Abstract
Purpose
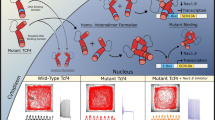
Pitt Hopkins Syndrome (PTHS) is a rare genetic disorder caused by mutations of a specific gene, transcription factor 4 (TCF4), located on chromosome 18. PTHS results in individuals that have moderate to severe intellectual disability, with most exhibiting psychomotor delay. PTHS also exhibits features of autistic spectrum disorders, which are characterized by the impaired ability to communicate and socialize. PTHS is comorbid with a higher prevalence of epileptic seizures which can be present from birth or which commonly develop in childhood. Attenuated or absent TCF4 expression results in increased translation of peripheral ion channels Kv7.1 and Nav1.8 which triggers an increase in after-hyperpolarization and altered firing properties.
Methods
We now describe a high throughput screen (HTS) of 1280 approved drugs and machine learning models developed from this data. The ion channels were expressed in either CHO (KV7.1) or HEK293 (Nav1.8) cells and the HTS used either 86Rb+ efflux (KV7.1) or a FLIPR assay (Nav1.8).
Results
The HTS delivered 55 inhibitors of Kv7.1 (4.2% hit rate) and 93 inhibitors of Nav1.8 (7.2% hit rate) at a screening concentration of 10 μM. These datasets also enabled us to generate and validate Bayesian machine learning models for these ion channels. We also describe a structure activity relationship for several dihydropyridine compounds as inhibitors of Nav1.8.
Conclusions
This work could lead to the potential repurposing of nicardipine or other dihydropyridine calcium channel antagonists as potential treatments for PTHS acting via Nav1.8, as there are currently no approved treatments for this rare disorder.






Similar content being viewed by others
References
Brockschmidt A, Todt U, Ryu S, Hoischen A, Landwehr C, Birnbaum S, et al. Severe mental retardation with breathing abnormalities (Pitt-Hopkins syndrome) is caused by haploinsufficiency of the neuronal bHLH transcription factor TCF4. Hum Mol Genet. 2007;16(12):1488–94.
Zweier C, Peippo MM, Hoyer J, Sousa S, Bottani A, Clayton-Smith J, et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt-Hopkins syndrome). Am J Hum Genet. 2007;80(5):994–1001.
Amiel J, Rio M, de Pontual L, Redon R, Malan V, Boddaert N, et al. Mutations in TCF4, encoding a class I basic helix-loop-helix transcription factor, are responsible for Pitt-Hopkins syndrome, a severe epileptic encephalopathy associated with autonomic dysfunction. Am J Hum Genet. 2007;80(5):988–93.
Pitt D, Hopkins I. A syndrome of mental retardation, wide mouth and intermittent overbreathing. Australian Paediatric J. 1978;14(3):182–4.
Rannals MD, Hamersky GR, Page SC, Campbell MN, Briley A, Gallo RA, et al. Psychiatric risk Gene transcription factor 4 regulates intrinsic excitability of prefrontal neurons via repression of SCN10a and KCNQ1. Neuron. 2016;90(1):43–55.
Bagal SK, Marron BE, Owen RM, Storer RI, Swain NA. Voltage gated sodium channels as drug discovery targets. Channels (Austin). 2015;9(6):360–6.
Swanwick RS, Pristera A, Okuse K. The trafficking of Na(V)1.8. Neurosci Lett. 2010;486(2):78–83.
Entrez Gene: sodium channel. Available from: https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=ShowDetailView&TermToSearch=6336. Accessed 13 Jul 2019
Catterall WA, Perez-Reyes E, Snutch TP, Striessnig J. International Union of Pharmacology. XLVIII. Nomenclature and structure-function relationships of voltage-gated calcium channels. Pharmacol Rev. 2005;57(4):411–25.
Plummer NW, Meisler MH. Evolution and diversity of mammalian sodium channel genes. Genomics. 1999;57(2):323–31.
Rabert DK, Koch BD, Ilnicka M, Obernolte RA, Naylor SL, Herman RC, et al. A tetrodotoxin-resistant voltage-gated sodium channel from human dorsal root ganglia, hPN3/SCN10A. Pain. 1998;78(2):107–14.
Akopian AN, Sivilotti L, Wood JN. A tetrodotoxin-resistant voltage-gated sodium channel expressed by sensory neurons. Nature. 1996;379(6562):257–62.
Akopian ANSV, England S, Okuse K, Ogata N, Ure J, Smith A, et al. Wood JN the tetrodotoxin-resistant sodium channel SNS has a specialized function in pain pathways. Nat Neurosci. 1999;2:541–8.
