Abstract
Reactive nitrogen species, such as nitric oxide (NO), exert their biological activity in large part through post-translational modification of cysteine residues, forming S-nitrosothiols. This chemical reaction proceeds via a process that we and our colleagues have termed protein S-nitrosylation. Under conditions of normal NO production, S-nitrosylation regulates the activity of many normal proteins. However, in degenerative conditions characterized by nitrosative stress, increased levels of NO lead to aberrant S-nitrosylation that contributes to the pathology of the disease. Thus, S-nitrosylation has been implicated in a wide range of cellular mechanisms, including mitochondrial function, proteostasis, transcriptional regulation, synaptic activity, and cell survival. In recent years, the research area of protein S-nitrosylation has become prominent due to improvements in the detection systems as well as the demonstration that protein S-nitrosylation plays a critical role in the pathogenesis of neurodegenerative and other neurological disorders. To further promote our understanding of how protein S-nitrosylation affects cellular systems, guidelines for the design and conduct of research on S-nitrosylated (or SNO-)proteins would be highly desirable, especially for those newly entering the field. In this review article, we provide a strategic overview of designing experimental approaches to study protein S-nitrosylation. We specifically focus on methods that can provide critical data to demonstrate that an S-nitrosylated protein plays a (patho-)physiologically-relevant role in a biological process. Hence, the implementation of the approaches described herein will contribute to further advancement of the study of S-nitrosylated proteins, not only in neuroscience but also in other research fields.



Similar content being viewed by others
Abbreviations
- GSNO:
-
S-Nitrosoglutathione
- MS:
-
Mass spectrometry
- NO:
-
Nitric oxide
- NOS:
-
NO synthase
- SNO:
-
S-Nitrosylation or S-nitrosothiol
- SNOC:
-
S-Nitrosocysteine
References
Martinez-Ruiz A, Cadenas S, Lamas S (2011) Nitric oxide signaling: classical, less classical and nonclassical mechanisms. Free Radic Biol Med 51:17–29
Hess DT, Matsumoto A, Kim SO, Marshall HE, Stamler JS (2005) Protein S-nitrosylation: purview and parameters. Nat Rev Mol Cell Biol 6:150–166
Nakamura T, Tu S, Akhtar MW, Sunico CR, Okamoto S, Lipton SA (2013) Aberrant protein S-nitrosylation in neurodegenerative diseases. Neuron 78:596–614
Smith BC, Marletta MA (2012) Mechanisms of S-nitrosothiol formation and selectivity in nitric oxide signaling. Curr Opin Chem Biol 16:498–506
Seth D, Stamler JS (2011) The SNO-proteome: causation and classifications. Curr Opin Chem Biol 15:129–136
Stamler JS, Lamas S, Fang FC (2001) Nitrosylation. The prototypic redox-based signaling mechanism. Cell 106:675–683
Stamler JS, Toone EJ, Lipton SA, Sucher NJ (1997) (S)NO signals: translocation, regulation, and a consensus motif. Neuron 18:691–696
Lipton SA, Choi YB, Pan ZH, Lei SZ, Chen HS, Sucher NJ, Loscalzo J, Singel DJ, Stamler JS (1993) A redox-based mechanism for the neuroprotective and neurodestructive effects of nitric oxide and related nitroso-compounds. Nature 364:626–632
Mannick JB, Hausladen A, Liu L, Hess DT, Zeng M, Miao QX, Kane LS, Gow AJ, Stamler JS (1999) Fas-induced caspase denitrosylation. Science 284:651–654
Nott A, Watson PM, Robinson JD, Crepaldi L, Riccio A (2008) S-nitrosylation of histone deacetylase 2 induces chromatin remodelling in neurons. Nature 455:411–415
Tenneti L, D’Emilia DM, Lipton SA (1997) Suppression of neuronal apoptosis by S-nitrosylation of caspases. Neurosci Lett 236:139–142
Cho DH, Nakamura T, Fang J, Cieplak P, Godzik A, Gu Z, Lipton SA (2009) S-nitrosylation of Drp1 mediates β-amyloid-related mitochondrial fission and neuronal injury. Science 324:102–105
