Abstract
It is widely accepted that Ca2+ATPase family proteins play important roles in plant environmental stress responses. However, up to now, most researches are limited in the reference plants Arabidopsis and rice. The function of Ca2+ATPases from non-reference plants was rarely reported, especially its regulatory role in carbonate alkaline stress responses. Hence, in this study, we identified the P-type II Ca2+ATPase family genes in soybean genome, determined their chromosomal location and gene architecture, and analyzed their amino acid sequence and evolutionary relationship. Based on above results, we pointed out the existence of gene duplication for soybean Ca2+ATPases. Then, we investigated the expression profiles of the ACA subfamily genes in wild soybean (Glycine soja) under carbonate alkaline stress, and functionally characterized one representative gene GsACA1 by using transgenic alfalfa. Our results suggested that GsACA1 overexpression in alfalfa obviously increased plant tolerance to both carbonate alkaline and neutral salt stresses, as evidenced by lower levels of membrane permeability and MDA content, but higher levels of SOD activity, proline concentration and chlorophyll content under stress conditions. Taken together, for the first time, we reported a P-type II Ca2+ATPase from wild soybean, GsACA1, which could positively regulate plant tolerance to both carbonate alkaline and neutral salt stresses.










Similar content being viewed by others
References
Bates L, Waldren R, Teare I (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Baxter I, Tchieu J, Sussman MR, Boutry M, Palmgren MG, Gribskov M, Harper JF, Axelsen KB (2003) Genomic comparison of P-type ATPase ion pumps in Arabidopsis and rice. Plant Physiol 132:618–628
Baxter A, Mittler R, Suzuki N (2014) ROS as key players in plant stress signalling. J Exp Bot 65:1229–1240
Bonza MC, De Michelis MI (2011) The plant Ca2+-ATPase repertoire: biochemical features and physiological functions. Plant Biol (Stuttg) 13:421–430
Bose J, Pottosin II, Shabala SS, Palmgren MG, Shabala S (2011) Calcium efflux systems in stress signaling and adaptation in plants. Front Plant Sci 2:85
Boursiac Y, Harper JF (2007) The origin and function of calmodulin regulated Ca2+ pumps in plants. J Bioenerg Biomembr 39:409–414
Cao J, Xu D, Wang D, Wu R, Zhang L, Zhu H, He Q, Yang B (2009) ROS-driven Akt dephosphorylation at Ser-473 is involved in 4-HPR-mediated apoptosis in NB4 cells. Free Radic Biol Med 47:536–547
Cerana M, Bonza MC, Harris R, Sanders D, De Michelis MI (2006) Abscisic acid stimulates the expression of two isoforms of plasma membrane Ca2+-ATPase in Arabidopsis thaliana seedlings. Plant Biol (Stuttg) 8:572–578
Chung WS, Lee SH, Kim JC, Heo WD, Kim MC, Park CY, Park HC, Lim CO, Kim WB, Harper JF, Cho MJ (2000) Identification of a calmodulin-regulated soybean Ca2+-ATPase (SCA1) that is located in the plasma membrane. Plant Cell 12:1393–1407
Dean WL, Tanford C (1977) Reactivation of lipid-depleted Ca2+-ATPase by a nonionic detergent. J Biol Chem 252:3551–3553
Ding M, Hou P, Shen X, Wang M, Deng S, Sun J, Xiao F, Wang R, Zhou X, Lu C, Zhang D, Zheng X, Hu Z, Chen S (2010) Salt-induced expression of genes related to Na+/K+ and ROS homeostasis in leaves of salt-resistant and salt-sensitive poplar species. Plant Mol Biol 73:251–269
Dodd AN, Kudla J, Sanders D (2010) The language of calcium signaling. Annu Rev Plant Biol 61:593–620
DuanMu H, Wang Y, Bai X, Cheng S, Deyholos MK, Wong GK, Li D, Zhu D, Li R, Yu Y, Cao L, Chen C, Zhu Y (2015) Wild soybean roots depend on specific transcription factors and oxidation reduction related genesin response to alkaline stress. Funct Integr Genomics 15:651–660
