Abstract
The interaction between metal nanoparticles and bacteria belongs to the central issues in a dynamically growing bionanotechnological research. Herein, we investigated the adhesion efficiency of gold nanoparticles (30 nm) for various bacterial strains, both Gram-positive (Bacillus subtilis, Staphylococcus carnosus) and Gram-negative (Neisseria subflava, Stenotrophomonas maltophilia). The thorough microscopic (SEM/TEM) observations revealed that the nanoparticles do not penetrate into the bacterial cells but adhere to the walls. Large differences in the adhered nanoparticles amount were observed for the investigated strains (B. subtilis >> S. carnosus > N. subflava > S. maltophilia). A direct correlation between the number of the attached nanoparticles and the ζ-potential of the bacterial strains was found, and the results were rationalized in terms of the DLVO model. The calculated DLVO energy profiles revealed that the activation barriers for the adhesion process are rather small (1.45–1.55 kT), and the primary energy minima of 120–170 kT are favorable for the effective adsorption process. The established linear correlation between the nanoparticles adhered to the cell surface and the size of the critical volume around the bacterial cell, where the attraction forces dominate, implies that the observed dramatic differences in the attachment efficiency result from the availability of the nanoparticles in the critical volume of the surrounding suspensions. Owing to non-specific interactions governed by the ζ-potential mainly, the obtained results can be readily extended for the other bacteria–nanoparticle systems, providing a rational background for future advances in bacteria detection and thorough characterization via SERS method as well as for nanoparticles assemblies towards nanoelectronics.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the wide range of microorganisms, bacteria are extensively investigated for many decades in the context of their surface colonization ability, leading to the formation of biofilms and the associated hazard of infection (Donlan and Costerton 2002). Recently, there is a growing research interest in the interaction between bacteria and nanoparticles of various chemical nature (metallic, oxides, sulfates, etc.) (Barnes et al. 2013; von Moos et al. 2014; Calatayud et al. 2014). Such investigations have a strong impact in many areas of nanoscience, including toxicology, pharmacology, nanomedicine, biotechnology, environmental sciences, chemistry, and even geology (Rai et al. 2009; Nath and Banerjee 2013; Judy and Bertsch 2014; Wadhwani et al. 2016). Bacteria are unicellular prokaryotic organisms surrounded by the cell membrane, cell wall, and cell envelope in some cases (Silhavy et al. 2010). In general, there are three main morphological types of bacterial cells that can be distinguished: spheres (cocci), rods (bacilli), and spirals (spirilla) (Young 2006). Depending on the composition of their cell walls, bacteria are divided into two major categories: Gram-negative (G−) and Gram-positive). In Fig. 1, simplified differences in the cell wall structure of those two groups are shown.
A major component of the bacterial cell walls, in both G+ and G− bacteria, is the polymer peptidoglycan. The G+ bacteria have thick cell walls, which consist of a large multilayer region of peptidoglycan with a wall teichoic acid (WTA) and lipoteichoic acid (LTA), attached to the peptidoglycan layer and to the cell membrane, respectively. The G− bacteria have a thin layer of the peptidoglycan, situated between the cell and outer membranes, in the region called periplasmic space (the specific feature of G−), filled with gel-like periplasm. Instead of teichoic acids, the G− bacteria synthesize lipopolysaccharides (LPS), which form distinct domains (marked in green) within the outer phospholipids bilayer (Fig. 1) (Malanovic and Lohner 2016). Bacterial cell wall plays a crucial role in the cell protection against the negative impact of physical, chemical, and biological factors, controlling the characteristic shape of the cells. The additional vital function of the cell walls associated with the adhesion to any surface owing to the presence of adhesins, i.e., specific macromolecules responsible for adhesive interactions at the interface (Moriarty et al. 2011; Braslavsky et al. 2016).
