Abstract
Spalt-like transcription factor 4 (SALL4), a oncogene, is known to participate in multiple carcinomas, and is up-regulated in glioma. However, its actual role and underlying mechanisms in the development of glioma remain unclear. The present study explored the molecular functions of SALL4 in promoting cell proliferation in glioma. The expression level of SALL4 in 69 human glioma samples and six non-tumor brain tissues was determined using real-time polymerase chain reaction (PCR). Then, we transfected U87 and U251 cell lines with siRNA, and assessed cellular proliferation and cell cycle to understand the function of SALL4, and the relationship between SALL4, PTEN and PI3K/AKT pathway. PCR confirmed that the expression of SALL4 was higher in the glioma samples than non-tumor brain tissues. Cellular growth and proliferation were dramatically reduced following inhibition of SALL4 expression. Western blot showed increase in PTEN expression when SALL4 was silenced, which in turn depressed the activation of PI3K/AKT pathway, suggesting that PTEN was a downstream target of SALL4 in glioma development. Therefore, SALL4 could act as a proto-oncogene by regulating the PTEN/PI3K/AKT signaling pathway, thereby facilitating proliferation of glioma cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Malignant glioma has the highest incidence among human primary brain tumors, and is characterized by high mortality rate, recurrence and malignancy. In spite of comprehensive therapies, the prognosis and survival of glioma patients remain poor [1]. Malignant growth, high proliferation of glioma cells and high infiltration that makes full surgical resection impossible are the predominant reasons for poor prognosis and survival. Like other types of tumors, the causes of glioma are varied, and include activation of oncogenes. The embryonic stem cell (ESC) gene SALL4 has been recently identified as a new target for cancer therapy.
SALL4 is the human homolog of Drosophila spalt (sal) mapped to chromosome 20q13 and encodes a C2H2 zinc-finger transcription factor [2], which is important for maintenance of pluripotent and self-renewal properties of ESCs [3]. With the same function of oncogenes, SALL4 participates in cell proliferation, apoptosis, cycle, invasion, drug resistance, and the formation and evolution [4,5,6,7] of multiple human solid tumors, such as hematopoiesis, hepatocellular carcinoma, lung cancer, myelodysplastic syndrome [8,9,10].
Phosphatase and tension homolog (PTEN), is a tumor suppressor whose expression is very low in various human tumors [11,12,13]. The PI3K/AKT signaling pathway is a well-known pathway in the regulation of tumorigenesis, and is significantly activated in glioma [14]. PTEN contributes in antagonizing PI3K [15], thereby weakening AKT activation [16], which could suppress down-stream products thereby inducing cell cycle arrest in the G1 phase by increasing ki-67 expression [15] and decreasing cyclin D1 expression [17].
Based on the important function of PI3K/AKT signaling in glioma development [18, 19] and the crosstalk between SALL4 and PTEN [20], we found that SALL4 mRNA expression was significantly higher in glioma specimens than in non‑cancerous brain samples. SALL4 expression may promote the formation of glioma, but the underlying mechanism remains unclear. The present study was based on the hypothesis that SALL4 could suppress PTEN, thereby strengthening PI3K/AKT signaling.
Materials and methods
Human tissue samples
Specimens were collected from patients who underwent surgical removal of brain tumors at the Department of Neurosurgery, Brain and Nerve Research Laboratory of The First Affiliated Hospital of Soochow University (Suzhou, China) from 2009 to 2012. Six non-tumor brain samples were collected from patients without brain tumors who underwent traumatic brain injury or arteriovenous malformation, which needed resection of a small part of their brain tissues to lower the intracranial hypertension and increase treatment outcome. Thirty-seven female and 32 male glioma patients were included. Among them, 17 had grade II (diffuse astrocytoma), 26 had grade III (anaplastic astrocytoma), and 26 had grade IV (primary brain glioblastoma), according to the 2007 WHO classification system. The mean age of the patients at the time of surgical resection were 46.9 years for men and 44.9 years for women. The mean age was 40.62 ± 15.64 years for grade II, 43.89 ± 15.21 for grade III and 48.12 ± 14.97 years for grade IV. All samples were collected and immediately stored in liquid nitrogen after resection. This study was approved by the local ethics committee of The First Affiliated Hospital of Soochow University, and all patients gave informed consent for the usage of their samples in the study.
