Abstract
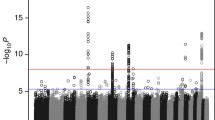
Although genome-wide association studies have identified several susceptibility loci for adult glioma, little is known regarding the potential contribution of genetic variation in the human leukocyte antigen (HLA) region to glioma risk. HLA associations have been reported for various malignancies, with many studies investigating selected candidate HLA polymorphisms. However, no systematic analysis has been conducted in glioma patients, and no investigation into potential non-additive effects has been described. We conducted comprehensive genetic analyses of HLA variants among 1746 adult glioma patients and 2312 controls of European-ancestry from the GliomaScan Consortium. Genotype data were generated with the Illumina 660-Quad array, and we imputed HLA alleles using a reference panel of 5225 individuals in the Type 1 Diabetes Genetics Consortium who underwent high-resolution HLA typing via next-generation sequencing. Case-control comparisons were adjusted for population stratification using ancestry-informative principal components. Because alleles in different loci across the HLA region are linked, we created multigene haplotypes consisting of the genes DRB1, DQA1, and DQB1. Although none of the haplotypes were associated with glioma in additive models, inclusion of a dominance term significantly improved the model for multigene haplotype HLA-DRB1*1501-DQA1*0102-DQB1*0602 (P = 0.002). Heterozygous carriers of the haplotype had an increased risk of glioma [odds ratio (OR) 1.23; 95% confidence interval (CI) 1.01–1.49], while homozygous carriers were at decreased risk compared with non-carriers (OR 0.64; 95% CI 0.40–1.01). Our results suggest that the DRB1*1501-DQA1*0102-DQB1*0602 haplotype may contribute to the risk of glioma in a non-additive manner, with the positive dominance effect partly explained by an epistatic interaction with HLA-DRB1*0401-DQA1*0301-DQB1*0301.

Similar content being viewed by others
References
Ostrom QT, Gittleman H, Farah P et al (2013) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2006–2010. Neuro-Oncology 15:ii1–ii56. doi:10.1093/neuonc/not151
Ostrom QT, Bauchet L, Davis FG et al (2014) The epidemiology of glioma in adults: a state of the science review. Neuro-Oncology 16:896–913. doi:10.1093/neuonc/nou087
Wiemels JL, Wilson D, Patel C et al (2009) IgE, allergy, and risk of glioma: update from the san francisco bay area adult glioma study in the Temozolomide Era. Int J Cancer J Int Cancer 125:680–687. doi:10.1002/ijc.24369
Amirian ES, Zhou R, Wrensch MR et al (2016) Approaching a scientific consensus on the association between allergies and glioma risk: a report from the glioma international case-control study. Cancer Epidemiol Biomark 25:282–290. doi:10.1158/1055-9965.EPI-15-0847
Egeberg A, Hansen PR, Gislason GH, Thyssen JP (2016) Association of rosacea with risk for glioma in a Danish Nationwide Cohort Study. JAMA Dermatol 152:541–545. doi:10.1001/jamadermatol.2015.5549
Brenner AV, Linet MS, Fine HA et al (2002) History of allergies and autoimmune diseases and risk of brain tumors in adults. Int J Cancer 99:252–259. doi:10.1002/ijc.10320
Hinds DA, McMahon G, Kiefer AK et al (2013) A genome-wide association meta-analysis of self-reported allergy identifies shared and allergy-specific susceptibility loci. Nat Genet 45:907–911. doi:10.1038/ng.2686
Chang ALS, Raber I, Xu J et al (2015) Assessment of the genetic basis of rosacea by genome-wide association study. J Invest Dermatol 135:1548–1555. doi:10.1038/jid.2015.53