Gautier R, Douguet D, Antonny B, Drin G (2008) HELIQUEST: a web server to screen sequences with specific alpha-helical properties. Bioinformatics 24:2101–2102
Ge Y, Li Y, Zhu YM, Bai X, Lv DK, Guo D, Ji W, Cai H (2010) Global transcriptome profiling of wild soybean (Glycine soja) roots under NaHCO3 treatment. BMC Plant Biol 10:153
George L, Romanowsky SM, Harper JF, Sharrock RA (2008) The ACA10 Ca2+-ATPase regulates adult vegetative development and inflorescence architecture in Arabidopsis. Plant Physiol 146:716–728
Giannopolitis CN, Ries SK (1977) Superoxide dismutases: I. Occurrence in higher plants. Plant Physiol 59:309–314
Guo R, Yang Z, Li F, Yan C, Zhong X, Liu Q, Xia X, Li H, Zhao L (2015) Comparative metabolic responses and adaptive strategies of wheat (Triticum aestivum) to salt and alkali stress. BMC Plant Biol 15:170
Hanikenne M, Baurain D (2013) Origin and evolution of metal P-type ATPases in Plantae (Archaeplastida). Front Plant Sci 4:544
Haro R, Garciadeblas B, Rodriguez-Navarro A (1991) A novel P-type ATPase from yeast involved in sodium transport. FEBS Lett 291:189–191
He X, Sambe MA, Zhuo C, Tu Q, Guo Z (2015) A temperature induced lipocalin gene from Medicago falcata (MfTIL1) confers tolerance to cold and oxidative stress. Plant Mol Biol 87:645–654
Huda KM, Banu MS, Garg B, Tula S, Tuteja R, Tuteja N (2013a) OsACA6, a P-type IIB Ca2+ATPase promotes salinity and drought stress tolerance in tobacco by ROS scavenging and enhancing the expression of stress-responsive genes. Plant J 76:997–1015
Huda KM, Banu MS, Tuteja R, Tuteja N (2013b) Global calcium transducer P-type Ca2+-ATPases open new avenues for agriculture by regulating stress signalling. J Exp Bot 64:3099–3109
Hwang I, Harper JF, Liang F, Sze H (2000a) Calmodulin activation of an endoplasmic reticulum-located calcium pump involves an interaction with the N-terminal autoinhibitory domain. Plant Physiol 122:157–168
Hwang I, Sze H, Harper JF (2000b) A calcium-dependent protein kinase can inhibit a calmodulin-stimulated Ca2+ pump (ACA2) located in the endoplasmic reticulum of Arabidopsis. Proc Natl Acad Sci USA 97:6224–6229
Ishitani M, Xiong L, Lee H, Stevenson B, Zhu JK (1998) HOS1, a genetic locus involved in cold-responsive gene expression in Arabidopsis. Plant Cell 10:1151–1161
Iwano M, Igarashi M, Tarutani Y, Kaothien-Nakayama P, Nakayama H, Moriyama H, Yakabe R, Entani T, Shimosato-Asano H, Ueki M, Tamiya G, Takayama S (2014) A pollen coat-inducible autoinhibited Ca2+-ATPase expressed in stigmatic papilla cells is required for compatible pollination in the Brassicaceae. Plant Cell 26:636–649
Kamrul Huda KM, Yadav S, Akhter Banu MS, Trivedi DK, Tuteja N (2013) Genome-wide analysis of plant-type II Ca2+ATPases gene family from rice and Arabidopsis: potential role in abiotic stresses. Plant Physiol Biochem 65:32–47
Kamrul Huda KM, Akhter Banu MS, Yadav S, Sahoo RK, Tuteja R, Tuteja N (2014) Salinity and drought tolerant OsACA6 enhances cold tolerance in transgenic tobacco by interacting with stress-inducible proteins. Plant Physiol Biochem 82:229–238
Kichtenthaler H, Wellburn A (1983) Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvent. Biochem Soc Trans 603:591–593
Kleist T, Luan S (2015) Constant change: dynamic regulation of membrane transport by calcium signaling networks keeps plants in tune with their environment. Plant Cell Environ. doi:10.1111/pce.12599
Kudla J, Batistic O, Hashimoto K (2010) Calcium signals: the lead currency of plant information processing. Plant Cell 22:541–563