Cell cultures and treatments
The U87MG and U251MG were obtained from the Cell Bank Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). Cells were maintained in DMEM (Hyclone, Thermo Fisher Scientific, USA) supplemented with 10% FBS (Gibco, Invitrogen, USA) at 37 °C under a humidified atmosphere of 5% CO2.
siRNA transfection
For down-regulation of SALL4, 50 pmol/l SALL4-siRNA, filtrating the best one from three different kings of SALL4-siRNA (siRNA-1:5-CCGAAAGCAUCAA GUCAAATT-3;5-UUUGACUUGAUGCUUUCGGTT-3. siRNA-2:5-GUCUCUGGAUGCCUGAAATT-3; 5-UUUCAAGGCAUCCAGAGACTT-3. siRNA-3:5-GUGGCCAACACUAAUGUGATT-3; 5-UCACAUUAGUGUUGGCCACTT-3) were transfected into the cells using Lipofectamine 2000 (invitrogen) according to the manufacturer’s instructions. The siRNA vectors were are purchased from Shanghai Genepharma Co., Ltd. The transfection rates of two human glioma cell lines U87 and U251 were determined by flow cytometry. Transfection ratio >80% was used for the experiments (the U87 transfection efficiency was 97.8% and the U251 transfection efficiency was 99.8%).
Quantitative RT-PCR
RNA from cells and specimens was extracted by TRIzol reagent (Invitrogen, USA), and quantified by spectrophotometer. Only mRNA with 260/280 ratios of 1.9–2.0 were used for the experiments. Relative levels of mRNA were examined using SYBR green real-time quantitative RT-PCR (qRT-PCR) (LightCycle r480 Roche, Switzerland), and normalized by GAPDH mRNA. qRT-PCR results were calculated using the 2−△△CT method, data analyses were performed in triplicate. Three times independent experiments are repeated and all data are presented as means and standard errors of the means.
Flow cytometric analysis
Glioma cell lines U87 and U251 were cultured in 6-well plates, at a cell density of 2 × 105 per well, to ensure a transfection density of 80–90%. After culturing for 24 h, transfection efficiency was detected. Cells were transfected with siRNA negative control, SALL4-siRNA and SALL4-siRNA-bpv (PTEN inhibitor bpv were add into SALL4-siRNA group), and incubated for 48 h before the cell cycle test. Both transfection efficiency and cell cycle were measured by flow cytometer and repeated three times this experiments.
CCK-8 assay
The proliferation of cells was detected by cell counting kit (CCK-8, Dojindo, China). siRNA negative control, SALL4-siRNA and SALL4-siRNA-bpv were transfected into the U87 and U251 glioma cells. After incubating for 12, 24, 48 and 72 h, CCK-8 was added and the absorbance was measured at 450 nm (Thermo, USA) after incubating for another 2 h. The OD was calculated as mean ± SD of three measurements per sample.
Western blotting
Fourty-eight hours after SALL4-siRNA and negative control siRNA transfection or 24 h before addition of PTEN inhibitors pten (bpv), the whole protein lysate was prepared from cells by radioimmunoprecipitation assay lysis buffer containing 50 mM Tris/HCl pH 7.5, 0.1% sodium dodecyl sulfate (SDS), 1% Triton-X 100, 0.5–1% sodium deoxycholate, 150 mM sodium chloride, and protease inhibitors (Roche). Bicinchoninic acid (BCA) protein assay kit (Beyotime Institute of Biotechnology) was used to calculate the protein concentration. Equal amounts of protein samples were separated with 6–10% SDS-PAGE and transferred onto nitrocellulose membrane, under constant voltage of 60 V. The membrane was blocked with 5% bovine serum albumin (BSA) (Amresco, USA) for 2 h at room temperature, and then incubated with 1:1000 primary antibodies overnight at 4 °C and 1:3000 HRP-conjugated secondary antibody (Beyotime Institute of Biotechnology) for 2 h followed by washing with PBST three times for 5 min each time. The signal was detected using enhanced chemiluminescence (ECL) (Thermo). The primary antibodies used in this study were anti-SALL4, anti-cyclin D1 (Abcam, Japan), anti-PTEN, anti-PI3K, anti-p-PI3K, anti-AKT, anti-p-AKT, and anti-GAPDH (CST, USA). Repeated three times this experiments.
Immunofluorescence analysis
U87 and U251 cells were transfected with SALL4-siRNA, negative control and SALL4-siRNA-bpv, treated with bpv, fixed with 4% paraformaldehyde for 30 min, and blocked with BSA (Amresco) for 30 min. The cells were incubated with anti-PTEN, anti-p-PI3K (CST, USA) primary antibodies at 4 °C overnight, and then incubated with tetramethylrhodamine isothiocyanate-labeled secondary antibody (diluted 1:500) at 37 °C for 30 min. The cells were stained with DAPI, and imaged with a fluorescence microscope (OLYMPUS BX50/BXFLA/DP70; Olympus Co., Japan). Three times independent experiments are repeated.
