Abstract
AT-rich DNA elements play an important role in regulating cell-specific gene expression. One of the AT-rich DNA binding proteins, SATB1 is a novel type of transcription factor that regulates gene expression in the hematopoietic lineage through chromatin modification. Using DNA-affinity purification followed by mass spectrometry we identified and isolated a related protein, SATB2 from the developing rat cerebral cortex. SATB2 shows homology to SATB1 and the rat protein is practically identical to the mouse and human SATB2. Using competitive EMSA, we show that recombinant SATB2 protein binds with high affinity and specificity to AT-rich dsDNA. Using RT-PCR, Western analysis and immunohistochemistry we demonstrate that SATB2 expression is restricted to a subset of postmitotic, differentiating neurons in the rat neocortex at ages E16 and P4. We suggest that similar to its homologue SATB1, SATB2 is also involved in regulating gene expression through altering chromatin structure in differentiating cortical neurons.
Similar content being viewed by others
Introduction
Proper differentiation of progenitors into diverse neuronal phenotypes of the adult brain greatly depends on the precise regulation of cell-specific gene expression. Although many transcription factors that control the process have been identified, we still lack the complete understanding about the molecular requirements of this critical developmental process. One approach toward the identification of regulatory cis- and trans-acting factors is to screen for development- and cell-specific interactions between nuclear proteins and cis-regulatory DNA elements of genes of interest [1]. Because of its contrasting pattern of expression in the adult brain, the enkephalin (ENK) gene offers an excellent model for such approach. The majority of enkephalinergic neurons—nerve cells that express the ENK gene and use the various enkephalin peptides as neurotransmitters—are found in the neostriatum whereas most other regions, such as the cerebral cortex are practically devoid of ENK+ neurons [2]. Enkephalin peptides are endogenous opiate ligands and they pharmacologically compete with and mimic the effects of opiate drugs (such as morphine) [3, 4].
The transcriptional regulation of cell specific expression of genes, such as the ENK gene in the developing brain is envisaged as a cascade of spatiotemporally coordinated interactions between trans-acting factors and cis-regulatory DNA elements of genes [5]. These trans-acting factors are differentially expressed in progenitors/differentiating neurons at progressive stages of development and can either activate or repress gene transcription resulting in the cell-specific expression pattern of the gene in the adult brain. Previous work using a rat model indicated that 2.7 kb 5′ upstream region of the ENK gene is both necessary and sufficient to govern correct cell-specific expression of the gene [6]. Our recent studies using the cre-lox “knock-in” strategy also indicated that the proximal 1.4 kb of the mouse ENK gene is sufficient for organ-specificity but farther upstream DNA elements are required for development- and cell-specific expression of the ENK gene [7]. This 2.7 kb region of the rat, mouse and human ENK genes contains an AT-rich DNA sequence [8–10] and interfering with the binding of proteins to this AT-rich DNA element resulted in an altered reporter gene expression [1, 11, 12].
AT-rich DNA sequences are tightly associated with the nuclear matrix or scaffold and they form the core regions of MAR/SAR DNA [13–15]. AT-rich DNA binding proteins include nucleolin [16], HMGI/Y [17] and the thymus-specific developmental regulator SATB1 [18, 19]. Multiple in vitro and in vivo studies have demonstrated that SATB1 plays a major role in regulating hematopoietic differentiation through chromatin modifications [20–22].
Because of the demonstrated role of AT-rich DNA elements in regulating cell- and tissue-specific gene expression e.g. [17, 22–24], we set out to identify its binding proteins that are expressed in the developing rat brain. Here we report the identification, isolation and characterization of a SATB1-related protein, called SATB2 and its pattern of spatial and temporal expression in the rat brain. During our final experiments and preparation of this manuscript, a group using subtractive cloning isolated the mouse SATB2 gene from embryonic mouse cortex [25].