Trowsdale J, Knight JC (2013) Major histocompatibility complex genomics and human disease. Annu Rev Genomics Hum Genet 14:301–323. doi:10.1146/annurev-genom-091212-153455
Bateman AC, Howell WM (1999) Human leukocyte antigens and cancer: is it in our genes? J Pathol 188:231–236. doi:10.1002/(SICI)1096-9896(199907)188:3
Walsh KM, Codd V, Smirnov IV et al (2014) Variants near TERT and TERC influencing telomere length are associated with high-grade glioma risk. Nat Genet 46:731–735
Melin BS, Barnholtz-Sloan JS, Wrensch MR et al (2017) Genome-wide association study of glioma subtypes identifies specific differences in genetic susceptibility to glioblastoma and non-glioblastoma tumors. Nat Genet 49:789–794
Rajaraman P, Melin BS, Wang Z et al (2012) Genome-wide association study of glioma and meta-analysis. Hum Genet 131:1877–1888. doi:10.1007/s00439-012-1212-0
Lenz TL, Deutsch AJ, Han B et al (2015) Widespread non-additive and interaction effects within HLA loci modulate the risk of autoimmune diseases. Nat Genet 47:1085–1090. doi:10.1038/ng.3379
Goyette P, Boucher G, Mallon D et al (2015) High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat Genet 47:172–179. doi:10.1038/ng.3176
The International Multiple Sclerosis Genetics Consortium (2015) Class II HLA interactions modulate genetic risk for multiple sclerosis. Nat Genet 47:1107–1113
Guerini FR, Agliardi C, Zanzottera M et al (2005) Human leukocyte antigen distribution analysis in North Italian brain glioma patients:an association with HLA-DRB1*14. J Neurooncol 77:213–217. doi:10.1007/s11060-005-9032-x
Machulla HKG, Steinborn F, Schaaf A et al (2001) Brain glioma and human leukocyte antigens (HLA)—is there an association. J Neurooncol 52:253–261. doi:10.1023/A:1010612327647
La Torre D, Maugeri R, Angileri FF et al (2009) Human leukocyte antigen frequency in human high-grade gliomas: a case-control study in Sicily. Neurosurgery 64:1082–1088; Discussion 1088–1089. doi:10.1227/01.NEU.0000345946.35786.92
Tang J, Shao W, Dorak MT et al (2005) Positive and negative associations of human leukocyte antigen variants with the onset and prognosis of adult glioblastoma multiforme. Cancer Epidemiol Biomark Amp Prev 14:2040. doi:10.1158/1055-9965.EPI-05-0136
Song W, Ruder AM, Hu L et al (2009) Genetic epidemiology of glioblastoma multiforme: confirmatory and new findings from analyses of human leukocyte antigen alleles and motifs. PLoS ONE 4:e7157. doi:10.1371/journal.pone.0007157
Bassig BA, Inskip PD, Burdette L et al (2011) Selected human leukocyte antigen class II polymorphisms and risk of adult glioma. J Neuroimmunol 233:185–191. doi:10.1016/j.jneuroim.2010.11.005
Guja C, Guja L, Nutland S et al (2004) Type 1 diabetes genetic susceptibility encoded by HLA DQB1 genes in Romania. J Cell Mol Med 8:249–256
Laaksonen M, Pastinen T, Sjoroos M et al (2002) HLA class II associated risk and protection against multiple sclerosis-a Finnish family study. J Neuroimmunol 122:140–145
Jia X, Han B, Onengut-Gumuscu S et al (2013) Imputing amino acid polymorphisms in human leukocyte antigens. PLoS ONE 8:e64683. doi:10.1371/journal.pone.0064683
Rich SS, Concannon P, Erlich H et al (2006) The type 1 diabetes genetics consortium. Ann N Y Acad Sci 1079:1–8. doi:10.1196/annals.1375.001
Chang CC, Chow CC, Tellier LC et al (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4:7. doi:10.1186/s13742-015-0047-8
Price AL, Patterson NJ, Plenge RM et al (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909. doi:10.1038/ng1847
Whitacre CC (2001) Sex differences in autoimmune disease. Nat Immunol 2:777–780. doi:10.1038/ni0901-777