Cummins TR, Sheets PL, Waxman SG. The roles of sodium channels in nociception: implications for mechanisms of pain. Pain. 2007;131(3):243–57.
Nardi A, Damann N, Hertrampf T, Kless A. Advances in targeting voltage-gated sodium channels with small molecules. ChemMedChem. 2012;7(10):1712–40.
Georgijevic ML. Molecular genetics in the hereditary form of long QT syndrome. Med Pregl. 2000;53(1–2):51–4.
Harmer SC, Tinker A. The role of abnormal trafficking of KCNE1 in long QT syndrome 5. Biochem Soc Trans. 2007;35(Pt 5):1074–6.
Peroz D, Rodriguez N, Choveau F, Baro I, Merot J, Loussouarn G. Kv7.1 (KCNQ1) properties and channelopathies. J Physiol. 2008;586(7):1785–9.
Goldman AM, Glasscock E, Yoo J, Chen TT, Klassen TL, Noebels JL. Arrhythmia in heart and brain: KCNQ1 mutations link epilepsy and sudden unexplained death. Sci Transl Med. 2009;1(2):2ra6.
Roepke TK, Kanda VA, Purtell K, King EC, Lerner DJ, Abbott GW. KCNE2 forms potassium channels with KCNA3 and KCNQ1 in the choroid plexus epithelium. FASEB J. 2011;25(12):4264–73.
Anantpadma M, Lane T, Zorn KM, Lingerfelt MA, Clark AM, Freundlich JS, et al. Ebola virus Bayesian machine learning models enable new in vitro leads. ACS Omega. 2019;4(1):2353–61.
Dalecki AG, Zorn KM, Clark AM, Ekins S, Narmore WT, Tower N, et al. High-throughput screening and Bayesian machine learning for copper-dependent inhibitors of Staphylococcus aureus. Metallomics. 2019;11(3):696–706.
Hernandez HW, Soeung M, Zorn KM, Ashoura N, Mottin M, Andrade CH, et al. High throughput and computational repurposing for neglected diseases. Pharm Res. 2018;36(2):27.
Lane T, Russo DP, Zorn KM, Clark AM, Korotcov A, Tkachenko V, Reynolds RC, Perryman AL, Freundlich JS, Ekins S. Comparing and Validating Machine Learning Models for Mycobacterium tuberculosis Drug Discovery. Molecular pharmaceutics. 2018.
Russo DP, Zorn KM, Clark AM, Zhu H, Ekins S. Comparing multiple machine learning algorithms and metrics for estrogen receptor binding prediction. Mol Pharm. 2018;15(10):4361–70.
Sandoval PJ, Zorn KM, Clark AM, Ekins S, Wright SH. Assessment of substrate-dependent ligand interactions at the organic cation transporter OCT2 using six model substrates. Mol Pharmacol. 2018;94(3):1057–68.
Wang PF, Neiner A, Lane TR, Zorn KM, Ekins S, Kharasch ED. Halogen substitution influences ketamine metabolism by cytochrome P450 2B6: in vitro and computational approaches. Mol Pharm. 2019;16(2):898–906.
Zorn KM, Lane TR, Russo DP, Clark AM, Makarov V, Ekins S. Multiple machine learning comparisons of HIV cell-based and reverse transcriptase data sets. Mol Pharm. 2019;16(4):1620–32.
Clark AM. Molecular Notebook. Available from: http://molmatinf.com/MolNote/.
Clark AM, Dole K, Coulon-Spector A, McNutt A, Grass G, Freundlich JS, et al. Open source bayesian models: 1. Application to ADME/Tox and drug discovery datasets. J Chem Inf Model. 2015;55:1231–45.
Clark AM, Ekins S. Open source Bayesian models: 2. Mining a "big dataset" to create and validate models with ChEMBL. J Chem Inf Model. 2015;55:1246–60.
Bajwa PJ, Alioua A, Lee JW, Straus DS, Toro L, Lytle C. Fenofibrate inhibits intestinal cl- secretion by blocking basolateral KCNQ1 K+ channels. Am J Physiol Gastrointest Liver Physiol. 2007;293(6):G1288–99.
Publications mentioning the Prestwick Chemical library. Available from: http://www.prestwickchemical.com/libraries-publications.html. Accessed 13 Jul 2019
Huang BR, Chang PC, Yeh WL, Lee CH, Tsai CF, Lin C, et al. Anti-neuroinflammatory effects of the calcium channel blocker nicardipine on microglial cells: implications for neuroprotection. PLoS One. 2014;9(3):e91167.
Bachmeier C, Beaulieu-Abdelahad D, Mullan M, Paris D. Selective dihydropyiridine compounds facilitate the clearance of beta-amyloid across the blood-brain barrier. Eur J Pharmacol. 2011;659(2–3):124–9.