In the adhesion process, the molecular properties of the bacterial cell surface are of crucial importance. Besides hydrophobicity, the surface charge defines the prime variable typically used for the evaluation of the bacterial adhesion process (Spriano et al. 2017). Usually, bacteria exhibit an overall negative charge when measured in water suspension at physiological pH. In the case of the G+ bacteria, the phosphoryl groups located in the LTA and WTA tails play an important role in the generation of the net negative charge of the cells. For the G− bacteria, the negative charge results from the ionization of the phosphoryl and 2-keto-3-deoxyoctonate carboxylate groups present in the LPS chains (Wilson et al. 2001). However, the surface charge measured as a ζ-potential (zeta potential) depends on the bacterial strain and its particular surface characteristics (Kłodzińska et al. 2010; Khelissa et al. 2017). The electrochemical behavior of bacterial cell surfaces is particularly important for the application of nanoparticles (NPs) as antimicrobial agents (Ramasamy et al. 2016; Mmola et al. 2016), in the detection of viruses, bacteria, and spores (Sajjanar et al. 2015; Donmez et al. 2017) or even for the production of NPs inside the microorganism (Nangia et al. 2009; Durán et al. 2011; Deplanche et al. 2012; Srivastava et al. 2013; Jung et al. 2018).
Following the rough definition (The European Commission 2011), objects are defined as NPs when their dimension is smaller than 100 nm. Owing to their high stability and spectacular catalytic and plasmonic behavior, one of the best characterized and intensively investigated are nanoparticles of gold (AuNPs) (Carrière 2012). Because of their biocompatibility, AuNPs are also widely investigated in medicine, including cancer therapies, pharmacology, and advanced diagnostics (Dykman and Khlebtsov 2011; Saha et al. 2012; Gatea et al. 2015; Sajjanar et al. 2015; Bozorgmehr et al. 2016). Bare AuNPs generally exhibit weak antimicrobial properties (Zhang et al. 2015), but ones functionalized with antibiotics or other ligands they can be successfully used as targeting drug delivery systems (Pradeepa et al. 2016; Faisal et al. 2018), antimicrobial agents (Bresee et al. 2011; Li et al. 2014; Ramasamy et al. 2014), or for photothermal therapy and tissue imaging (Loeb et al. 2018; Yang et al. 2018; Zhu et al. 2018).
However, the growing applications of NPs unavoidably lead to their systematic accumulation in the environment. Although some studies suggest weak toxicity of AuNPs (Ramasamy et al. 2016; Faisal et al. 2018), the majority of scientific reports reveal negative impacts on aquatic organisms, mammals, and humans (Bar-Ilan et al. 2009; Bozich et al. 2014; von Moos et al. 2014; Aruoja et al. 2015; McCracken et al. 2016). Therefore, there is an urgent need to identify and quantify the principal parameters governing the NPs interaction with the biological surfaces such as bacterial cell walls (Xing et al. 2018). Most of the previous studies are focused on the interaction between NPs and bacteria, leading to an apoptotic disintegration of the bacterial cell walls (cells death) (Chwalibog et al. 2010; Miller et al. 2016; López-Lorente et al. 2019). Yet, we believe that such knowledge may be used for designing microorganism-based effective systems, not only for the mere capture of NPs from the environment but also for the development of new functionalities associated with a controlled dispersion of NPs of various size, shape, and chemical nature (Polish Patents Applications P.426130 from 29 June 2018 and P.424760 from 5 March 2018).
The aim of this study was to substantiate the interaction of the G+ and G− bacteria with the gold nanoparticles and to identify the key descriptors responsible for the nanoparticles attachment efficiency. The results are accounted for in terms of the DLVO (Derjaguin–Landau–Verwey–Overbeek) model.
Materials and methods
Bacterial strains
The reference strains used in the study were Staphylococcus carnosus DSM 20501 (Deutsche Sammlung von Mikroorganismen und Zellkulturen), Stenotrophomonas maltophilia ATCC®13636 (American Type Culture Collection), Neisseria subflava ATCC® 14221, and Bacillus subtilis ATCC® 6633. All selected for investigation are environmental, non-pathogenic, and aerobic bacteria, and are used in a variety of applications.