Experimental procedures
DNA affinity purification and protein identification by electrospray tandem mass spectrometry
DNA affinity purification was performed according to Kumar et al. [26] with some modifications. Two hundred micrograms of nuclear extract or recombinant proteins were precleared with 60 μl streptavidin-agarose beads (PIERCE, Rockford, IL) washed in buffer Z (25 mM HEPES (pH 7.9), 20% glycerol, 0.1% Igepal, 0.1 M KCl, 12.5 mM MgCl2, 1 mM DTT, 0.1 μM ZnCl2) After separation by centrifuging, 300 pmol competitor ds- or ssDNA or equal volume of 1× annealing buffer (50 mM Tris–HCl (pH: 7.5), 10 mM MgCl2) were added in the presence of 0.1 μg/μl poly[d(I-C)], 1 mM Na-Orthovanadate and Complete Mini protease inhibitors (Roche, Indianapolis, IN), and incubated under gentle rotation for 15 min at 4°C. 30 pmol of biotinylated ds- or ssDNA (rATENK or rATENK mut, for sequence information, see Table 1) were added to the recovered nuclear extract, and mixed by gentle rotation for 2 h at 4°C. Sixty microliters of washed streptavidin-agarose bead slurry was added and the suspension was further incubated, gently rotating, for 2 h at 4°C. The mixture was centrifuged briefly, the pelleted beads were washed three times in buffer Z and beads were boiled in the presence of LDS sample buffer (Invitrogen, Carlsbad, CA) under reducing conditions. The bound proteins were separated on NuPage 4–12% Bis–Tris gels. In gel digestion and analysis of peptides by mass spectrometry were performed as previously described [27]. Briefly, peptides were excised, reduced with 4 mM DTT at room temperature for 1 h, alkylated with 55 mM iodoacetamide for 1 h at room temperature in the dark, and digested with trypsin overnight at 37°C. Digested peptides were analyzed on an ion trap mass spectrometer LCQ-DECA-XP plus equipped with a nanoelectrospray source and an online Surveyor HPLC (Thermo Finnigan). Proteins were identified by searching the resulting MS/MS fragmentation spectra against non-redundant GenBank protein sequence database using TurboSequest. Peptides were filtered using following criteria (Xcorr >1.9 (z=1), 2.3 (z=2) and 3.5 (z=3) and delta Cn> 0.1).
Cloning and plasmid construction
DNA fragments encoding the SATB2 protein were amplified from an embryonic rat brain library in pGAD vector (Promega, Madison, WI) using the primers 5′S2F and 3′S2F (Table 1). PCR products were cloned into pEAK10 vector (Edge Biosystems, Gaithersburg, MD) through the EcoRI and NotI cloning sites. Plasmids were sequenced and recombinant rSATB2 was overexpressed in PEAK cells using the PEAK Rapid Expression Kit (Edge Biosystems, Gaithersburg, MD) and purified according to the manufacturer’s instructions.
Electrophoretic mobility shift assays (EMSA)
EMSA were performed as previously described [28]. Oligonucleotides representing the AT-rich DNA region of the rENK gene (rATENK F, rATENK R, or mutant oligos: rATENK mutF, rATENK MutR, see Table 1) were chemically synthesized radioactively labelled and annealed. In the competition assay, unlabeled DNA fragments were used in 100× excess prior to the addition of the nuclear extract (“head-to-head” competition). For supershift assay 1 μl of polyclonal rabbit antibody or rabbit preimmune-serum were used. Antibodies were mixed with 2 μg of recombinant SATB2 protein in the presence of 0.1 μg/μl of poly[d(I-C)]. The reaction mixture was incubated on ice for 30 min then radioactively labelled double stranded rATENK probes were added and the mixture was further incubated for 10 min on ice and analyzed using an agarose thin-layer gel system and [29] analyzed in PhosphorImager (Molecular Dynamics, Carlsbad, CA).
Semi-quantitative reverse transcription PCR
Total RNA was purified from rat cortex dissected at different developmental ages and from P2 thymus using the Trizol reagent (Invitrogen, Carlsbad, CA). Two micrograms of RNA was reverse transcribed with the Promega Reverse Transcription System (Madison, WI). The obtained cDNA was diluted 10-fold and served as a template in the following PCR. A SATB2-specific DNA fragment was amplified by using the primers 5′S2A and 3′S2A by YieldAce polymerase (Stratagene, La Jolla, CA). After an initial denaturizing step at 95°C for 5 min, the following temperature profile was followed during 35 cycles: 30 s at 95°C, 30 s at 55°C and 1 min at 72°C. The amplicons were subjected to gel electrophoresis on a 2% agarose gel and bands were visualised under UV light after Ethydium Bromide staining, photographed using Kodak Gel Electrophoresis and Documentation System 120.
Antibody production and testing
Antibody production was performed at ProteinTech Group Inc. (Chicago, IL). Based on the predicted amino acid sequence, the peptide: CAEEESAEKNKVAPAETDQR was synthesized using FMOC chemistry on solid phase peptide synthesizer (Symphony, Protein Technologies, Inc. Arizona). Synthesis took place from C-terminal to N-terminal. The C-terminal fmoc-amino acid (Anaspec, Inc California) was attached to an insoluble support resin via an acid labile linker. Fmoc group of this amino acid was de-protected by 20% piperidine. The second fmoc-amino acid was coupled to the c-terminal residue using 0.1 M HBTU in DMF containing 0.4 M 4-Methyl morpholine for 60 min and later washed to remove all unbound amino acid. Fmoc group was again removed and the cycle was continued to generate the peptide. The resin bound peptide is de-protected of its side chain protection groups as well as cleaved from the resin using Trifluoroacetic acid (TFA). Ethyl ether is added to precipitate the peptide from the TFA solution. The peptide is then characterized by MALDI mass spec (Voyager, DE PRO, Applied Biosystems). The Cys amino acid added onto the peptide was used to conjugate the peptide to KLH using a heterobifunctional cross linker 3-Maleimidobenzoic acid N-Hydroxysuccinimide ester (MBS). Unbound MBS was removed by PD 10 column. KLH bound maleimide was reacted with the peptide at pH ~7.3 for 2–24 h. Afterwards, unbound peptide was removed by PD-10 column and lyophilized. Rabbits were immunized with 2 mg of peptide-KLH in Complete Freund’s Adjuvant. They received booster with 2 mg of peptide-KLH in Incomplete Freund’s Adjuvant in 30 days. Three more boosts were administered in 2 weeks interval thereafter. The titer was tested by ELISA using the synthetic peptide and the final serum was affinity purified and tested by Western analysis and by immunohistochemistry.
