biol Rep 33:490–504
Kang G, Li G, Wang L, Wei L, Yang Y, Wang P, Yang Y, Wang Y, Feng W, Wang C, Guo T (2015) Hg–responsive proteins identified in wheat seedlings using iTRAQ analysis and the role of ABA in Hg stress. J Proteome Res 14:249–267
Kosová K, Vítámvás P, Prášil IT, Renaut J (2011) Plant proteome changes under abiotic stress–contribution of proteomics studies to understanding plant stress response. J Proteome 74:1301–1322
Kosová K, Vítámvás P, Urban MO, Prášil IT, Renaut J (2018) Plant abiotic stress proteomics: the major factors determining alterations in cellular proteome. Front Plant Sci 9:122
Krapp A, Berthomé R, Orsel M, Mercey-Boutet S, Yu A, Castaings L, Elftieh S, Major H, Renou JP, Daniel-Vedele F (2011) Arabidopsis roots and shoots show distinct temporal adaptation patterns toward nitrogen starvation. Plant Physiol 157:1255–1282
Kuang Q, Zhang S, Wu P, Chen Y, Li M, Jiang H, Wu G (2017) Global gene expression analysis of the response of physic nut (Jatropha curcas L.) to medium- and long-term nitrogen deficiency. PLoS One 12:e0182700
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60
Li G, Peng X, Xuan H, Wei L, Yang Y, Guo T, Kang G (2013) Proteomic analysis of leaves and roots of common wheat (Triticum aestivum L.) under copper-stress conditions. J Proteome Res 12:4846–4861
Li A, Liu D, Wu J, Zhao X, Hao M, Geng S, Yan J, Jiang X, Zhang L, Wu J, Yin L, Zhang R, Wu L, Zheng Y, Mao L (2014) mRNA and small RNA transcriptomes reveal insights into dynamic homoeolog regulation of allopolyploid heterosis in nascent hexaploid wheat. Plant Cell 26:1878–1900
Li G, Wu Y, Liu G, Xiao X, Wang P, Gao T, Xu M, Han Q, Wang Y, Guo T, Kang G (2017) Large-scale proteomics combined with transgenic experiments demonstrates an important role of jasmonic acid in potassium deficiency response in wheat and rice. Mol Cell Proteomics 16:1889–1905
Liang C, Tian J, Liao H (2013) Proteomics dissection of plant responses to mineral nutrient deficiency. Proteomics 13:624–636
Ling HQ, Zhao S, Liu D, Wan J, Sun H, Zhang C et al (2013) Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 496:87–90
Liu G, Wu Y, Xu M, Gao T, Wang P, Wang L, Guo T, Kang G (2016) Virus-induced gene silencing identifies an important role of the TaRSR1 transcription factor in starch synthesis in bread wheat. Int J Mol Sci 17:1557
Ma TL, Wu WH, Wang Y (2012) Transcriptome analysis of rice root responses to potassium deficiency. BMC Plant Biol 12:161
Ma C, Zhou J, Chen G, Bian Y, Lv D, Li X, Wang Z, Yan Y (2014) iTRAQ-based quantitative proteome and phosphoprotein characterization reveals the central metabolism changes involved in wheat grain development. BMC Genomics 15:1–20
Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y (2008) RNA-Seq: An assessment of technical reproducibility and comparison with gene expression arrays. Genome Res 18:1509–1517
Meise P, Jozefowicz AM, Uptmoor R, Mock HP, Ordon F, Schum A (2017) Comparative shoot proteome analysis of two potato (Solanum tuberosum L.) genotypes contrasting in nitrogen deficiency responses in vitro. J Proteome 166:68–82
Møller ALB, Pedas P, Andersen B, Svensson B, Schjoerring JK, Finnie C (2011) Responses of barley root and shoot proteomes to long-term nitrogen deficiency, short-term nitrogen starvation and ammonium. Plant Cell Environ 34:2024–2037
Nawaz MA, Chen C, Shireen F, Zheng Z, Sohail H, Afzal M, Ali MA, Bie Z, Huang Y (2018) Genome-wide expression profiling of leaves and roots of watermelon in response to low nitrogen. BMC Genomics 19:456
Nazir M, Pandey R, Siddiqi TO, Ibrahim MM, Qureshi MI, Abraham G, Vengavasi K, Ahamad A (2015) Nitrogen-deficiency stress induces protein expression differentially in low-N tolerant and low-N sensitive maize genotypes. Front Plant Sci 7:298
Oka M, Shimoda Y, Sato N, Inoue J, Yamazaki T, Shimomura N, Fujiyama H (2012) Abscisic acid substantially inhibits senescence of cucumber plants (Cucumis sativus) grown under low nitrogen conditions. J Plant Physiol 169:789–796
Qin L, Walk TC, Han P, Chen L, Zhang S, Li Y, Hu X, Xie L, Yang Y, Liu J, Lu X, Yu C, Tian J, Shaff JE, Kochian LV, Liao X, Liao H (2019) Adaption of roots to nitrogen deficiency revealed by 3D quantification and proteomic analysis. Plant Physiol 179:329–347