Culture preparation
The strains were incubated in Bacto™ Tryptic Soy Broth (TSB, Becton Dickinson) for 24 h (S. carnosus, B. subtilis) or 48 h (S. maltophilia, N. subflava) at 37 °C under gentle shaking. Bacteria were harvested by centrifugation at 4200 rpm for 5 min and washed three times with deionized water. Afterwards, the cell pellets were re-suspended in deionized water and diluted to an optical density OD600 of 0.5 corresponding to the cell concentration of ~ 3 × 108 CFU/mL (colony forming units) according to McFarland standards. Bacterial suspensions were plated on agar plates and counted to confirm the cells concentration independently. The prepared suspensions (~ 3 × 108 CFU/mL) were used in all the protocols, described in details in the following subsections: “Characterization of bacterial zeta potential,” “Scanning electron microscopy,” “Transmission electron microscopy.”
Characterization of bacterial zeta potential
The zeta potential of the investigated bacteria in water suspensions was determined via electrophoretic light scattering in Zetasizer Nano ZS (Malvern Instruments, Malvern) equipment, always in the same conditions, i.e., temperature, growth phase of bacteria, preparation time, and pH. The ζ-potential value is an average of at least three independent experimental series for each bacterial strain (more than 15 independent measurements). The ζ- potential values of bacterial suspensions were measured before each experiment, following the protocols described in details in the “Scanning electron microscopy” and “Transmission electron microscopy” subsections.
Synthesis of gold nanoparticles
HAuCl4 solution (99%) supplied by Sigma-Aldrich and trisodium citrate dihydrate (99%) supplied by Avantor Performance Materials Poland S.A. were used as substrates for the preparation of gold nanoparticles as described elsewhere in more detail (Bastús et al. 2011). Shortly, gold seeds were synthesized by injecting HAuCl4 (1 mL, 25 mM) into sodium citrate (150 mL, 2.2 mM) at 100 °C. Once the synthesis was finished, the reaction mixture was cooled down to 90 °C. Then, 1 mL of sodium citrate solution (60 mM) and once more 1 mL of a HAuCl4 solution (25 mM) were sequentially added with the time interval of 2 min. The resulting particles were coated with negatively charged citrate ions (capping agent) and hence are well suspended in H2O. After 30 min, the reaction was finished, and aliquots of 2 mL were extracted for further characterization by nanoparticle tracking analysis (NTA) and transmission electron microscopy (TEM).
Finally, the prepared suspension of AuNPs was contacted with various bacterial strains for 15 min, following the protocols described in details in the “Scanning electron microscopy” and “Transmission electron microscopy” subsections.
Nanoparticle tracking analysis
The AuNP size distribution was determined using the LM10 Nanosight instrument (Malvern Instruments Ltd) equipped with a sCMOS camera (Hamamatsu Photonics, Hamamatsu, Japan) and a 450-nm blue laser. The obtained data were processed with NTA software (version 3.1 Build 3.1.45). The samples were sonicated for 15 min before analysis. The viscosity settings for water were automatically corrected for the experiment temperature (22 °C). The size distribution was measured with the camera shutter setting of 1232 and the gain of 219. A single experiment consisted of 3 movies at 25 frames/s and a duration of 30 s.
Characterization of gold nanoparticles in suspension
The zeta potential (ζ-potential) of the suspended gold nanoparticles was measured via electrophoretic light scattering by means of a Zetasizer Nano ZS (Malvern Instruments). The pH of the suspension was 6.5.