Hara MR, Agrawal N, Kim SF, Cascio MB, Fujimuro M, Ozeki Y, Takahashi M, Cheah JH, Tankou SK, Hester LD, Ferris CD, Hayward SD, Snyder SH, Sawa A (2005) S-Nitrosylated GAPDH initiates apoptotic cell death by nuclear translocation following Siah1 binding. Nat Cell Biol 7:665–674
Nakamura T, Wang L, Wong CC, Scott FL, Eckelman BP, Han X, Tzitzilonis C, Meng F, Gu Z, Holland EA, Clemente AT, Okamoto S, Salvesen GS, Riek R, Yates JR III, Lipton SA (2010) Transnitrosylation of XIAP regulates caspase-dependent neuronal cell death. Mol Cell 39:184–195
Ryan SD, Dolatabadi N, Chan SF, Zhang X, Akhtar MW, Parker J, Soldner F, Sunico CR, Nagar S, Talantova M, Lee B, Lopez K, Nutter A, Shan B, Molokanova E, Zhang Y, Han X, Nakamura T, Masliah E, Yates JR 3rd, Nakanishi N, Andreyev AY, Okamoto S, Jaenisch R, Ambasudhan R, Lipton SA (2013) Isogenic human iPSC Parkinson’s model shows nitrosative stress-induced dysfunction in MEF2-PGC1α transcription. Cell 155:1351–1364
Uehara T, Nakamura T, Yao D, Shi ZQ, Gu Z, Ma Y, Masliah E, Nomura Y, Lipton SA (2006) S-Nitrosylated protein-disulphide isomerase links protein misfolding to neurodegeneration. Nature 441:513–517
Jaffrey SR, Erdjument-Bromage H, Ferris CD, Tempst P, Snyder SH (2001) Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat Cell Biol 3:193–197
Seneviratne U, Godoy LC, Wishnok JS, Wogan GN, Tannenbaum SR (2013) Mechanism-based triarylphosphine-ester probes for capture of endogenous RSNOs. J Am Chem Soc 135:7693–7704
Gaston B, Reilly J, Drazen JM, Fackler J, Ramdev P, Arnelle D, Mullins ME, Sugarbaker DJ, Chee C, Singel DJ et al (1993) Endogenous nitrogen oxides and bronchodilator S-nitrosothiols in human airways. Proc Natl Acad Sci USA 90:10957–10961
Lei SZ, Pan ZH, Aggarwal SK, Chen HS, Hartman J, Sucher NJ, Lipton SA (1992) Effect of nitric oxide production on the redox modulatory site of the NMDA receptor-channel complex. Neuron 8:1087–1099
Zhang Y, Hogg N (2005) S-Nitrosothiols: cellular formation and transport. Free Radic Biol Med 38:831–838
Hirota Y, Ishida H, Genka C, Obama R, Matsuyama S, Nakazawa H (2001) Physiological concentration of nitric oxide induces positive inotropic effects through cGMP pathway in isolated rat ventricular myocytes. Jpn J Physiol 51:455–461
Pinsky DJ, Patton S, Mesaros S, Brovkovych V, Kubaszewski E, Grunfeld S, Malinski T (1997) Mechanical transduction of nitric oxide synthesis in the beating heart. Circ Res 81:372–379
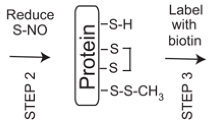
Forrester MT, Foster MW, Benhar M, Stamler JS (2009) Detection of protein S-nitrosylation with the biotin-switch technique. Free Radic Biol Med 46:119–126
Forrester MT, Foster MW, Stamler JS (2007) Assessment and application of the biotin switch technique for examining protein S-nitrosylation under conditions of pharmacologically induced oxidative stress. J Biol Chem 282:13977–13983
Shi ZQ, Sunico CR, McKercher SR, Cui J, Feng GS, Nakamura T, Lipton SA (2013) S-Nitrosylated SHP-2 contributes to NMDA receptor-mediated excitotoxicity in acute ischemic stroke. Proc Natl Acad Sci USA 110:3137–3142
Gaj T, Gersbach CA, Barbas CF 3rd (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405
Acknowledgments
This work was supported in part by NIH Grants P30 NS076411, R01 NS086890, R01 ES017462, P01 HD029587, and R21 NS080799 (SAL), the Brain and Behavior Research Foundation (SAL), and the Michael J. Fox Foundation (SAL and TN).
Author information
Authors and Affiliations
Corresponding author
Additional information
Special Issue: In honor of Dr. Philip Beart.
Rights and permissions
About this article
Cite this article
Nakamura, T., Lipton, S.A. Nitrosative Stress in the Nervous System: Guidelines for Designing Experimental Strategies to Study Protein S-Nitrosylation. Neurochem Res 41, 510–514 (2016). https://doi.org/10.1007/s11064-015-1640-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-015-1640-z