Statistical analysis
Statistical analyses were conducted with GraphPad, PRISM4.0 software (GraphPad, USA) and SPSS 13.0 software (SPSS Inc., USA). Student’s t-test or ANOVA were used to test the difference between the groups. P < 0.05 was considered to be statistically significant.
Results
Expression of SALL4 mRNA in glioma samples and non-tumor brain tissues
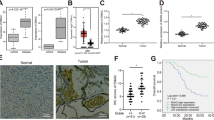
qPCR was performed to quantify the expression of SALL4 in 69 glioma samples and six non-tumor brain tissues. The expression of SALL4 was higher in the glioma samples than in the non-tumor brain tissues, and increased with the increase in degree of malignancy in glioma (P < 0.05; Fig. 1a). SALL4 level was detected in different types of glioma cell lines(SHG139, SHG44, U87, U251, A172) and the more malignant the cell, the higher was the expression of SALL4. U87 and U251 cell lines were selected for the present study (Fig. 1b).
a shows qRT-PCR analysis (data were reported as 2−△△CT) of the expression of SALL4 in six non-tumor brain tissues and 69 glioma tissues. SALL4 expression was markedly higher in glioma than in non-tumor brain tissues and increased with the increase in degree of malignancy in glioma. b SALL4 expression in glioma cell lines was up-regulated as compared to normal brain tissues (NBT including N1, N2 and N3). N1, N2 and N3 small nuclear RNA was used as an internal control. *P < 0.05, **P < 0.01. Higher grade glioma cells had higher level of SALL4
Relationship between SALL4 and PTEN in glioma
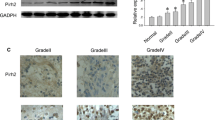
PTEN was shown to be regulated by SALL4 in some types of tumors. Li found the cancer suppressor gene PTEN was obviously down-regulated in glioma [21]. To estimate the effect of SALL4 on PTEN, firstly, the most efficient siRNA was selected from the three siRNA sequences (P < 0.05; Fig. 2a). Secondly, mRNA of U87 and U251 cells transfected with SALL4-siRNA were used for qRT-PCR, which showed that the expression of PTEN was apparently up-regulated when SALL4 was blocked (P < 0.05; Fig. 2b), implying a negative correlation between PTEN and SALL4. Western blot also demonstrated that PTEN protein was significantly higher in cells transfected with SALL4-siRNA as compared to siRNA negative control and blank groups (P < 0.05; Fig. 2c, d). Based on these findings, the expression of SALL4 might suppress the mRNA and protein levels of PTEN.
siRNA-SALL4 reduces proliferation of glioma cells
Rapid proliferation of glioma cells leads to poor prognosis and short survival. Cellular proliferation assays in U87 and U251 cells were conducted using CCK-8 to explore the influence SALL4 on growth of glioma cells. SALL4 expression was decreased after transfection with SALL4-siRNA, which in turn resulted in significant decline in proliferation of glioma cells transfected with SALL4-siRNA (P < 0.05; Fig. 3a, b). However, this inhibition of proliferation was reversed by the PTEN inhibitor phen (bpv), here, this group we entitled SALL4-siRNA-bpv. Therefore, SALL4 can promote proliferation of glioma cells, and down-regulation of SALL4 could suppress cell growth.
a Cell counting kit (CCK-8) was used to detect cellular proliferation in U87 and U251 cells at 6, 24, 48, and 72 h after siRNA transfection. b Shows the OD value at the time of 72 h and data were reported as means ± SD (*P < 0.01). c, d Blocking SALL4 induced U87 and U251 at the time of 48 h, cell cycle arrest in the G1 phase, which decreased cells in the S phase, increased cells in the G1 phase. Cell cycle in the cells treated with SALL4-siRNA and phen (bpv) were similar to blank and negative control groups (P < 0.05)
Down-regulation of SALL4 could induce cell cycle arrest at G1 phase
SALL4 could work as a promoter for tumor formation in various human tumors including glioma, but the mechanism was unclear. Cell cycle analysis was performed in U87 and U251 cells to better understand the cell cycle modulation. Outcomes revealed that G1 phase was increased and S phase was decreased (P < 0.05; Fig. 3c, d) in cells where SALL4 was blocked. And in cells with SALL4-siRNA-bpv, the repression effect was significantly relieved. Hence, inhibition of SALL4 expression in glioma by SALL4-siRNA arrested the cell cycle at G1 phase, and inhibited cell proliferation as seen by increased percentage of G1 phase cells and decreased S phase cells.
