Abstract
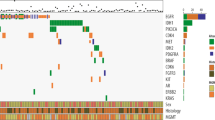
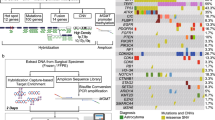
Gliomas consist of multiple histologic and molecular subtypes with different clinical phenotypes and responsiveness to treatment. However, enrollment criteria for clinical trials still largely do not take into account these underlying molecular differences. We have incorporated a high-throughput tumor genotyping program based on the ABI SNaPshot platform as well as other molecular diagnostic tests into the standard evaluation of glioma patients in order to assess whether prospective molecular profiling would allow rational patient selection onto clinical trials. From 218 gliomas we prospectively collected SNaPshot genotyping data on 68 mutated loci from 15 key cancer genes along with data from clinical assays for gene amplification (EGFR, PDGFRA, MET), 1p/19q co-deletion and MGMT promoter methylation. SNaPshot mutations and focal gene amplifications were detected in 38.5 and 47.1 % of glioblastomas, respectively. Genetic alterations in EGFR, IDH1 and PIK3CA closely matched frequencies reported in recent studies. In addition, we identified events that are rare in gliomas although are known driver mutations in other cancer types, such as mutations of AKT1, BRAF and KRAS. Patients with genetic alterations that activate signaling pathways were enrolled onto genetically selective clinical trials for malignant glioma as well as for other solid cancers. High-throughput molecular profiling incorporated into the routine clinical evaluation of glioma patients may enable the rational selection of patients for targeted therapy clinical trials and thereby improve the likelihood that such trials succeed.


Similar content being viewed by others
References
Druker B, Talpaz M, Resta D, Peng B, Buchdunger E, Ford J, Lydon N, Kantarjian H, Capdeville R, Ohno-Jones S, Sawyers C (2001) Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N Engl J Med 344:1031–1037
Lynch T, Bell D, Sordella R, Gurubhagavatula S, Okimoto R, Brannigan B, Harris P, Haserlat S, Supko J, Haluska F, Louis D, Christiani D, Settleman J, Haber D (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350:2129–2139
Bollag G, Hirth P, Tsai J, Zhang J, Ibrahim PN, Cho H, Spevak W, Zhang C, Zhang Y, Habets G, Burton EA, Wong B, Tsang G, West BL, Powell B, Shellooe R, Marimuthu A, Nguyen H, Zhang KY, Artis DR, Schlessinger J, Su F, Higgins B, Iyer R, D’Andrea K, Koehler A, Stumm M, Lin PS, Lee RJ, Grippo J, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA, Chapman PB, Flaherty KT, Xu X, Nathanson KL, Nolop K (2010) Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature 467:596–599
Kwak EL, Bang YJ, Camidge DR, Shaw AT, Solomon B, Maki RG, Ou SH, Dezube BJ, Janne PA, Costa DB, Varella-Garcia M, Kim WH, Lynch TJ, Fidias P, Stubbs H, Engelman JA, Sequist LV, Tan W, Gandhi L, Mino-Kenudson M, Wei GC, Shreeve SM, Ratain MJ, Settleman J, Christensen JG, Haber DA, Wilner K, Salgia R, Shapiro GI, Clark JW, Iafrate AJ (2010) Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med 363:1693–1703
Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA, O’Dwyer PJ, Lee RJ, Grippo JF, Nolop K, Chapman PB (2010) Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med 363:809–819
Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A, Fleming T, Eiermann W, Wolter J, Pegram M, Baselga J, Norton L (2001) Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med 344:783–792