Western analysis
Twelve micrograms of proteins were separated on NuPage 4–12% Bis–Tris gels, transferred onto PVDF membranes by electroblotting (Invitrogen, Carlsbad, CA) followed by incubating the membranes with various primary antibodies overnight at 4°C. Immunoreactive protein bands were visualized using SuperSignal West Pico Chemiluminescent Kit (PIERCE, Rockford, IL). The membranes were stripped using the Re-Blot Plus Mild Antibody Stripping Solution (Chemicon, Temecula, CA) and re-probed with anti-GAPDH antibody (Sigma-Aldrich, St. Louis, MO) as described above.
Immunohistochemistry
Rat embryos were obtained from timed pregnancies (Zivic Miller Laboratories, Pittsburgh, PA). Following anesthesia, brains were removed and snap frozen on powdered dry ice. Anesthetized animals at gestation ages E17 and P1 were injected with 50 mg/kg BrdU i.p. and brains were removed and frozen 24 h later as above. Twenty micrograms frozen coronal sections containing the cerebral cortex, striatum, septum (or their primordia) were cut and processed for immunohistochemistry using anti-rSATB2 antibody at 1:200 dilution and anti-BrdU monoclonal antibody (Becton Dickinson Immunocytometry Systems, San Jose, CA) at 1:100 dilution. After washing, sections were processed for immunofluorescent detection using FITC-conjugated secondary antibodies (1:1000, Molecular Probes, Eugene, Oregon). Sections were viewed in an Olympos 70 IX microscope equipped with a Spot digital camera or in a Leica DM RXA microscope (Biomedical Instrumentation Center, USUHS) using appropriate fluorescent channels and images were merged using TIFFany3 software to create the final pictures.
Results
Identification of SATB2 as AT-rich DNA binding protein expressed in the developing rat cortex
To isolate proteins that specifically bind to the AT-rich DNA element of the rENK gene, we immobilized rATENK dsDNA fragments and performed DNA-affinity purification using nuclear extracts form neonatal rat cerebral cortex. Bound proteins were separated by gel electrophoresis (Fig. 1), individual bands were cut out and analyzed by mass spectrometry. As expected, we found several known ubiquitous AT-binding proteins (Table 2). These include DNA topoisomerase II A and B [30], poly(ADP-ribose) polymerase I [31, 32], hnRNP U (SAF-A) [33, 34] and hnRNPK [35]. These proteins are known to be either part of the nuclear matrix or frequently associated with it indicating that rATENK binds to the nuclear matrix. In addition, our mass spectrometry analysis also found that the hematopoietic regulator SATB1 also expressed in the developing cortex and binds to rATENK dsDNA [18, 19, 25, 36]. Finally, we identified a homologue to SATB1 originally called KIAA1034 [37, 38]. Because of its homology to SATB1, the protein was renamed as “special AT binding protein 2” or SATB2.
Identification of AT-rich DNA binding protein following DNA affinity purification. Nuclear extract from developing rat cortex (from P2) was used for DNA affinity purification. The binding fraction was loaded on a 4–12% gradient polyacrylamide gel, was subjected to electrophoresis and stained with SyproRuby stain. After visualization the proteins were identified by mass spectrometry. These bands are marked with arrows. Abbreviations: TOP2B: DNA-Topoisomerase II beta; TOP2A: DNA-Topoisomerase II alpha, PARP: poly(ADP-ribose) polymerase, SATB1: special AT-rich sequence binding protein 1, SATB2: special AT-rich sequence binding protein 2 (or similar to SATB1). hnRNP U or K: heterogenous ribonucleoprotein U or K
Because during the time of our DNA affinity studies there was no information about the neuronal expression of SATB2 and because its homologue SATB1 represents a novel type of transcriptional regulator, we decided to characterize SATB2 and analyze its expression in the rat brain.
To verify the identity of SATB2 as the AT-rich DNA binding protein expressed in the developing rat cortex, we isolated its cDNA clone from a neonatal rat brain library using PCR cloning. Comparison of the deducted amino acid sequences of rat (rSATB2), mouse (mSATB2) and human (hSATB2) proteins has revealed high level of homology (Fig. 2). Amino acid substitutions of rSATB2 compared to the mouse of the human protein (marked with stars) were found on the C-terminus outside of known functional domains.