Down-regulation of SALL4 could suppress the activation of PTEN/PI3K/AKT signaling pathway
Down-regulation of SALL4 could increase the expression of PTEN. PTEN is an anti-tumor gene that depresses the PI3K/AKT signaling pathway in many cancers, but its role in glioma is unknown. In the present study, western blot results suggested that p-PI3K, p-AKT and cyclin D1 levels in cells transfected with SALL4-siRNA were significantly lower than in cells transfected with negative control, while the levels were moderate in SALL4-siRNA-bpv transfected cells (P < 0.05; Fig. 4a). Therefore, down-regulation of SALL4 expression suppressed activation of PTEN/PI3K/AKT pathway, which in turn impeded cell proliferation.
a The expression of SALL4, PTEN, PI3K, p-PI3K, AKT, p-AKT, cyclin D1 and GAPDH in different groups were detected by western blot analysis. The activity of PI3K/AKT signaling pathway was obviously decreased in cells transfected with SALL4-siRNA. b Immunofluorescence analysis data showed the expression of PTEN and p-PI3K after different treatment showed by bar graph
PTEN and p-PI3K level was altered after blocking SALL4 in glioma cells
Immunohistochemistry showed that the expression of PTEN was increased, and p-PI3K was decreased in U87 and U251 cells after knocking down SALL4 as compared to the negative control group. When PTEN inhibitor phen (bpv) was added to the cells transfected with SALL4-siRNA, the protein level was similar to the negative control group (P < 0.05; Fig. 5a–d).
Discussion
SALL4 is known to be abnormally expressed in multiple human tumors. In our study, abnormal expression of SALL4 was confirmed in glioma samples. Furthermore, blocking SALL4 increased PTEN expression and restrained the activation of PI3K/AKT pathway, thus suppressing cellular growth and proliferation of glioma.
SALL4 is expressed in the early stages of fetal development, and then diminishes as differentiation proceeds, with very low levels found in adults [22]. Up- or down-regulation of SALL4 is involved in cell proliferation, invasion, drug resistance, apoptosis, and other processes in some malignancies by targeting related genes [2, 23,24,25,26]. Owing to the strong proliferative function of SALL4 on cells and tissues, its correlation with various tumors was examined. SALL4 was aberrantly elevated in multiple carcinomas, such as leukemia, germ cell tumors, liver cancer and gastric cancer [2, 27, 28], acting as an oncogene and biomarker [23]. AJAY considered that SALL4 may be an extremely useful diagnostic marker in lung cancer. SALL4 played an important role in ESCs and human tumors. Abnormally high expression of SALL4 was closely related to tumor formation and prognosis in hematopoiesis and leukemogenesis, but SALL4 was also required for DNA damage response in ESCs, ensuring their stability during expansion [29]. SALL4 was a potent stimulator for the expansion of human hematopoietic stem/progenitor cells, esophageal squamous cell and gastric cancers [30, 31]. Over-expression of SALL4 enhanced gastric cancer cell proliferation and migration, whereas knocking down SALL4 reversed these effects [27]. Park found that SALL4 expression was significantly associated with a poor overall survival as compared to SALL4-negative HCCs [32], and implied poor prognosis in human hepatocellular and endometrial cancers. Down-regulation of SALL4 expression using small-hairpin RNA led to decreased in vitro myeloid colony-forming abilities and impaired in vivo engraftment in normal primary CD34+ cells [33]. SALL4 is a therapeutic target in intrahepatic cholangiocarcinoma (ICC) and endometrial cancer. However, its underlying mechanism during the development of ESCs and human tumors remains indistinct. Accumulating evidence reveals a crucial role for SALL4 in leukemogenesis due to its ability to promote proliferation and participation in Wnt/β-catenin pathways [34]. Wang believed that regulating NANOG, OCT4, and SOX2 may account for maintenance of pluripotent and self-renewal properties of ESCs. Hong W and Lauberth SM confirmed that by recruiting Mi-2/Nucleosome Remodeling and Deacetylase (NuRD) complex, SALL4 mediated transcription to impact cell growth [20].