Paez J, Janne P, Lee J, Tracy S, Greulich H, Gabriel S, Herman P, Kaye F, Lindeman N, Boggon T, Naoki K, Sasaki H, Fujii Y, Eck M, Sellers W, Johnson B, Meyerson M (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304:1497–1500
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522
Parsons D, Jones S, Zhang X, Lin J, Leary R, Angenendt P, Mankoo P, Carter H, Siu I, Gallia G, Olivi A, McLendon R, Rasheed B, Keir S, Nikolskaya T, Nikolsky Y, Busam D, Tekleab H, Diaz LJ, Hartigan J, Smith D, Strausberg R, Marie S, Shinjo S, Yan H, Riggins G, Bigner D, Karchin R, Papadopoulos N, Parmigiani G, Vogelstein B, Velculescu V, Kinzler K (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812
Hegi M, Diserens A, Gorlia T, Hamou M, de Tribolet N, Weller M, Kros J, Hainfellner J, Mason W, Mariani L, Bromberg J, Hau P, Mirimanoff R, Cairncross J, Janzer R, Stupp R (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352:997–1003
Yan H, Parsons D, Jin G, McLendon R, Rasheed B, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins G, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler K, Velculescu V, Vogelstein B, Bigner D (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J, Ducray F, Hallani SE, Boisselier B, Mokhtari K, Hoang-Xuan K, Delattre JY (2009) Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 27:4150–4154
Jansen M, Yip S, Louis DN (2010) Molecular pathology in adult gliomas: diagnostic, prognostic, and predictive markers. Lancet Neurol 9:717–726
Dias-Santagata D, Akhavanfard S, David SS, Vernovsky K, Kuhlmann G, Boisvert SL, Stubbs H, McDermott U, Settleman J, Kwak EL, Clark JW, Isakoff SJ, Sequist LV, Engelman JA, Lynch TJ, Haber DA, Louis DN, Ellisen LW, Borger DR, Iafrate AJ (2010) Rapid targeted mutational analysis of human tumours: a clinical platform to guide personalized cancer medicine. EMBO Mol Med 2:146–158
Stupp R, Mason W, van den Bent M, Weller M, Fisher B, Taphoorn M, Belanger K, Brandes A, Marosi C, Bogdahn U, Curschmann J, Janzer R, Ludwin S, Gorlia T, Allgeier A, Lacombe D, Cairncross J, Eisenhauer E, Mirimanoff R (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996
Palmisano W, Divine K, Saccomanno G, Gilliland F, Baylin S, Herman J, Belinsky S (2000) Predicting lung cancer by detecting aberrant promoter methylation in sputum. Cancer Res 60:5954–5958
Cancer Genome Atlas Research Network (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455:1061–1068
Chi A, Batchelor T, Kwak E et al (2011) Rapid radiographic and clinical improvement after treatment of a MET-amplified recurrent glioblastoma with a MET inhibitor (crizotinib). J Clin Oncol 29(15_suppl):2072
Humphrey P, Wong A, Vogelstein B, Friedman H, Werner M, Bigner D, Bigner S (1988) Amplification and expression of the epidermal growth factor receptor gene in human glioma xenografts. Cancer Res 48:2231–2238
Mizoguchi M, Betensky R, Batchelor T, Bernay D, Louis D, Nutt C (2006) Activation of STAT3, MAPK, and AKT in malignant astrocytic gliomas: correlation with EGFR status, tumor grade, and survival. J Neuropathol Exp Neurol 65:1181–1188
Chaffanet M, Chauvin C, Laine M, Berger F, Chedin M, Rost N, Nissou M, Benabid A (1992) EGF receptor amplification and expression in human brain tumours. Eur J Cancer 28:11–17
Gallia GL, Rand V, Siu IM, Eberhart CG, James CD, Marie SK, Oba-Shinjo SM, Carlotti CG, Caballero OL, Simpson AJ, Brock MV, Massion PP, Carson BS Sr, Riggins GJ (2006) PIK3CA gene mutations in pediatric and adult glioblastoma multiforme. Mol Cancer Res 4:709–714