Comparison of amino acid sequences for mouse, human and rat SATB2. The number refers to the position of amino acids. The black and grey boxes indicate the conserved “cut” and homeodomains. Dots indicate identical amino acids. Amino acid subsitutions in the rat are bold and marked with a star above. Possible SUMOilation sites are italicised and marked with an ‘s’ above. The 19 amino acid peptide sequence (AEEESAEKNKVAPAETDQR) used to raise the rat SATB2 specific antibody is underlined on the C-terminal of the protein
Similar to a recent publication [25] we found significant homology between SATB1 and SATB2 at a region between amino acids 355 and 483 of SATB1 (data not shown). This region, part of a cut domain, has been shown to be responsible for the high affinity binding of SATB1 to AT-rich DNA [18, 36].
Based on the predicted amino acid sequence, we synthesized a peptide unique to rSATB2 and raised antibody against the protein.
Characterization of rSATB2
To characterize the binding specificity of rSATB2 protein, we expressed the recombinant protein and performed competitive EMSA using rATENK dsDNA as probe. Comparing the mobility of protein–DNA complexes formed with the recombinant rSATB2 protein and nuclear extracts isolated from P2 cortex showed similar mobility (Fig. 3). However, the complex formed with the nuclear extract appeared as a broader band indicating the presence of multiple proteins in the complex. This finding is consistent with our DNA affinity purification and mass spectrometry data that identified these additional AT-rich DNA binding proteins (see Fig 1 and Table 2). Competitive EMSA showed that the binding of SATB2 to the AT-rich dsDNA probe was specific. Neither single stranded specific DNA competitors (rATENK–F or rATENK–R, see Table 1 for details) nor mutant single-stranded competitors (rATENK mut–F or rATENK mut–R) or double-stranded mutant competitor (rATENK mut) interfered with the binding. We also compared the binding of rSATB2 to the previously identified AT-rich DNA binding protein, SATB1. In the presence of 0.1 μg/μl of poly[d(I-C)] we use for EMSA, SATB2 did bind to rATENK dsDNA probe with an apparent higher affinity than that of the SATB1. In addition, consistent with the different molecular weights and amino acid compositions, the two complexes also showed different mobility.
Binding specificity of SATB2 to AT-rich DNA element. Competitive electro-mobility shift assay (EMSA) with different single- and doublestranded AT-rich and mutant DNA. Abbreviations: S2: recombinant rSATB2; S1: recombinant rSATB1; Cx: nuclear extract from P2 rat cortex; ds: double stranded rAT(ENK); sT: single stranded rAT(ENK)-F; sA: single stranded rAT(ENK)-R; dM: double stranded rAT(ENK) mut; sM: single stranded rAT(ENK)mut-F; S: anti-SATB2 antibody
Importantly, in the presence of rSATB2 antibody, the complex was supershifted confirming that the protein is rSATB2 (Fig. 3, lane: s).
Developmental expression of SATB2 mRNA and protein in the rat brain
We analyzed the developmental pattern of rSATB2 mRNA expression in the rat cerebral cortex by semi-quantitative RT-PCR using primers 5′S2A and 3′S2A (Table 1). As shown on Fig. 4a, we detected low levels of rSATB2 mRNA in the rat telencephalon at E14. The abundance of rSATB2 mRNA gradually increases in the developing cortex and reaches the highest abundance at P2. SATB2 mRNA levels were gradually decreasing after P2 and only very low level of rSATB2 mRNA was detected in the adult cortex. Interestingly, we detected no SATB2 mRNA in the neonatal thymus, a rich source of SATB1 mRNA at this developmental stage [39].
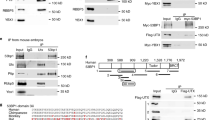
(a) SATB2 mRNA expression in the developing rat cortex. RT-PCR analysis of total RNA isolated from whole brains at different ages (E14-P28). Thy: total RNA derived from P2 thymus. (+): positive control where SATB2-coding plasmid was used as a template. (−): No template control. M: 100 bp molecular weight marker. S: anti-SATB2 antibody used for supershift assay. (b) SATB2 protein in the developing rat cortex. Western blot with anti-SATB2 antibody. All lanes contains equal amount of nuclear extracts isolated from whole brain from E14, or cortex from E18, P2 or adult rats. The antibody is specific to SATB2 and did not recognize any other protein. We used anti-GAPDH antibody as a loading control
We also analyzed the developmental distribution of SATB2 protein using Western blot analysis. As opposed to mRNA shown above, we did not detect SATB2 protein in the E14 cortical nuclear extract (Fig. 4b). We detected high level of SATB2 protein in nuclear extracts isolated from E18 cortex and the abundance of SATB2 protein was further increased at P2. No SATB2 protein was seen in the adult rat cortex. Overall, the observed developmental pattern of SATB2 mRNA and protein expressions showed the same trend. The apparent discrepancy between mRNA and protein levels are likely due to the lower sensitivity of Western analysis.