PI3K/AKT signaling is a classical pathway involved in the regulation of tumorigenesis, hypoxia-induced VM formation, migration, invasion, and proliferation due to increase in the expression of phosphorylated EGFR (pEGFR), PI3K (p-PI3K), and AKT (p-AKT) [35,36,37,38]. Inhibitors of PI3K, for example, LY 294002 and PTEN, blocked the PI3K/AKT signaling and suppressed growth of diverse malignant tumors, including gliomas [14, 39]. Phen (bpv) is a known PTEN inhibitor that antagonizes PTEN function, and reverses the effect of PTEN as a cancer suppressor gene. Phosphorylated PI3K/AKT can promote cell growth and decrease apoptosis via increased Bcl-2/BAX ratio in human chondrosarcoma (CS) [40]. Cyclin D1 can be induced by growth factors through activation of various signaling pathways including PI3K/AKT, NF-kB [41,42,43], and p-AKT plays a crucial role in inducing cell proliferation by modulation of cyclin D1 in primary murine keratinocytes [43, 44]. In the present study, PTEN expression was up- regulation when SALL4 was reduced by siRNA-SALL4, as a result, the inhibitory action of PTEN on PI3K/AKT signaling was weaken, thus receded the cycling D1 level which arrested the cell cycle at G1 phase, regulating glioma proliferation. So, our study disclosed that SALL4 can inhibit PTEN and promote PI3K/AKT pathway in glioma, similar to leukemia [24].
In our text, SALL4 was found lower expression in some kinds of glioma samples than non-tumor brain tissues, a certain extent like the SALL4-positive immunoreactivity was 58% in total 102 intrahepatic cholangiocarcinoma cases [45]. Expression has been also reported SALL4 does not seem to be expressed in all trophoblastic tumors [46], individual genetic difference may be responsible for this phenomenon based on our relatively small sample, the more samples the more valuable outcome can be obtain.
Conclusion
Our study showed significantly high expression of SALL4 mRNA in glioma specimens as compared to non-tumor samples using RT-PCR. Blocking SALL4 using SALL4-siRNA decreased proliferation of U87 and U251 cells, which was reversed by the addition of PTEN inhibitor phen (bpv). Furthermore, marked increase in PTEN mRNA and protein levels was seen in cells treated with siRNA-SALL4. These findings suggest that SALL4 can facilitate cell growth by suppressing PTEN expression in glioma cell lines. The PI3K/AKT activity was decreased in cells treated with pten (bpv) after transfecting with SALL4-siRNA, whereby p-PI3K and p-AKT protein levels were significantly increased as compared to cells transfected with SALL4-siRNA only. Therefore, blocking SALL4 could promote PTEN expression and restrain PI3K/AKT activity resulting in suppression of cell growth.
References
Avni D, Glucksam Y, Zor T (2012) The phosphatidylinositol 3-kinase (PI3K) inhibitor LY294002 modulates cytokine expression in macrophages via p50 nuclear factor kappaB inhibition, in a PI3K-independent mechanism. Biochem Pharmacol 83(1):106–114. doi:10.1016/j.bcp.2011.09.025
Yong KJ, Gao C, Lim JS, Yan B, Yang H, Dimitrov T, Kawasaki A, Ong CW, Wong KF, Lee S, Ravikumar S, Srivastava S, Tian X, Poon RT, Fan ST, Luk JM, Dan YY, Salto-Tellez M, Chai L, Tenen DG (2013) Oncofetal gene SALL4 in aggressive hepatocellular carcinoma. N Engl J Med 368(24):2266–2276. doi:10.1056/NEJMoa1300297
Gao C, Dimitrov T, Yong KJ, Tatetsu H, Jeong HW, Luo HR, Bradner JE, Tenen DG, Chai L (2013) Targeting transcription factor SALL4 in acute myeloid leukemia by interrupting its interaction with an epigenetic complex. Blood 121(8):1413–1421. doi:10.1182/blood-2012-04-424275
Chen YY, Li ZZ, Ye YY, Xu F, Niu RJ, Zhang HC, Zhang YJ, Liu YB, Han BS (2016) Knockdown of SALL4 inhibits the proliferation and reverses the resistance of MCF-7/ADR cells to doxorubicin hydrochloride. BMC Mol Biol 17(1):6. doi:10.1186/s12867-016-0055-y