Leclercq J, Martin F, Sanier C, Clement-Vidal A, Fabre D, Oliver G, Lardet L, Ayar A, Peyramard M, Montoro P (2012) Over-expression of a cytosolic isoform of the HbCuZnSOD gene in Hevea brasiliensis changes its response to a water deficit. Plant Mol Biol 80:255–272
Lee SM, Kim HS, Han HJ, Moon BC, Kim CY, Harper JF, Chung WS (2007) Identification of a calmodulin-regulated autoinhibited Ca2+-ATPase (ACA11) that is localized to vacuole membranes in Arabidopsis. FEBS Lett 581:3943–3949
Limonta M, Romanowsky S, Olivari C, Bonza MC, Luoni L, Rosenberg A, Harper JF, De Michelis MI (2014) ACA12 is a deregulated isoform of plasma membrane Ca2+-ATPase of Arabidopsis thaliana. Plant Mol Biol 84:387–397
Lucca N, Leon G (2012) Arabidopsis ACA7, encoding a putative auto-regulated Ca2+-ATPase, is required for normal pollen development. Plant Cell Rep 31:651–659
Mao GL, Xu X, Zeng J, Yue ZH, Yang SJ (2012) Effects of desulfurization waste on calcium distribution, Ca2+-ATPase activity, and antioxidant characteristics of rice leaf under alkali stress. Ying Yong Sheng Tai Xue Bao 23:363–368
Palmgren MG, Nissen P (2011) P-type ATPases. Annu Rev Biophys 40:243–266
Pedersen CN, Axelsen KB, Harper JF, Palmgren MG (2012) Evolution of plant P-type ATPases. Front Plant Sci 3:31
Peever TL, Higgins VJ (1989) Electrolyte leakage, lipoxygenase, and lipid peroxidation induced in tomato leaf tissue by specific and nonspecific elicitors from Cladosporium fulvum. Plant Physiol 90:867–875
Pittman JK, Hirschi KD (2003) Don’t shoot the (second) messenger: endomembrane transporters and binding proteins modulate cytosolic Ca2+ levels. Curr Opin Plant Biol 6:257–262
Qiu Y, Xi J, Du L, Suttle JC, Poovaiah BW (2012) Coupling calcium/calmodulin-mediated signaling and herbivore-induced plant response through calmodulin-binding transcription factor AtSR1/CAMTA3. Plant Mol Biol 79:89–99
Qudeimat E, Faltusz AM, Wheeler G, Lang D, Holtorf H, Brownlee C, Reski R, Frank W (2008) A PIIB-type Ca2+-ATPase is essential for stress adaptation in Physcomitrella patens. Proc Natl Acad Sci USA 105:19555–19560
Schiott M, Romanowsky SM, Baekgaard L, Jakobsen MK, Palmgren MG, Harper JF (2004) A plant plasma membrane Ca2+ pump is required for normal pollen tube growth and fertilization. Proc Natl Acad Sci USA 101:9502–9507
Seo PJ, Park MJ, Park CM (2013) Alternative splicing of transcription factors in plant responses to low temperature stress: mechanisms and functions. Planta 237:1415–1424
Shafi A, Chauhan R, Gill T, Swarnkar MK, Sreenivasulu Y, Kumar S, Kumar N, Shankar R, Ahuja PS, Singh AK (2015) Expression of SOD and APX genes positively regulates secondary cell wall biosynthesis and promotes plant growth and yield in Arabidopsis under salt stress. Plant Mol Biol 87:615–631
Shukla D, Huda KM, Banu MS, Gill SS, Tuteja R, Tuteja N (2014) OsACA6, a P-type 2B Ca2+ATPase functions in cadmium stress tolerance in tobacco by reducing the oxidative stress load. Planta 240:809–824
Singh A, Kanwar P, Yadav AK, Mishra M, Jha SK, Baranwal V, Pandey A, Kapoor S, Tyagi AK, Pandey GK (2014) Genome-wide expressional and functional analysis of calcium transport elements during abiotic stress and development in rice. FEBS J 281:894–915
Spalding EP, Harper JF (2011) The ins and outs of cellular Ca2+ transport. Curr Opin Plant Biol 14:715–720
Sun M, Sun X, Zhao Y, Zhao C, Duanmu H, Yu Y, Ji W, Zhu Y (2014) Ectopic expression of GsPPCK3 and SCMRP in Medicago sativa enhances plant alkaline stress tolerance and methionine content. PLoS ONE 9:e89578
