Abstract
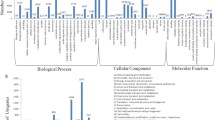
Pearl millet (Pennisetum glaucum), used as forage and grain crop is a stress tolerant species. Here we identify differentially regulated transcripts in response to abiotic (salinity, drought and cold) stresses from subtracted cDNA libraries by single-pass sequencing of cDNA clones. A total of 2,494 EST sequences were clustered and assembled into a collection of 1,850 unique sequences with 224 contigs and 1,626 singleton sequences. By sequence comparisons the putative functions of many ESTs could be assigned. Genes with stress related functions include those involved in cellular defense against abiotic stresses and transcripts for proteins involved in stress response signaling and transcription in addition to ESTs encoding unknown functions. These provide new candidate genes for investigation to elucidate their role in abiotic stress. The relative mRNA abundance of 38 selected genes, quantified using real time quantitative RT-PCR, demonstrated the existence of a complex gene regulatory network that differentially modulates gene expression in a kinetics-specific manner in response to different abiotic stresses. Notably, housekeeping and non-target genes were effectively reduced in these subtracted cDNA libraries constructed. These EST sequences are a rich source of stress-related genes and reveal a major part of the stress-response transcriptome that will provide the foundation for further studies into understanding Pennisetum’s adaptability to harsh environmental conditions.



Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Arimura G, Huber DP, Bohlmann J (2004) Forest tent caterpillars (Malacosoma disstria) induce local and systemic diurnal emissions of terpenoid volatiles in hybrid poplar (Populus trichocarpa x deltoids): cDNA cloning, functional characterization, and patterns of gene expression of (-)-germacrene D synthase, PtdTPS1. Plant J 37:603–616
Asamizu E, Nakamura Y, Sato S, Tabata S (2000) A large scale analysis of cDNA in Arabidopsis thaliana: generation of 12028 non redundant expressed sequence tags from normalized and size-selected cDNA library. DNA Res 7:175–180
Borkird C, Simoens C, Villarroel R, Vanmontagu M (1991) Gene-expression associated with water-stress adaptation of rice cells and identification of 2 genes as hsp 70 and ubiquitin. Physiol Plant 82:449–457
Buntjer J, Ruiter MD, Vallins B, Minarik M, Mahtani M, Schaik RV, Vos P, Eijk MV (2001) High throughput analysis of AFLP fragments on the MegaBASE capillary sequencer. In: Plant and animal genome IX conference. January 13–17, 2001, San Diego, CA, USA
Cabané M, Calvet P, Vincens P, Boudet AM (1993) Characterization of chilling-acclimation-related proteins in soybean and identification of one as a member of the heat shock protein (HSP70) family. Planta 190:346–353
Chen W, Provart NJ, Glazebrook J, Katagiri F, Chang HS, Eulgem T, Mauch F, Luan S, Zou G, Whitnam SA, Budworth PR, Tao Y, Xie Z, Chen X, Lam S, Kreps JA, Harper JF, Si-Ammour A, Mauch-Mani B, Heilein M, Kobayashi L, Hohn T, Dangl JL, Wang X, Zhu T (2002) Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14:559–574
Cho YJ, Meade JD, Walden JC, Chen X, Guo Z, Liang P (2001) Multicolor fluorescent differential display. BioTechniques 30:562
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Ciechanover A, Orian A, Schwartz AL (2000) Ubiquitin mediated proteolysis: biological regulation via destruction. Bioessay 22:442–451
Covitz PA, Smith LS, Long SR (1998) Expressed sequence tags from a root-hair-enriched Medicago truncatula cDNA library. Plant Physiol 117:1325–1332
Delseny M, Cooke R, Raynal M, Grellet F (1997) The Arabidopsis thaliana cDNA sequencing projects. FEBS Lett 403:221–224
Diatchenko L, Lau YF, Campbell AP, Chenchick A, Moqaddam F, Huang B, Lukyanov S, Konstantin L, Gurskaya N, Sverdlov E, Siebert PD (1996) Suppression subtractive hybridization: a method for generating differentially regulated or tissue-specific cDNA probes and libraries. Proc Natl Acad Sci USA 93:6025–6030
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Ewing B, Green P (1998) Base-calling of automated sequencer traces using Phred II. Error probabilities. Genome Res 8:186–194
Fernandes J, Brendel V, Gai X, Lal S, Chandler VL, Elumali RP, Galbraith DW, Pierson EA, Walbot V (2002) Comparison of RNA expression profiles based on maize expressed sequence tag frequency analysis and micro-array hybridization. Plant Physiol 128:896–910
Gilmour SJ, Sebolt AM, Salazar MP, Everard JD, Thomashow MF (2000) Overexpression of Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol 124:1854–1865