Zhang D, Jiang F, Wang X, Li G (2017) Knockdown of SALL4 inhibits proliferation, migration and invasion in osteosarcoma cells. Oncol Res Featur Preclin Clin Cancer Ther 25(5):763–771. doi:10.3727/096504016X14772402056137
He J, Zhang W, Zhou Q, Zhao T, Song Y, Chai L, Li Y (2013) Low-expression of microRNA-107 inhibits cell apoptosis in glioma by upregulation of SALL4. Int J Biochem cell Biol 45(9):1962–1973. doi:10.1016/j.biocel.2013.06.008
Liu L, Zhang J, Yang X, Fang C, Xu H, Xi X (2015) SALL4 as an epithelial-mesenchymal transition and drug resistance inducer through the regulation of c-Myc in endometrial cancer. PloS one 10(9):e0138515. doi:10.1371/journal.pone.0138515
Milanovich S, Peterson J, Allred J, Stelloh C, Rajasekaran K, Fisher J, Duncan SA, Malarkannan S, Rao S (2015) Sall4 overexpression blocks murine hematopoiesis in a dose-dependent manner. Exp Hematol 43(1):53–64. doi:10.1016/j.exphem.2014.09.004
Yin F, Han X, Yao SK, Wang XL, Yang HC (2016) Importance of SALL4 in the development and prognosis of hepatocellular carcinoma. World J Gastroenterol 22(9):2837–2843. doi:10.3748/wjg.v22.i9.2837
Liu H, Bai Y, Wang W, Guo L (2014) Role of SALL4 in regulating multi-drug resistance of small cell lung cancer and its clinical significance. Zhonghua bing li xue za zhi 43 (9):604–608. doi:10.3760/cma.j.issn.0529-5807.2014.09.007
Liu S, Ma X, Ai Q, Huang Q, Shi T, Zhu M, Wang B, Zhang X (2013) NOTCH1 functions as an oncogene by regulating the PTEN/PI3K/AKT pathway in clear cell renal cell carcinoma. Urol Oncol 31(6):938–948. doi:10.1016/j.urolonc.2011.07.006
Yong Ji, Mingfeng Zheng, Shugao Ye, Jingyu Chen, Yijiang Chen (2014) PTEN and Ki67 expression is associated with clinicopathologic features of non-small cell lung cancer. J Biomed Res 28 (6):462–467. doi:10.7555/JBR.27.20130084
Lin PC, Lin JK, Lin HH, Lan YT, Lin CC, Yang SH, Chen WS, Liang WY, Jiang JK, Chang SC (2015) A comprehensive analysis of phosphatase and tensin homolog deleted on chromosome 10 (PTEN) loss in colorectal cancer. World J Surg Oncol 13:186. doi:10.1186/s12957-015-0601-y
Dasari VR, Kaur K, Velpula KK, Gujrati M, Fassett D, Klopfenstein JD, Dinh DH, Rao JS (2010) Upregulation of PTEN in glioma cells by cord blood mesenchymal stem cells inhibits migration via downregulation of the PI3K/Akt pathway. PloS one 5(4):e10350. doi:10.1371/journal.pone.0010350
Ahmed MW, Kayani MA, Shabbir G, Ali SM, Shinwari WU, Mahjabeen I (2016) Expression of PTEN and its correlation with proliferation marker Ki-67 in head and neck cancer. Int J Biol Mark 31(2):e193–e203. doi:10.5301/jbm.5000196
Zhang LL, Mu GG, Ding QS, Li YX, Shi YB, Dai JF, Yu HG (2015) Phosphatase and tensin homolog (PTEN) represses colon cancer progression through inhibiting paxillin transcription via PI3K/AKT/NF-kappaB pathway. J Biol Chem 290(24):15018–15029. doi:10.1074/jbc.M115.641407
Feng XJ, Liu SX, Wu C, Kang PP, Liu QJ, Hao J, Li HB, Li F, Zhang YJ, Fu XH, Zhang SB, Zuo LF (2014) The PTEN/PI3K/Akt signaling pathway mediates HMGB1-induced cell proliferation by regulating the NF-kappaB/cyclin D1 pathway in mouse mesangial cells. Am J Physiol Cell Physiol 306(12):C1119–C1128. doi:10.1152/ajpcell.00385.2013
Nicoletti NF, Erig TC, Zanin RF, Pereira TC, Bogo MR, Campos MM, Morrone FB (2014) Mechanisms involved in kinin-induced glioma cells proliferation: the role of ERK1/2 and PI3K/Akt pathways. J Neurooncol 120(2):235–244. doi:10.1007/s11060-014-1549-4
Yang J, Yang Q, Yu J, Li X, Yu S, Zhang X (2016) SPOCK1 promotes the proliferation, migration and invasion of glioma cells through PI3K/AKT and Wnt/beta-catenin signaling pathways. Oncol Rep 35(6):3566–3576. doi:10.3892/or.2016.4757
Lu J, Jeong HW, Kong N, Yang Y, Carroll J, Luo HR, Silberstein LE, Yupoma, Chai L (2009) Stem cell factor SALL4 represses the transcriptions of PTEN and SALL1 through an epigenetic repressor complex. PloS one 4(5):e5577. doi:10.1371/journal.pone.0005577