Knobbe CB, Reifenberger J, Reifenberger G (2004) Mutation analysis of the Ras pathway genes NRAS, HRAS, KRAS and BRAF in glioblastomas. Acta Neuropathol 108:467–470
Jeuken J, van den Broecke C, Gijsen S, Boots-Sprenger S, Wesseling P (2007) RAS/RAF pathway activation in gliomas: the result of copy number gains rather than activating mutations. Acta Neuropathol 114:121–133
Basto D, Trovisco V, Lopes JM, Martins A, Pardal F, Soares P, Reis RM (2005) Mutation analysis of B-RAF gene in human gliomas. Acta Neuropathol 109:207–210
Yip S, Miao J, Cahill DP et al (2009) MSH6 mutations arise in glioblastomas during temozolomide therapy and mediate temozolomide resistance. Clin Cancer Res 15(14):4622–4629
Snuderl M, Fazlollahi L, Le LP et al (2011) Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 20(6):810–817
Engelman JA, Chen L, Tan X, Crosby K, Guimaraes AR, Upadhyay R, Maira M, McNamara K, Perera SA, Song Y, Chirieac LR, Kaur R, Lightbown A, Simendinger J, Li T, Padera RF, Garcia-Echeverria C, Weissleder R, Mahmood U, Cantley LC, Wong KK (2008) Effective use of PI3K and MEK inhibitors to treat mutant Kras G12D and PIK3CA H1047R murine lung cancers. Nat Med 14:1351–1356
Bamford S, Dawson E, Forbes S, Clements J, Pettett R, Dogan A, Flanagan A, Teague J, Futreal PA, Stratton MR, Wooster R (2004) The COSMIC (catalogue of somatic mutations in cancer) database and website. Br J Cancer 91:355–358
Peraud A, Watanabe K, Schwechheimer K, Yonekawa Y, Kleihues P, Ohgaki H (1999) Genetic profile of the giant cell glioblastoma. Lab Invest 79:123–129
Peraud A, Watanabe K, Plate KH, Yonekawa Y, Kleihues P, Ohgaki H (1997) p53 Mutations versus EGF receptor expression in giant cell glioblastomas. J Neuropathol Exp Neurol 56:1236–1241
Meyer-Puttlitz B, Hayashi Y, Waha A, Rollbrocker B, Bostrom J, Wiestler OD, Louis DN, Reifenberger G, von Deimling A (1997) Molecular genetic analysis of giant cell glioblastomas. Am J Pathol 151:853–857
Duerr EM, Rollbrocker B, Hayashi Y, Peters N, Meyer-Puttlitz B, Louis DN, Schramm J, Wiestler OD, Parsons R, Eng C, von Deimling A (1998) PTEN mutations in gliomas and glioneuronal tumors. Oncogene 16:2259–2264
Wang SI, Puc J, Li J, Bruce JN, Cairns P, Sidransky D, Parsons R (1997) Somatic mutations of PTEN in glioblastoma multiforme. Cancer Res 57:4183–4186
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, Puc J, Miliaresis C, Rodgers L, McCombie R, Bigner SH, Giovanella BC, Ittmann M, Tycko B, Hibshoosh H, Wigler MH, Parsons R (1997) PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275:1943–1947
Acknowledgments
Preliminary data of this manuscript was presented at the 2011 American Association of Neurology Annual Meeting in Honolulu, Hawaii. Andrew S. Chi is supported by a Joan Ambriz American Brain Tumor Association Basic Research Fellowship and an Early Career research award from the Ben and Catherine Ivy Foundation. Ethical Standards: All results were obtained from the medical record of patients and collected in an IRB-approved patient database. Molecular testing was performed as part of routine medical care in CLIA-certified clinical laboratories. All molecular tests comply with the current laws of the USA.
Conflicts of interest
The authors have no conflicts of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chi, A.S., Batchelor, T.T., Dias-Santagata, D. et al. Prospective, high-throughput molecular profiling of human gliomas. J Neurooncol 110, 89–98 (2012). https://doi.org/10.1007/s11060-012-0938-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-012-0938-9