Quan X, Zeng J, Ye L, Chen G, Han Z, Munawar J, Zhang G (2016) Transcriptome profiling analysis for two Tibetan wild barley genotypes in responses to low nitrogen. BMC Plant Biol 16:30
Ristova D, Carré C, Pervent M, Medici A, Kim GJ, Scalia D et al (2016) Combinatorial interaction network of transcriptomic and phenotypic responses to nitrogen and hormones in the Arabidopsis thaliana root. Sci Signal 9:451
Schlüter H, Apweiler R, Holzhütter HG, Jungblut PR (2009) Finding one's way in proteomics: a protein species nomenclature. Chem Cent J 3:11
Scofield SR, Huang L, Brandt AS, Gill BS (2005) Development of a virus-induced gene-silencing system for hexaploid wheat and its use in functional analysis of the Lr21-mediated leaf rust resistance pathway. Plant Physiol 138:2165–2173
Shi W, Xu W, Li S, Zhao X, Dong G (2010) Responses of two rice cultivars differing in seedling–stage nitrogen use efficiency to growth under low–nitrogen conditions. Plant Soil 326:291–302
Song J, Jiang L, Jameso PE (2012) Co-ordinate regulation of cytokinin gene family members during flag leaf and reproductive development in wheat. BMC Plant Biol 12:78
Torabi S, Wissuwa M, Heidari M, Naghavi MR, Gilany K, Hajirezaei MR, Omidi M, Yazdi-Samadi B, Ismail AM, Salekdeh GH (2009) A comparative proteome approach to decipher the mechanism of rice adaptation to phosphorus deficiency. Proteomics 9:159–170
Tufan HA, Stefanato FL, McGrann GRD, MacCormack R, Boyd LA (2011) The barley stripe mosaic virus system used for virus-induced gene silencing in cereals differentially affects susceptibility to fungal pathogens in wheat. J Plant Physiol 168:990–994
Vogel C, Marcotte EM (2012) Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet 13:227–232
Wang X, Bian Y, Cheng K, Zou H, Sun S, He J (2012) A comprehensive differential proteomic study of nitrate deprivation in Arabidopsis reveals complex regulatory networks of plant nitrogen responses. J Proteome Res 11:2301–2315
Wei Z, Zeng X, Qin C, Wang Y, Bai L, Xu Q et al (2016) Comparative transcriptome analysis revealed genes commonly responsive to varied nitrate stress in leaves of tibetan hulless barley. Front Plant Sci 7:298
Wisniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6:359–362
Wu H, Sparks C, Amoah B, Jones HD (2003) Factors influencing successful Agrobacterium-mediated genetic transformation of wheat. Plant Cell Rep 21:659–668
Yang W, Yoon J, Choi H, Fan Y, Chen R, An G (2015) Transcriptome analysis of nitrogen-starvation responsive genes in rice. BMC Plant Biol 15:31
Zhang N, Huo W, Zhang L, Chen F, Cui D (2016) Identification of winter-responsive proteins in bread wheat using proteomics analysis and virus-induced gene silencing. Mol Cell Proteomics 15:2954–2969
Zhang Y, Zhou Y, Chen S, Liu J, Fan K, Li Z, Liu Z, Lin W (2019) Gibberellins play dual roles in response to phosphate starvation of tomato seedlings, negatively in shoots but positively in roots. J Plant Physiol 234–245:145–153
Zhao T, Zhao S, Chen H, Zhao Q, Hu Z, Hou B, Xia G (2006) Transgenic wheat frogeny resistant to powdery mildew generated by Agrobacterium inoculumto the basal portion of wheat seedling. Plant Cell Rep 25:1199–1204
Acknowledgements
This study was financially supported by the Science and Technology Innovation Program for Increase in Yield and Efficiency of Food Crop (2016YFD0300105) and the National Key Technology Support Program (2015BAD26B01).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Ad C. Borstlap.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Table S1
(DOC 33 kb)
Supplementary Table S2
(XLS 3280 kb)
Supplementary Table S3
(XLS 3016 kb)
Supplementary Table S4
(XLS 1956 kb)
Supplementary Table S5
(XLS 535 kb)
Supplementary Table S6
(XLS 90 kb)
Supplementary Table S7
(DOC 32 kb)
Supplementary Fig. S1
(DOC 41 kb)
Supplementary Fig. S2
(DOC 409 kb)
Rights and permissions
About this article
Cite this article
Kang, G., Wu, Y., Li, G. et al. Proteomics combined with BSMV-VIGS methods identified some N deficiency-responsive protein species and ABA role in wheat seedling. Plant Soil 444, 177–191 (2019). https://doi.org/10.1007/s11104-019-04260-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-019-04260-1