Li XT, Wang HZ, Wu ZW, Yang TQ, Zhao ZH, Chen GL, Xie XS, Li B, Wei YX, Huang YL, Zhou YX, Du ZW (2015) miR-494-3p Regulates cellular proliferation, invasion, migration, and apoptosis by PTEN/AKT signaling in human glioblastoma cells. Cell Mol Neurobiol 35(5):679–687. doi:10.1007/s10571-015-0163-0
Kinoshita S, Sho M, Enomoto Y, Koyama F, Akahori T, Nagai M, Ito M, Morita K, Ohbayashi C, Nakajima Y (2014) SALL4 positive fetal gut-like adenocarcinoma of the duodenum. Pathol Int 64(11):581–584. doi:10.1111/pin.12211
Wang F, Guo Y, Chen Q, Yang Z, Ning N, Zhang Y, Xu Y, Xu X, Tong C, Chai L, Cui W (2013) Stem cell factor SALL4, a potential prognostic marker for myelodysplastic syndromes. J Hematol Oncol 6(1):73. doi:10.1186/1756-8722-6-73
Wang F, Zhao W, Kong N, Cui W, Chai L (2014) The next new target in leukemia: the embryonic stem cell gene. Mol Cell Oncol 1(4):e969169. doi:10.4161/23723548.2014.969169
Ma Y, Cui W, Yang J, Qu J, Di C, Amin HM, Lai R, Ritz J, Krause DS, Chai L (2006) SALL4, a novel oncogene, is constitutively expressed in human acute myeloid leukemia (AML) and induces AML in transgenic mice. Blood 108(8):2726–2735. doi:10.1182/blood-2006-02-001594
Li A, Jiao Y, Yong KJ, Wang F, Gao C, Yan B, Srivastava S, Lim GS, Tang P, Yang H, Tenen DG, Chai L (2015) SALL4 is a new target in endometrial cancer. Oncogene 34(1):63–72. doi:10.1038/onc.2013.529
Zhang L, Xu Z, Xu X, Zhang B, Wu H, Wang M, Zhang X, Yang T, Cai J, Yan Y, Mao F, Zhu W, Shao Q, Qian H, Xu W (2014) SALL4, a novel marker for human gastric carcinogenesis and metastasis. Oncogene 33(48):5491–5500. doi:10.1038/onc.2013.495
Gao C, Kong NR, Chai L (2011) The role of stem cell factor SALL4 in leukemogenesis. Crit Rev Oncog 16(1–2):117–127
Xiong J, Todorova D, Su NY, Kim J, Lee PJ, Shen Z, Briggs SP, Xu Y (2015) Stemness factor Sall4 is required for DNA damage response in embryonic stem cells. J Cell Biol 208(5):513–520. doi:10.1083/jcb.201408106
Liao W, Aguila JR, Yao Y, Yang J, Zieve G, Jiang Y, Avila C, Senzel L, Lai R, Xu D, Dai W, Ma Y (2013) Enhancing bone marrow regeneration by SALL4 protein. J Hematol Oncol 6:84. doi:10.1186/1756-8722-6-84
Islam F, Gopalan V, Wahab R, Smith RA, Lam AK (2015) Cancer stem cells in oesophageal squamous cell carcinoma: identification, prognostic and treatment perspectives. Critical Rev Oncol/Hematol 96 (1):9–19. doi:10.1016/j.critrevonc.2015.04.007
Park H, Lee H, Seo AN, Cho JY, Choi YR, Yoon Y-S, Han H-S, Park YN, Kim H (2015) SALL4 expression in hepatocellular carcinomas is associated with EpCAM-positivity and a poor prognosis. J Pathol Transl Med 49(5):373–381. doi:10.4132/jptm.2015.07.09
Gao C, Kong NR, Li A, Tatetu H, Ueno S, Yang Y, He J, Yang J, Ma Y, Kao GS, Tenen DG, Chai L (2013) SALL4 is a key transcription regulator in normal human hematopoiesis. Transfusion 53(5):1037–1049. doi:10.1111/j.1537-2995.2012.03888.x
Bohm J, Sustmann C, Wilhelm C, Kohlhase J (2006) SALL4 is directly activated by TCF/LEF in the canonical Wnt signaling pathway. Biochem Biophys Res Commun 348(3):898–907. doi:10.1016/j.bbrc.2006.07.124
Zhang X, Song Q, Wei C, Qu J (2015) LRIG1 inhibits hypoxia-induced vasculogenic mimicry formation via suppression of the EGFR/PI3K/AKT pathway and epithelial-to-mesenchymal transition in human glioma SHG-44 cells. Cell Stress Chaperones 20 (4):631–641. doi:10.1007/s12192-015-0587-y
Thapa N, Choi S, Tan X, Wise T, Anderson RA (2015) Phosphatidylinositol phosphate 5-kinase igamma and phosphoinositide 3-kinase/Akt signaling couple to promote oncogenic growth. J Biol Chem 290(30):18843–18854. doi:10.1074/jbc.M114.596742
Ke Z, Caiping S, Qing Z, Xiaojing W (2015) Sonic hedgehog-Gli1 signals promote epithelial-mesenchymal transition in ovarian cancer by mediating PI3K/AKT pathway. Medical oncology 32(1):368. doi:10.1007/s12032-014-0368-y