Distribution and characterization of SATB2 expressing cells in the rat brain
To obtain information about the cell-specific expression of SATB2 in the rat brain, we performed immunohistochemical analysis using the same SATB2 antibody as in Western analysis. We found that SATB2 immunoreactive cells were restricted to the developing cortex, no immunoreactive cells were found in any of other brain regions. SATB2 immunoreactive cells were first detected at E16 in the superficial layers of the developing cortex (Fig. 5). At E18, the number of SATB2 immunoreactive cells and the apparent staining intensity increased substantially. SATB2 expressing cells were present in the marginal zone, cortical plate and intermediate zone. Interestingly, SATB2 (+) cells were mostly restricted to the neocortex and cells at similar laminar locations in the piriform cortex were not expressing SATB2. This finding is consistent with a recent observation in the mouse brain using in situ hybridization histochemical analysis also showing that SATB2 mRNA expression is absent from the developing piriform cortex [25]. The number of immunoreactive cells and apparent staining intensity further increased in the rat neocortex at P2 and then sharply declined by P4.
Distribution of SATB2 expressing cells in the rat cortex. (A)–(D) Cross-sections derived from developing rat brains at ages E16, E18, P2 and P4 were immunostained with the SATB2 antibody (red). The low magnification images show immunoreactive cells that increase in numbers and staining intensity in the neocortex between E16 and P2 and decreases thereafter. (E)–(H) SATB2 expressing cells (red) and proliferating, BrdU (green) are spatially separated at E18 (E–F). The insert on (E) indicate the location of the higher magnification image (f). SATB2+ (red) cells are located in the intermediate zone (IMZ), in the cortical plate (CP) and in the marginal zone (MZ). BrdU immunoreactive cells (green) cells are located in the ventricular (VZ) and subventricular zones (SVZ). Some BrdU+ cells are present in the IMZ but they do not express SATB2. At P2 (G–H) the separation between SATB2+ and BrdU+ cells is vene more marked. Although there are both SATB2+ and BrdU+ cells in the IMZ, there are no double labelled cells were found. Abbreviation: LV, lateral ventricle
The observed spatio-temporal distribution of SATB2 positive cells in the cortex suggested that they are postmitotic, differentiating neurons. To test this hypothesis, we performed SATB2/BrdU double immunohistochemistry at ages E18 and P2. We found a clear boundary between BrdU and SATB2 immunoreactive cells at both ages investigated and we found no double-labelled cells. Consistent with previous observations, BrdU+ proliferating cells populated the ventricular and the subventricular zones [40, 41] whereas SATB2 immunoreactive cells were located at the marginal zone and cortical plate. The few BrdU+ cells we detected in the intermediate zone at E18 were not expressing SATB2 and similarly, all SATB2 immunoreactive cells at this location were BrdU negative. At P2 when the number of proliferating cells is substantially reduced and restricted to small number of cells at the ventricular zone, the separation of BrdU positive and SATB2 expressing cells was even more distinct. These findings demonstrated that SATB2 is expressed in a subpopulation of postmitotic, differentiating neurons in the rat cerebral cortex and that these cells were born before E17 in the rat. Our findings are consistent with a recent report showing that SATB2 is expressed in BrdU negative, postmitotic cells in the mouse cerebral cortex [25].
Discussion
Toward better understanding the transcriptional control of cell-specific gene expression during neuronal differentiation we have used the ENK gene as model and employed the “sample & probe” approach [1]. The approach is based on screening for development- and cell-specific interactions between nuclear proteins and cis-regulatory DNA elements of genes of interest followed by in vivo verification of the interaction [1, 11, 28, 42]. One of the cis-regulatory DNA elements we have identified is an AT-rich DNA site present on the upstream regulatory region of the mouse, rat and human ENK genes [8]. In the rat the AT-rich sequence is a perfect repeat (ATT)19. Although, AT-rich sequences were found at similar upstream locations on the mouse and human ENK genes, they lack the regularity/repetitive nature of the rat sequence. This is consistent with the finding that AT-rich DNA elements do not have consensus sequence in a classical sense [23, 43, 44]. DNA sequences that are composed of higher than 70% of A or T nucleotides and longer than 36 nucleotides form specific binding sites on the minor grove of the AT-rich DNA and the precise dimensions of these sites are determined by the frequency of A- and T-nucleotides. These DNA sites display unique structural features including base un-pairing which may contribute to their unique regulatory mechanism [45]. AT-rich DNA elements form the core of MARs/SARs which are frequent in the genome [44]. Accordingly, AT-rich DNA binding proteins such as SATB1 and SATB2 have many potential binding sites present on large numbers of genes [14, 46, 47].