Goff SA, Ricke D, Lan TH, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D, Hutchison D, Martin C, Katagiri F, Lange BM, Moughamer T, Xia Y, Budworth P, Zhong J, Miguel T, Paszkowski U, Zhang S, Colbert M, Sun WL, Chen L, Cooper B, Park S, Wood TC, Mao L, Quail P, Wing R, Dean R, Yu Y, Zharkikh A, Shen R, Sahasrabudhe S, Thomas A, Cannings R, Gutin A, Pruss D, Reid J, Tavtigian S, Mitchell J, Eldredge G, Scholl T, Miller RM, Bhatnagar S, Adey N, Rubano T, Tusneem N, Robinson R, Feldhaus J, Macalma T, Oliphant A, Briggs S (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296:92–100
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Hartl FU, Hayer-Hartl M (2002) Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295:1852–1858
Haslbeck M (2002) sHsps and their role in the chaperone network. Cell Mol Life Sci 59:1649–1657
Hirt H (2000) Connecting oxidative stress, auxin, and cell cycle regulation through a plant mitogen-activated protein kinase pathway. Proc Natl Acad Sci USA 97:2405–2407
Houde M, Belcaid M, Ouellet F, Danyluk J, Monroy AF, Dryanova A, Gulick P, Bergeron A, Laroche A, Links M, McCarthy L, Crosby WL, Sarhan F (2006) Wheat EST resources for functional genomics of abiotic stress. BMC Genomics 7:149
Hubank M, Schatz DG (1994) Identifying differences in mRNA expression by representational difference analysis of cDNA. Nucleic Acids Res 22:5640–5648
Ingram J, Bartels D (1996) The molecular basis of dehydration tolerance in plants. Annu Rev Plant Physiol Plant Mol Biol 47:377–403
International Rice Genome Sequencing Project (2005) The map-based sequence of the rice genome. Nature 436:793–800
Kawasaki S, Borchert C, Deyholos M, Wang H, Brazille S, Kawai K, Galbraith D, Bohnert HJ (2001) Gene expression profiles during the initial phase of salt stress in rice. Plant Cell 13:889–905
Kizis D, Lumbreras V, Pages M (2001) Role of AP2/EREBP transcription factors in gene regulation during abiotic stress. FEBS Lett 498:187–189
Knight H, Knight MR (2001) Abiotic stress signalling pathways: specificity and cross-talk. Trends Plant Sci 6:262–267
Lecourieux D, Ranjeva R, Pugin A (2006) Calcium in plant defence-signalling pathways. New Phytol 171:249–269
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Annu Rev Plant Physiol Plant Mol Biol 49:199–222
Liang P, Pardee AB (1995) Recent advances in differential display. Curr Opin Immunol 7:274–280
Lim CO, Kim HY, Kim MG, Lee SI, Chung WS, Park SH, Hwang I, Cho MJ (1996) Expressed sequence tags of Chinese cabbage flower bud cDNA. Plant Physiol 111:577–588
Mahalingam R, Gomez-Buitrago A, Eckardt N, Shah N, Guevara-Garcia A, Day P, Raina R, Fedoroff NV (2003) Characterizing the stress/defense transcriptome of Arabidopsis. Genome Biol 4:R20
Miernyk JA (1999) Protein folding in the plant cell. Plant Physiol 121:695–703
Mishra RN, Ramesha A, Kaul T, Nair S, Sopory SK, Reddy MK (2005) A modified cDNA subtraction to identify differentially expressed genes from plants with universal application to other eukaryotes. Anal Biochem 345:149–157
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7:405–410
Neven LG, Haskell DW, Guy CL, Denslow N, Klein PA, Green LG, Silverman A (1992) Association of 70-kilodalton heat-shock cognate proteins with acclimation to cold. Plant Physiol 99:1362–1369
Pareek A, Singla SL, Grover A (1995) Immunological evidence for accumulation of two high-molecular-weight (104 and 90 kDa) HSPs in response to different stresses in rice and in response to high temperature stress in diverse plant genera. Plant Mol Biol 29:293–301
Pfaffl MW, Horgan GW, Dempfle L (2002) Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30:e36
Picard D (2002) Heat-shock protein 90, a chaperone for folding and regulation. Cell Mol Life Sci 59:1640–1648
Popova OV, Ismailov SF, Popova TN, Dietz KJ, Golldack D (2002) Salt-induced expression of NADP-dependent isocitrate dehydrogenase and ferredoxin-dependent glutamate synthase in Mesembryanthemum crystallinum. Planta 215:906–913
Pratt LH, Liang C, Shah M, Sun F, Wang H, Reid SP, Gingle AR, Paterson AH, Wing R, Dean R, Klein R, Nguyen HT, Ma H-M, Zhao X, Morishige DT, Mullet JE, Pratt M-MC (2005) Sorghum expressed sequence tags identify signature genes for drought, pathogenesis, and skotomorphogenesis from a Milestone set of 16,801 unique transcripts. Plant Physiol 139:869–884
Putney SD, Herligh WD, Schimmel P (1983) A new troponin T and cDNA clones for 13 different muscle proteins, found by shotgun sequencing. Nature 302:718–721
Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, Ito Y, Yoshiwara K, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133:1755–1767