Luo Q, Wang C, Jin G, Gu D, Wang N, Song J, Jin H, Hu F, Zhang Y, Ge T, Huo X, Chu W, Shu H, Fang J, Yao M, Gu J, Cong W, Qin W (2015) LIFR functions as a metastasis suppressor in hepatocellular carcinoma by negatively regulating phosphoinositide 3-kinase/AKT pathway. Carcinogenesis 36(10):1201–1212. doi:10.1093/carcin/bgv108
Villa-Moruzzi E (2013) PTPN12 controls PTEN and the AKT signalling to FAK and HER2 in migrating ovarian cancer cells. Mol Cell Biochem 375(1–2):151–157. doi:10.1007/s11010-012-1537-y
Fan Y, Yang F, Cao X, Chen C, Zhang X, Zhang X, Lin W, Wang X, Liang C (2016) Gab1 regulates SDF-1-induced progression via inhibition of apoptosis pathway induced by PI3K/AKT/Bcl-2/BAX pathway in human chondrosarcoma. Tumour Biol 37(1):1141–1149. doi:10.1007/s13277-015-3815-2
Liu W, Ren H, Ren J, Yin T, Hu B, Xie S, Dai Y, Wu W, Xiao Z, Yang X, Xie D (2013) The role of EGFR/PI3K/Akt/cyclinD1 signaling pathway in acquired middle ear cholesteatoma. Mediators Inflamm 2013(5):651207. doi:10.1155/2013/651207
Segrelles C, Ruiz S, Perez P, Murga C, Santos M, Budunova IV, Martinez J, Larcher F, Slaga TJ, Gutkind JS, Jorcano JL, Paramio JM (2002) Functional roles of Akt signaling in mouse skin tumorigenesis. Oncogene 21(1):53–64. doi:10.1038/sj.onc.1205032
Alam S, Pal A, Kumar R, Dwivedi PD, Das M, Ansari KM (2014) EGFR-mediated Akt and MAPKs signal pathways play a crucial role in patulin-induced cell proliferation in primary murine keratinocytes via modulation of cyclin D1 and COX-2 expression. Mol Carcinog 53(12):988–998. doi:10.1002/mc.22060
Bohm J, Kaiser FJ, Borozdin W, Depping R, Kohlhase J (2007) Synergistic cooperation of Sall4 and cyclin D1 in transcriptional repression. Biochem Biophys Res Commun 356(3):773–779. doi:10.1016/j.bbrc.2007.03.050
Deng G, Zhu L, Huang F, Nie W, Huang W, Xu H, Zheng S, Yi Z, Wan T (2015) SALL4 is a novel therapeutic target in intrahepatic cholangiocarcinoma. Oncotarget 6(29):27416–27426. doi:10.18632/oncotarget.4862
Miettinen M, Wang Z, McCue PA, Sarlomo-Rikala M, Rys J, Biernat W, Lasota J, Lee Y-S (2014) SALL4 expression in germ cell and non–germ cell tumors. Am J Surg Pathol 38(3):410–420. doi:10.1097/pas.0000000000000116
Acknowledgements
We thank Genechem for providing us the interference sequence and technical assistance. We thank our central lab for providing technical instruction and assistance. We thank Cell Bank Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China) for offering us glioma cells line. We thank The First Affiliated Hospital of Soochow University for providing us with glioma samples.
Funding
This work was partly supported by National Science Foundation of China (Grant Nos. 81372689, 81572475), Anti-Cancer Association Foundation of China (Grant CSNO-2013-MSD-009) and the foundation of Suzhou science and technology development plan item (Grant No. SYS201324).
Author information
Authors and Affiliations
Contributions
CL and HW conducted most of the experiments, analyzed the results, and wrote most of the paper. CS conducted the selection of topics, coordinated the study and wrote the paper with YZ. LS conducted the design of experiments program, analyzed the results. RY and HY conducted experiments on cell adhesion and spreading and the collection of glioma specimen. YL offered technical instruction and assistance with TS. YZ and ZD participated in the proofread of the paper. All the authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liu, C., Wu, H., Li, Y. et al. SALL4 suppresses PTEN expression to promote glioma cell proliferation via PI3K/AKT signaling pathway. J Neurooncol 135, 263–272 (2017). https://doi.org/10.1007/s11060-017-2589-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2589-3