Abstract
Purpose
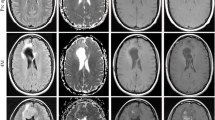
Texture analysis (TA) can quantify variations in surface intensity or patterns, including some that are imperceptible to the human visual system. The purpose of this study was to determine the diagnostic accuracy of radiomic based filtration-histogram TA to differentiate high-grade from low-grade gliomas by assessing tumor heterogeneity.
Methods
Patients with a histopathological diagnosis of glioma and preoperative 3T MRI imaging were included in this retrospective study. A region of interest was manually delineated on post-contrast T1 images. TA was performed using commercially available research software. The histogram parameters including mean, standard deviation, entropy, mean of the positive pixels, skewness, and kurtosis were analyzed at spatial scaling factors ranging from 0 to 6 mm. The parameters were correlated with WHO glioma grade using Spearman correlation. Areas under the curve (AUC) were calculated using ROC curve analysis to distinguish tumor grades.
Results
Of a total of 94 patients, 14 had low-grade gliomas and 80 had high-grade gliomas. Mean, SD, MPP, entropy and kurtosis each showed significant differences between glioma grades for different spatial scaling filters. Low and high-grade gliomas were best-discriminated using mean of 2 mm fine texture scale, with a sensitivity and specificity of 93% and 86% (AUC of 0.90).
Conclusions
Quantitative measurement of heterogeneity using TA can discriminate high versus low-grade gliomas. Radiomic data of texture features can provide complementary diagnostic information for gliomas.



Similar content being viewed by others
References
Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, Gronroos E, Martinez P, Matthews N, Stewart A, Tarpey P, Varela I, Phillimore B, Begum S, McDonald NQ, Butler A, Jones D, Raine K, Latimer C, Santos CR, Nohadani M, Eklund AC, Spencer-Dene B, Clark G, Pickering L, Stamp G, Gore M, Szallasi Z, Downward J, Futreal PA, Swanton C (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. New Eng J Med 366(10):883–892. https://doi.org/10.1056/NEJMoa1113205
Friedmann-Morvinski D (2014) Glioblastoma heterogeneity and cancer cell plasticity. Crit Rev Oncog 19(5):327–336
Inda MM, Bonavia R, Seoane J (2014) Glioblastoma multiforme: a look inside its heterogeneous nature. Cancers 6(1):226–239. https://doi.org/10.3390/cancers6010226
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109. https://doi.org/10.1007/s00401-007-0243-4
Raab SS, Grzybicki DM, Janosky JE, Zarbo RJ, Meier FA, Jensen C, Geyer SJ (2005) Clinical impact and frequency of anatomic pathology errors in cancer diagnoses. Cancer 104(10):2205–2213. https://doi.org/10.1002/cncr.21431
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577. https://doi.org/10.1148/radiol.2015151169
Hu LS, Ning S, Eschbacher JM, Gaw N, Dueck AC, Smith KA, Nakaji P, Plasencia J, Ranjbar S, Price SJ, Tran N, Loftus J, Jenkins R, O’Neill BP, Elmquist W, Baxter LC, Gao F, Frakes D, Karis JP, Zwart C, Swanson KR, Sarkaria J, Wu T, Mitchell JR, Li J (2015) Multi-parametric MRI and texture analysis to visualize spatial histologic heterogeneity and tumor extent in glioblastoma. PLoS ONE 10(11):e0141506. https://doi.org/10.1371/journal.pone.0141506
Miles KA, Ganeshan B, Griffiths MR, Young RC, Chatwin CR (2009) Colorectal cancer: texture analysis of portal phase hepatic CT images as a potential marker of survival. Radiology 250(2):444–452. https://doi.org/10.1148/radiol.2502071879
Parikh J, Selmi M, Charles-Edwards G, Glendenning J, Ganeshan B, Verma H, Mansi J, Harries M, Tutt A, Goh V (2014) Changes in primary breast cancer heterogeneity may augment midtreatment MR imaging assessment of response to neoadjuvant chemotherapy. Radiology 272(1):100–112. https://doi.org/10.1148/radiol.14130569
Win T, Miles KA, Janes SM, Ganeshan B, Shastry M, Endozo R, Meagher M, Shortman RI, Wan S, Kayani I, Ell PJ, Groves AM (2013) Tumor heterogeneity and permeability as measured on the CT component of PET/CT predict survival in patients with non-small cell lung cancer. Clin Cancer Res 19(13):3591–3599. https://doi.org/10.1158/1078-0432.ccr-12-1307
Yip C, Landau D, Kozarski R, Ganeshan B, Thomas R, Michaelidou A, Goh V (2014) Primary esophageal cancer: heterogeneity as potential prognostic biomarker in patients treated with definitive chemotherapy and radiation therapy. Radiology 270(1):141–148. https://doi.org/10.1148/radiol.13122869