Both SATB2 and SATB1 lack the AT-hook, found in ubiquitous AT-rich DNA binders such as HMGI/Y protein [17, 24, 36, 48]. Similar to a recent publication [25] we also found high level of homology between rSATB1 and rSATB2 including a region previously identified as responsible for high affinity binding of SATB1 to AT-rich DNA elements [18, 36]. Although the original binding site for SATB1 is a classical MAR/SAR element [19] that is significantly longer than the AT-rich element of the rENK gene [11, 12] both proteins bound to this DNA element with similar affinity.
Although we used the AT-rich DNA element of the rENK gene to DNA affinity purify SATB2, the role of SATB2 in regulating ENK gene expression is currently unknown. AT-rich DNA sequences are frequent in the genome thus the AT-rich DNA element of the ENK gene can be one of the many potential binding site for SATB2. Until a genetic model (SATB2 “knock-out”) is available, the role of SATB2 in regulating ENK gene expression will remain unclear. Previous studies showed that SATB2 (earlier known as KIAA1034) involved in regulating the expression of the immunoglobulin mu gene and being involved in palate formation [37, 38].
Studies have shown that the first AT-specific binding protein, SATB1 exerts its regulatory role by regulating chromatin structure. In the lymphoid lineage, SATB1 interacts with chromatin remodellers and it is involved in organizing the chromatin structure thereby controlling the accessibility of multiple transcription factors [20, 49, 50] suggesting that SATB1 acts as a “global” coordinator of cellular differentiation in the hematopoietic lineage. Chromatin remodelling is emerging as a critical mechanism of coordinating the expression of large numbers if genes [51–53]. Another example of such regulator expressed both in the hematopoietic and in the neuronal lineages is Ikaros [39, 54, 55]. Ikaros binds to short nucleic acid motif and not to AT-rich DNA but controls phenotypic differentiation by regulating the expression of large numbers of genes by chromatin remodelling in the hematopoietic system [56]. In the developing brain, Ikaros is specifically expressed by both proliferating and by differentiating striatal neurons between E14 and E18 and Ikaros mutation results in reduced numbers of striatal ENK mRNA+ neurons [54].
Our immunohistochemical data and a recent study [25] showed that SATB2 is abundantly expressed in a subset of postmitotic differentiating neurons in the neocortex implicating SATB2 in regulating gene expression during cortical development. We believe that both SATB1 and SATB2 are major regulators of gene expression during cortical development [19, 25]. Our own pilot data (not shown) and a recent publication [25] demonstrate that in the developing brain, the pattern of expression of SATB1 and SATB2 is not overlapping suggesting that they regulate the differentiation of distinct populations of neurons. Accordingly, SATB1 and SATB2 can be used as markers to identify distinct subsets of differentiating cortical neurons.
In summary, SATB2 represents a novel type of nuclear regulatory molecule expressed in the developing brain. Understanding the role of SATB2 in cortical development along with the identification of its downstream target genes and the molecular mechanism of SATB2 regulation are important goals for future studies.
References
Dobi A, Palkovits M, Ring MA, Eitel A, Palkovits CG, Lim F, Agoston DV (1997) Sample and probe: a novel approach for identifying development-specific cis-elements of the enkephalin gene. Brain Res Mol Brain Res 52:98–111
Loughlin SE, Leslie FM, Fallon JH (1995) Endogenous opioid systems In: Paxinos G (eds) The Rat Nervous System. Academic Press, San Diego, pp. 975–1001
Akil H, Owens C, Gutstein H, Taylor L, Curran E, Watson S (1998) Endogenous opioids: overview and current issues. Drug Alcohol Depend 51:127–140
Olson GA, Olson RD, Kastin AJ (1995) Endogenous opiates: 1994. Peptides 16:1517–1555
He X, Rosenfeld MG (1991) Mechanisms of complex transcriptional regulation: implications for brain development. Neuron 7:183–196
Kaplitt MG, Kwong AD, Kleopoulos SP, Mobbs CV, Rabkin SD, Pfaff DW (1994) Preproenkephalin promoter yields region-specific and long-term expression in adult brain after direct in vivo gene transfer via a defective herpes simplex viral vector. Proc Natl Acad Sci USA 91:8979–8983
Le Y, Gagneten S, Larson T, Santha E, Dobi A, v Agoston D, Sauer B (2003) Far-upstream elements are dispensable for tissue-specific proenkephalin expression using a Cre-mediated knock-in strategy. J Neurochem 84:689–697