Jenkins RB, Xiao Y, Sicotte H et al (2012) A low-frequency variant at 8q24.21 is strongly associated with risk of oligodendroglial tumors and astrocytomas with IDH1 or IDH2 mutation. Nat Genet 44:1122–1125. doi:10.1038/ng.2388
Grier JT, Batchelor T (2006) Low-grade gliomas in adults. Oncologist 11:681–693. doi:10.1634/theoncologist.11-6-681
Eckel-Passow JE, Lachance DH, Molinaro AM et al (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. New Engl J Med 372:2499–2508. doi:10.1056/NEJMoa1407279
Traherne JA (2008) Human MHC architecture and evolution: implications for disease association studies. Int J Immunogenet 35:179–192. doi:10.1111/j.1744-313X.2008.00765.x
Mignot E, Lin L, Rogers W et al (2001) Complex HLA-DR and -DQ interactions confer risk of narcolepsy-cataplexy in three ethnic groups. Am J Hum Genet 68:686–699
Fernando MMA, Stevens CR, Sabeti PC et al (2007) Identification of two independent risk factors for lupus within the MHC in United Kingdom Families. PLoS Genet 3:e192. doi:10.1371/journal.pgen.0030192
Erlich H, Valdes AM, Noble J et al (2008) HLA DR-dq haplotypes and genotypes and type 1 diabetes risk. Diabetes 57:1084. doi:10.2337/db07-1331
Schmidt H, Williamson D, Ashley-Koch A (2007) HLA-DR15 haplotype and multiple sclerosis: a HuGE review. Am J Epidemiol 165:1097–1109. doi:10.1093/aje/kwk118
Hildesheim A, Schiffman M, Scott DR et al (1998) Human leukocyte antigen class I/II alleles and development of human papillomavirus-related cervical neoplasia: results from a case-control study conducted in the United States. Cancer Epidemiol Biomark Prev Publ Am Assoc Cancer Res Cosponsored Am Soc Prev Oncol 7:1035–1041
Dziurzynski K, Chang SM, Heimberger AB et al (2012) Consensus on the role of human cytomegalovirus in glioblastoma. Neuro-Oncol 14:246–255. doi:10.1093/neuonc/nor227
Amirian ES, Scheurer ME, Zhou R et al (2016) History of chickenpox in glioma risk: a report from the glioma international case–control study (GICC). Cancer Med 5:1352–1358. doi:10.1002/cam4.682
Rose AM, Bell LCK (2012) Epistasis and immunity: the role of genetic interactions in autoimmune diseases. Immunology 137:131–138. doi:10.1111/j.1365-2567.2012.03623.x
Wiemels JL, Wiencke JK, Sison JD et al (2002) History of allergies among adults with glioma and controls. Int J Cancer 98:609–615. doi:10.1002/ijc.10239
Acknowledgements
The results published here are, in part, based upon data obtained from dbGaP Study Accession phs000652.v1.p1: “Cohort-based Genome-Wide Association Study of Glioma (GliomaScan)” which was supported by intramural funds from the NCI and federal funds from the NCI under Contract N01-CO-12400. The authors additionally acknowledge use of the British 1958 Birth Cohort DNA collection, funded by the Medical Research Council Grant G0000934 and the Wellcome Trust Grant 068545/Z/02. This research uses resources provided by the Type 1 Diabetes Genetics Consortium (T1DGC); a collaborative clinical study sponsored by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK); National Institute of Allergy and Infectious Diseases (NIAID); National Human Genome Research Institute (NHGRI); National Institute of Child Health and Human Development; Juvenile Diabetes Research Foundation International (JDRF), supported by U01 DK062418.
Funding
This work was supported by the National Institutes of Health T32CA151022-06 (C.Z.), R25T CA112355 (J.S.W.), and The Sontag Foundation (K.M.W.)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, C., de Smith, A.J., Smirnov, I.V. et al. Non-additive and epistatic effects of HLA polymorphisms contributing to risk of adult glioma. J Neurooncol 135, 237–244 (2017). https://doi.org/10.1007/s11060-017-2569-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2569-7