Wang S, Meng M, Zhang X, Wu C, Wang R, Wu J, Sami MU, Xu K (2018) Texture analysis of diffusion weighted imaging for the evaluation of glioma heterogeneity based on different regions of interest. Oncol Lett 15(5):7297–7304. https://doi.org/10.3892/ol.2018.8232
Juntu J, Sijbers J, De Backer S, Rajan J, Van Dyck D (2010) Machine learning study of several classifiers trained with texture analysis features to differentiate benign from malignant soft-tissue tumors in T1-MRI images. J Magn Reson Imaging 31(3):680–689. https://doi.org/10.1002/jmri.22095
Skogen K, Schulz A, Dormagen JB, Ganeshan B, Helseth E, Server A (2016) Diagnostic performance of texture analysis on MRI in grading cerebral gliomas. Eur J Radiol 85(4):824–829. https://doi.org/10.1016/j.ejrad.2016.01.013
Cohen JF, Korevaar DA, Altman DG, Bruns DE, Gatsonis CA, Hooft L, Irwig L, Levine D, Reitsma JB, de Vet HC, Bossuyt PM (2016) STARD 2015 guidelines for reporting diagnostic accuracy studies: explanation and elaboration. BMJ Open 6(11):e012799. https://doi.org/10.1136/bmjopen-2016-012799
Ganeshan B, Miles KA (2013) Quantifying tumour heterogeneity with CT. Cancer Imaging 13:140–149. https://doi.org/10.1102/1470-7330.2013.0015
Ganeshan B, Goh V, Mandeville HC, Ng QS, Hoskin PJ, Miles KA (2013) Non-small cell lung cancer: histopathologic correlates for texture parameters at CT. Radiology 266(1):326–336. https://doi.org/10.1148/radiol.12112428
Chen B, Zhang R, Gan Y, Yang L, Li W (2017) Development and clinical application of radiomics in lung cancer. Radiat Oncol 12(1):154. https://doi.org/10.1186/s13014-017-0885-x
Stoyanova R, Takhar M, Tschudi Y, Ford JC, Solorzano G, Erho N, Balagurunathan Y, Punnen S, Davicioni E, Gillies RJ, Pollack A (2016) Prostate cancer radiomics and the promise of radiogenomics. Transl Cancer Res 5(4):432–447
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue R, Even AJG, Jochems A, van Wijk Y, Woodruff H, van Soest J, Lustberg T, Roelofs E, van Elmpt W, Dekker A, Mottaghy FM, Wildberger JE, Walsh S (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14(12):749–762. https://doi.org/10.1038/nrclinonc.2017.141
Ryu YJ, Choi SH, Park SJ, Yun TJ, Kim JH, Sohn CH (2014) Glioma: application of whole-tumor texture analysis of diffusion-weighted imaging for the evaluation of tumor heterogeneity. PLoS ONE 9(9):e108335. https://doi.org/10.1371/journal.pone.0108335
Raja R, Sinha N, Saini J, Mahadevan A, Rao KN, Swaminathan A (2016) Assessment of tissue heterogeneity using diffusion tensor and diffusion kurtosis imaging for grading gliomas. Neuroradiology 58(12):1217–1231. https://doi.org/10.1007/s00234-016-1758-y
Donahue MJ, Blakeley JO, Zhou J, Pomper MG, Laterra J, van Zijl PC (2008) Evaluation of human brain tumor heterogeneity using multiple T1-based MRI signal weighting approaches. Magn Reson Med 59(2):336–344. https://doi.org/10.1002/mrm.21467
Ng F, Kozarski R, Ganeshan B, Goh V (2013) Assessment of tumor heterogeneity by CT texture analysis: can the largest cross-sectional area be used as an alternative to whole tumor analysis? Eur J Radiol 82(2):342–348. https://doi.org/10.1016/j.ejrad.2012.10.023
Lubner MG, Stabo N, Lubner SJ, del Rio AM, Song C, Halberg RB, Pickhardt PJ (2015) CT textural analysis of hepatic metastatic colorectal cancer: pre-treatment tumor heterogeneity correlates with pathology and clinical outcomes. Abdom Imaging 40(7):2331–2337. https://doi.org/10.1007/s00261-015-0438-4
Chen W, Giger ML, Li H, Bick U, Newstead GM (2007) Volumetric texture analysis of breast lesions on contrast-enhanced magnetic resonance images. Magn Reson Med 58(3):562–571. https://doi.org/10.1002/mrm.21347
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Funding
Funding for this study was provided by Pilot grant, Department of Radiology, University of Cincinnati. Dr. Vagal has received research funding unrelated to this study from R01 NH/NINDS NS100417, R01 NIH/NINDS NS30678.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ditmer, A., Zhang, B., Shujaat, T. et al. Diagnostic accuracy of MRI texture analysis for grading gliomas. J Neurooncol 140, 583–589 (2018). https://doi.org/10.1007/s11060-018-2984-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2984-4