Agoston D, Santha E, Shieh G, Lala R, Dobi A (1998) Isolation and structural and genetic analysis of the mouse enkephalin gene and its d(AC/TG)n repeats. DNA Seq 9:217–226
Joshi J, Sabol SL (1991) Proenkephalin gene expression in C6 rat glioma cells: potentiation of cyclic adenosine 3′, 5′-monophosphate-dependent transcription by glucocorticoids. Mol Endocrinol 5:1069–1080
Van Koughnet K, Smirnova O, Hyman SE, Borsook D (1999) Proenkephalin transgene regulation in the paraventricular nucleus of the hypothalamus by lipopolysaccharide and interleukin-1beta. J Comp Neurol 405:199–215
Agoston DV, Dobi A (2000) Complexity of transcriptional control in neuropeptide gene expression; enkephalin gene regulation during neurodevelopment. Biochem Soc Trans 28:446–451
Dobi L, Szemes M, Lee C, Palkovits M, Lim F, Gyorgy A, Mahan MA, Agoston DV (2005) AUF1 is expressed in cell- and development-specific manner in the developing brain, binds to AT-rich dsDNA and regulates enkephalin gene expression. J Biol Chem (submitted)
Ishii K, Laemmli UK (2003) Structural and dynamic functions establish chromatin domains. Mol Cell 11:237–248
Laemmli UK, Tjian R (1996) A nuclear traffic jam—unraveling multicomponent machines and compartments. Curr Opin Cell Biol 8:299–302
Saitoh Y, Laemmli UK (1993) From the chromosomal loops and the scaffold to the classic bands of metaphase chromosomes. Cold Spring Harb Symp Quant Biol 58:755–765
Dickinson LA, Kohwi ST (1995) Nucleolin is a matrix attachment region DNA-binding protein that specifically recognizes a region with high base-unpairing potential. Mol Cell Biol 15:456–465
Reeves R (2000) Structure and function of the HMGI(Y) family of architectural transcription factors. Environ Health Perspect 108:803–809
Dickinson LA, Dickinson CD, Kohwi ST (1997) An atypical homeodomain in SATB1 promotes specific recognition of the key structural element in a matrix attachment region. J Biol Chem 272:11463–11470
Dickinson LA, Joh T, Kohwi Y, Kohwi-Shigematsu T (1992) A tissue-specific MAR/SAR DNA-binding protein with unusual binding site recognition. Cell 70:631–645
Nakayama Y, Mian IS, Kohwi-Shigematsu T, Ogawa T (2005) A Nuclear targeting determinant for SATB1, a Genome Organizer in the T cell lineage. Cell Cycle 4:
Nie H, Maika SD, Tucker PW, Gottlieb PD (2005) A role for SATB1, a nuclear matrix association region-binding protein, in the development of CD8SP thymocytes and peripheral T lymphocytes. J Immunol 174:4745–4752
Yasui D, Miyano M, Cai S, Varga-Weisz P, Kohwi-Shigematsu T (2002) SATB1 targets chromatin remodelling to regulate genes over long distances. Nature 419:641–645
Boulikas T (1995) Chromatin domains and prediction of MAR sequences. Int Rev Cytol
Wilsker D, Patsialou A, Dallas PB, Moran E (2002) ARID proteins: a diverse family of DNA binding proteins implicated in the control of cell growth, differentiation, and development. Cell Growth Differ 13:95–106
Britanova O, Akopov S, Lukyanov S, Gruss P, Tarabykin V (2005) Novel transcription factor Satb2 interacts with matrix attachment region DNA elements in a tissue-specific manner and demonstrates cell-type-dependent expression in the developing mouse CNS. Eur J Neurosci 21:658–668
Kumar NV, Bernstein LR (2001) A new analytical scale DNA affinity binding assay for analyses of specific protein–DNA interactions. Anal Biochem 299:203–210
Hellman U, Wernstedt C, Gonez J, Heldin CH (1995) Improvement of an “In-Gel” digestion procedure for the micropreparation of internal protein fragments for amino acid sequencing. Anal Biochem 224:451–455
Dobi A, Palkovits M, Palkovits CG, Ring MA, Agoston D (2000) Septamer element-binding proteins in neuronal and glial differentiation. J Neurosci 20:1073–1084
Dobi A, Debnam W, Dalgard C, Owusu A, von Agoston D (2002) Mammalian expression cloning of nucleic acid binding proteins by agarose thin-layer gelshift clone selection. Biotechniques 33:868–872
Kas E, Poljak L, Adachi Y, Laemmli UK (1993) A model for chromatin opening: stimulation of topoisomerase II and restriction enzyme cleavage of chromatin by distamycin. Embo J 12:115–126
Galande S, Kohwi-Shigematsu T (2000) Caught in the act: binding of Ku and PARP to MARs reveals novel aspects of their functional interaction. Crit Rev Eukaryot Gene Expr 10:63–72
Soldatenkov VA, Potaman VN (2004) DNA-binding properties of poly(ADP-ribose) polymerase: a target for anticancer therapy. Curr Drug Targets 5:357–365
Galande S (2002) Chromatin (dis)organization and cancer: BUR-binding proteins as biomarkers for cancer. Curr Cancer Drug Targets 2:157–190