Ekins S, Williams AJ, Krasowski MD, Freundlich JS. In silico repositioning of approved drugs for rare and neglected diseases. Drug Discov Today. 2011;16:298–310.
Ekins S, Freundlich J, Clark A, Anantpadma M, Davey R, Madrid P. Machine learning models identify molecules active against Ebola virus in vitro. F1000 Research. 2015;4:1091.
Ekins S, de Lage Siqueira-Neto J, McCall LI, Sarker M, Yadav M, Ponder EL, et al. Machine learning models and pathway genome data base for trypanosoma cruzi drug discovery. PLoS Negl Trop Dis. 2015;9(6):e0003878.
Ekins S. Progress in computational toxicology. J Pharmacol Toxicol Methods. 2014;69(2):115–40.
Dong Z, Ekins S, Polli JE. Structure-activity relationship for FDA approved drugs as inhibitors of the human sodium taurocholate cotransporting polypeptide (NTCP). Mol Pharm. 2013;10(3):1008–19.
Astorga B, Ekins S, Morales M, Wright SH. Molecular determinants of ligand selectivity for the human multidrug and toxin extrusion proteins, MATE1 and MATE-2K. J Pharmacol Exp Ther. 2012;341(3):743–55.
Pan Y, Li L, Kim G, Ekins S, Wang H, Swaan PW. Identification and validation of novel hPXR activators amongst prescribed drugs via ligand-based virtual screening. Drug Metabolism Disposition: Biological Fate Chemicals. 2011;39:337–44.
Zientek M, Stoner C, Ayscue R, Klug-McLeod J, Jiang Y, West M, et al. Integrated in silico-in vitro strategy for addressing cytochrome P450 3A4 time-dependent inhibition. Chem Res Toxicol. 2010;23(3):664–76.
Ekins S, Williams AJ, Xu JJ. A predictive ligand-based Bayesian model for human drug induced liver injury. Drug Metabolism Disposition: Biological Fate Chemicals. 2010;38:2302–8.
Diao L, Ekins S, Polli JE. Quantitative structure activity relationship for inhibition of human organic cation/carnitine transporter. Mol Pharm. 2010;7:2120–30.
Zheng X, Ekins S, Raufman JP, Polli JE. Computational models for drug inhibition of the human apical sodium-dependent bile acid transporter. Mol Pharm. 2009;6(5):1591–603.
Ekins S, Kortagere S, Iyer M, Reschly EJ, Lill MA, Redinbo MR, et al. Challenges predicting ligand-receptor interactions of promiscuous proteins: the nuclear receptor PXR. PLoS Comput Biol. 2009;5(12):e1000594.
Anon. Assay Central video. Available from: https://www.youtube.com/watch?v=aTJJ6Tyu4bY&feature=youtu.be. Accessed 13 Jul 2019
Anon. Assay Central Website. Available from: www.assaycentral.org.
Clark AM, Sarker M, Ekins S. New target predictions and visualization tools incorporating open source molecular fingerprints for TB Mobile 2.0. Aust J Chem. 2014;6:38.
Anon. PF-06305591 Available from: https://clinicaltrials.gov/ct2/results?cond=&term=PF06305591&cntry1=&state1=&SearchAll=Search+all+studies&recrs=. Accessed 13 Jul 2019
Bagal SK, Brown AD, Kemp MI, Klute W, Sanz LM, Marron BE, Miller DC, Skerrat E, Suto MJ, West CW. Chemical Compounds. In: US, editor.: Pfizer Limited; 2013.
ACKNOWLEDGMENTS AND DISCLOSURES
The work at Icagen was funded by the Pitt Hopkins Research Foundation. SE kindly acknowledges NIH funding R43GM122196 and R44GM122196-02A1 “Centralized assay datasets for modelling support of small drug discovery organizations” from NIH/ NIGMS which supported development of Assay Central. We also kindly thank Dr. Alex Clark for assistance with Assay Central and Dr. Mary Lingerfelt, Dr. Aaron McMurtray and Dr. Kimberly Goodspeed for discussions. S.E. is owner, and K.M.Z., are employees of Collaborations Pharmaceuticals Inc. J.G. was an intern at Collaborations Pharmaceuticals Inc. A.G., B.A. and Z.L. are employees of Icagen. Models are available from authors upon request.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ekins, S., Gerlach, J., Zorn, K.M. et al. Repurposing Approved Drugs as Inhibitors of Kv7.1 and Nav1.8 to Treat Pitt Hopkins Syndrome. Pharm Res 36, 137 (2019). https://doi.org/10.1007/s11095-019-2671-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11095-019-2671-y