Reddy AR, Ramakrishna AC, Sekhar AC, Nagablushana I, Ravindrababu P, Bonaldo MF, Soares MB, Bennetzen JL (2002) Novel genes are enriched in normalized cDNA libraries from drought-stressed seedlings of rice (Oryza sativa L. sub sp. Indica cv. Nagina 22). Genome 45:204–211
Rensink W, Hart A, Liu J, Ouyang S, Zismann V, Buell CR (2005) Analyzing the potato abiotic stress transcriptome using expressed sequence tags. Genome 48:598–605
Roy NV, Paepe AD, Speleman F (2003) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:Research 0034.1–0034.11
Rudd S (2003) Expressed sequence tags: alternative or complement to whole genome sequence? Trends Plant Sci 8:321–329
Sanam-Mishra N, Pham XH, Sopory SK, Tuteja N (2005) Pea DNA helicase 45 overexpression in tobacco confers high salinity tolerance without affecting yield. Proc Natl Acad Sci USA 102:509–514
Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K, Carninci P, Hayashizaki Y, Shinozaki K (2001) Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13:61–72
Seki M, Ishida J, Narusaka M, Fujita M, Nanjo T, Umezawa T, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002a) Monitoring the expression pattern of around 7,000 Arabidopsis genes under ABA treatments using a full-length cDNA microarray. Funct Integr Genomics 2:282–291
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002b) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115:327–334
Sterky F, Regan S, Karlsson J, Hertzberg M, Rohde A, Holmberg A, Amini B, Bhalerao R, Larsson M, Villarroel R, Van Montagu M, Sandberg G, Olsson O, Teeri TT, Boerjan W, Gustafsson P, Uhlen M, Sundberg B, Lundeberg J (1998) Gene discovery in the wood-forming tissues of poplar: analysis of 5,692 expressed sequence tags. Proc Natl Acad Sci USA 95:13330–13335
Takatsuji H (1999) Zinc-finger proteins: the classical zinc finger emerges in contemporary plant science. Plant Mol Biol 39:1073–1078
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Wagner D (2003) Chromatin regulation of plant development. Curr Opin Plant Biol 6:20–28
Wang W, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Xiong L, Lee MW, Yang Y (2001) Identification of defense-related rice genes by suppression subtracted hybridization and differential screening. Mol Plant-Microbe Interact 14:685–692
Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:S165–183
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yazaki J, Shimatani Z, Hashimoto A, Nagata Y, Fujii F, Kojima K, Suzuki K, Taya T, Tonouchi M, Nelson C, Nakagawa A, Otomo Y, Murakami K, Matsubara K, Kawai J, Carninci P, Hayashizaki Y, Kikuchi S (2004) Transcriptional profiling of genes responsive to abscisic acid and gibberellin in rice: phenotyping and comparative analysis between rice and Arabidopsis. Physiol Genomics 17:87–100
Young JC, Barral JM, Ulrich-Hartl F (2003) More than folding: localized functions of cytosolic chaperones. Trends Biochem Sci 28:541–547
Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M, Liu J, Sun J, Tang J, Chen Y, Huang X, Lin W, Ye C, Tong W, Cong L, Geng J, Han Y, Li L, Li W, Hu G, Huang X, Li W, Li J, Liu Z, Li L, Liu J, Qi Q, Liu J, Li L, Li T, Wang X, Lu H, Wu T, Zhu M, Ni P, Han H, Dong W, Ren X, Feng X, Cui P, Li X, Wang H, Xu X, Zhai W, Xu Z, Zhang J, He S, Zhang J, Xu J, Zhang K, Zheng X, Dong J, Zeng W, Tao L, Ye J, Tan J, Ren X, Chen X, He J, Liu D, Tian W, Tian C, Xia H, Bao Q, Li G, Gao H, Cao T, Wang J, Zhao W, Li P, Chen W, Wang X, Zhang Y, Hu J, Wang J, Liu S, Yang J, Zhang G, Xiong Y, Li Z, Mao L, Zhou C, Zhu Z, Chen R, Hao B, Zheng W, Chen S, Guo W, Li G, Liu S, Tao M, Wang J, Zhu L, Yuan L, Yang H (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Zielinski RE (1998) Calmodulin and calmodulin-binding proteins in Plants. Annu Rev Plant Physiol Plant Mol Biol 49:697–725
Acknowledgements
This work was supported in part by NATP (Indian Council of Agricultural Research) and Department of Biotechnology (Ministry of Science and Technology, Government of India) research grants. R.N.M. was supported by Council of Scientific and Industrial Research (CSIR) fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence data reported here appears in the EMBL, GenBank and DDBJ Nucleotide Sequence Databases under the accession numbers CD724312 to CD726805.
Rights and permissions
About this article
Cite this article
Mishra, R.N., Reddy, P.S., Nair, S. et al. Isolation and characterization of expressed sequence tags (ESTs) from subtracted cDNA libraries of Pennisetum glaucum seedlings. Plant Mol Biol 64, 713–732 (2007). https://doi.org/10.1007/s11103-007-9193-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-007-9193-4