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tuteja N, Mahajan S (2007) Calcium signaling network in plants: an overview. Plant Signal Behav 2:79–85
Wang H, Wu Z, Han J, Zheng W, Yang C (2012) Comparison of ion balance and nitrogen metabolism in old and young leaves of alkali-stressed rice plants. PLoS ONE 7:e37817
Willems E, Leyns L, Vandesompele J (2008) Standardization of real-time PCR gene expression data from independent biological replicates. Anal Biochem 379:127–129
Yamada N, Theerawitaya C, Cha-um S, Kirdmanee C, Takabe T (2014) Expression and functional analysis of putative vacuolar Ca2+-transporters (CAXs and ACAs) in roots of salt tolerant and sensitive rice cultivars. Protoplasma 251:1067–1075
Yap KL, Kim J, Truong K, Sherman M, Yuan T, Ikura M (2000) Calmodulin target database. J Struct Funct Genomics 1:8–14
Zhang YM, Zhang HM, Liu ZH, Li HC, Guo XL, Li GL (2015) The wheat NHX antiporter gene TaNHX2 confers salt tolerance in transgenic alfalfa by increasing the retention capacity of intracellular potassium. Plant Mol Biol 87:317–327
Zhao Y, Lu Z, He L (2014) Effects of saline-alkaline stress on seed germination and seedling growth of Sorghum bicolor (L.) Moench. Appl Biochem Biotechnol 173:1680–1691
Acknowledgments
We thank members of the lab for discussions and comments on this manuscript. This work was supported by the National Natural Science Foundation of China (31171578), National basic scientific talent training fund projects (J1210069), the National Natural Science Foundation of China (31500204), the Natural Science Foundation of Heilongjiang Province (C2015035), and Graduate student innovation research project of Northeast Agricultural University (yjscx14056).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2015_426_MOESM1_ESM.tif
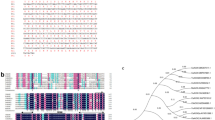
Supplementary Fig. S1 Gene architecture and phylogenetic analyses of soybean ECAs. a Phylogenetic relationship among ECAs in soybean. b Representation of intron-exon and domain structures in ECAs proteins. The intron-exon structures were from Phytozome website. The domains found in the ECA subfamily contained cation transporter/ATPase N-terminus (red), E1-E2 ATPase (orange), haloacid dehalogenase-like hydrolase (purple) and cation transporter/ATPase C-terminus (blue). (TIFF 84 kb)
11103_2015_426_MOESM2_ESM.tif
Supplementary Fig. S2 Multiple amino acid sequence alignment of ECAs from soybean and Arabidopsis. The figure showed the conserved motifs (red boxes) and the predicted transmembrane domains (designated as M1-M10) (marked by red lines on the top). The amino acid sequences were aligned by the ClustalX software. The existence and position of transmembrane domains were verified and marked according to previous study (Chung et al. 2000). (TIFF 1817 kb)
11103_2015_426_MOESM3_ESM.tif
Supplementary Fig. S3 Expression patterns of ACA subfamily genes in Glycine soja under 50 mM NaHCO3 (pH 8.5) treatment based on the microarray data. a Expression patterns of ACAs in Glycine soja roots. b Expression patterns of ACAs in Glycine soja leaves. The color saturation reflected the log2 fold change (LogFC) to 0 h for each gene. Green color indicated higher transcript levels than 0 h, whereas green color meant lower transcript levels. (TIFF 154 kb)
Rights and permissions
About this article
Cite this article
Sun, M., Jia, B., Cui, N. et al. Functional characterization of a Glycine soja Ca2+ATPase in salt–alkaline stress responses. Plant Mol Biol 90, 419–434 (2016). https://doi.org/10.1007/s11103-015-0426-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0426-7