Podgornaya OI, Voronin AP, Enukashvily NI, Matveev IV, Lobov IB (2003) Structure-specific DNA-binding proteins as the foundation for three-dimensional chromatin organization. Int Rev Cytol 224:227–296
Bomsztyk K, Denisenko O, Ostrowski J (2004) hnRNP K: one protein multiple processes. Bioessays 26:629–638
Nakagomi K, Kohwi Y, Dickinson LA, Kohwi-Shigematsu T (1994) A novel DNA-binding motif in the nuclear matrix attachment DNA-binding protein SATB1. Mol Cell Biol 14:1852–1860
Dobreva G, Dambacher J, Grosschedl R (2003) SUMO modification of a novel MAR-binding protein, SATB2, modulates immunoglobulin mu gene expression. Genes Dev 17:3048–3061
FitzPatrick DR, Carr IM, McLaren L, Leek JP, Wightman P, Williamson K, Gautier P, McGill N, Hayward C, Firth H, Markham AF, Fantes JA, Bonthron DT (2003) Identification of SATB2 as the cleft palate gene on 2q32-q33. Hum Mol Genet 12:2491–2501
Georgopoulos K, Moore DD, Derfler B (1992) Ikaros, an early lymphoid-specific transcription factor and a putative mediator for T cell commitment. Science 258:808–812
Jossin Y (2004) Neuronal migration and the role of reelin during early development of the cerebral cortex. Mol Neurobiol 30:225–251
Uylings HB (2000) Development of the cerebral cortex in rodents and man. Eur J Morphol 38:309–312
Dobi AL, Palkovits M, Palkovits CG, Santha E, van Agoston D (1995) Protein–DNA interactions during phenotypic differentiation. Mol Neurobiol 10:185–203
Boulikas T (1992) Homeotic protein binding sites, origins of replication, and nuclear matrix anchorage sites share the ATTA and ATTTA motifs. J Cell Biochem 50:111–123
Boulikas T (1993) Nature of DNA sequences at the attachment regions of genes to the nuclear matrix. J Cell Biochem 52:14–22
Bode J, Kohwi Y, Dickinson L, Joh T, Klehr D, Mielke C, Kohwi-Shigematsu T (1992) Biological significance of unwinding capability of nuclear matrix- associating DNAs. Science 255:195–197
Strick R, Laemmli UK (1995) SARs are cis DNA elements of chromosome dynamics: synthesis of a SAR repressor protein. Cell 83:1137–1148
Strick T, Allemand J, Croquette V, Bensimon D (2000) Twisting and stretching single DNA molecules. Prog Biophys Mol Biol 74:115–140
Kortschak RD, Tucker PW, Saint R (2000) ARID proteins come in from the desert. Trends Biochem Sci 25:294–299
Cai S, Han HJ, Kohwi-Shigematsu T (2003) Tissue-specific nuclear architecture and gene expression regulated by SATB1. Nat Genet 34:42–51
Wen J, Huang S, Rogers H, Dickinson LA, Kohwi-Shigematsu T, Noguchi CT (2005) SATB1 family protein expressed during early erythroid differentiation modifies globin gene expression. Blood 105:3330–3339
Brown KE (2003) Chromatin folding and gene expression: new tools to reveal the spatial organization of genes. Chromosome Res 11:423–433
de Laat W, Grosveld F (2003) Spatial organization of gene expression: the active chromatin hub. Chromosome Res 11:447–459
Recillas-Targa F, Razin SV (2001) Chromatin domains and regulation of gene expression: familiar and enigmatic clusters of chicken globin genes. Crit Rev Eukaryot Gene Expr 11:227–242
Agoston DV, Szemes M, Dobi A, Palkovits M, Georgopoulos K and, Ring M (2005) Ikaros, the Hematopoietic master regulator is expressed in developing striatal neurons and involved in enkephalinergic differentiation. J Neurosci submitted
Georgopoulos K, Bigby M, Wang JH, Molnar A, Wu P, Winandy S, Sharpe A (1994) The Ikaros gene is required for the development of all lymphoid lineages. Mol Cell Biol 14:7111–7123
Georgopoulos K (2002) Haematopoietic cell-fate decisions, chromatin regulation and ikaros. Nature Rev Immunol 2:162–174
Acknowledgements
The authors would like to thank Mr. Cheol Lee for his help and advice with histological techniques, Mr. Michael Lawson for the technical assistance with the immunohistochemical stainings and Dr. Jozsef Czege (BIC, USUHS) for his expert help with collecting and processing histological images. The work was funded by an intramural grant from USUHS.
Author information
Authors and Affiliations
Corresponding author
Additional information
The first two authors contributed equally to this work.
Special Issue Dedicated to Miklós Palkovits.
Rights and permissions
About this article
Cite this article
Szemes, M., Gyorgy, A., Paweletz, C. et al. Isolation and Characterization of SATB2, a Novel AT-rich DNA Binding Protein Expressed in Development- and Cell-Specific Manner in the Rat Brain. Neurochem Res 31, 237–246 (2006). https://doi.org/10.1007/s11064-005-9012-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-005-9012-8