Scanning electron microscopy
The investigated bacterial strains were cultured on a glass substrate and then fixed for the SEM observations according to the protocol described elsewhere (Chissoe et al. 1994; Fischer et al. 2012). Briefly, samples on a glass substrate were fixed in 3% buffered glutaraldehyde for 24 h, and then carefully washed two times with Dulbecco’s Phosphate Buffered Saline (DPBS). They were next dehydrated in the water-alcohol solutions with gradually increasing ethanol concentration (50, 60, 70, 80, 90, 96, and 100%) for 10 min each. Finally, the samples were dried using hexamethyldisilazane (HMDS). The prepared samples were mounted on a SEM holder by using adhesive carbon tape and carbon conductive paint. Prior to the observations, the specimens were coated with a thin layer of gold (approximately 15 nm) with the use of sputter-coater (Quorum Q150T S).
For SEM observations of the bacteria coated with Au nanoparticles, the preparation procedure was simplified to avoid removal of the adsorbed NPs from the bacterial walls. Therefore, the bacteria were cultured on a conductive ITO (indium tin oxide) substrate, the supernatant was removed, and 1 mL of AuNP suspension was added. After 15 min of incubation (room temperature) with the AuNPs, the solution was removed, and the bacterial samples were dried in the air.
The samples were characterized with the use of the field-emission scanning electron microscope (FE-SEM, Hitachi S-4700). The SEM microphotographs were analyzed using the Java open-source ImageJ software 1.51 k (Eliceiri et al. 2012). The obtained images allowed for the evaluation of the average area of individual bacterial cells and the number of the adsorbed AuNPs. The quantification of the adsorption efficiency (expressed as the number of NPs per μm2 of the bacterial surface area) was determined by analyzing at least ten bacterial cells from three independent experimental series for each of the investigated bacterial strain.
Transmission electron microscopy
Transmission electron microscopy (TEM) observations were carried out using a Tecnai Osiris instrument (FEI) with the X-FEG Schottky field emitter operated at an accelerating voltage of 200 kV. Samples for TEM characterization were prepared by the standard procedure described elsewhere (Romero-Brey and Bartenschlager 2015). The investigated bacterial strain suspensions (after the 15 min contact with the AuNPs suspension) were washed three times in DPBS, fixed in 3% buffered glutaraldehyde for 24 h. The pellets were washed three times in DPBS, rinsed with 1% osmium tetraoxide solution in DPBS for 2 h, and washed again with DPBS. Samples were dehydrated in the water-alcohol solutions with gradually increasing ethanol concentration (50, 60, 70, 80, 90, 96, and 100%) for 15 min each. The pellets were rinsed with propylene oxide (20 min) and incubated in 1:1 propylene oxide/resin ratio overnight (Durcupan, Sigma-Aldrich). Then, samples were incubated in 100% resin at 37 °C for 24 h and then 60 °C for 48 h. The samples were sectioned using an ultramicrotome (Leica) equipped with the glass-edged knife (Diatome). The ultrathin lamellas were placed onto Cu TEM slots with the carbon-coated membrane and stained with lead citrate and uranyl citrate for contrast enhancement.
Results and discussion
Bacteria characteristics
For the present study, only non-pathogenic bacteria, naturally occurring in the environment were used. The selected bacterial strains differ in their cell wall structure, zeta potential, cell morphology (spherical or rod-shaped), and growth kinetics (stationary phase after 24 or 48 h). The representative SEM microphotographs of the investigated bacterial cells are presented in Fig. 2.
In the upper panel, the groups of tested bacteria (S. maltophilia, N. subflava, S. carnosus, B. subtilis) are shown, whereas the lower panel illustrates the different morphologies of the corresponding single cells in more detail. S. maltophilia is a rod-shaped G− aerobic and motile bacteria occurring naturally in soil, water, and plants. In line with the literature data, the estimated average length of a single cell ranges between 0.5 and 1.5 μm. N. subflava is a non-pathogenic G−, aerobic coccus, grouped in pairs (diplococci) with an average diameter in the range of 0.5–1 μm. S. carnosus, a typical G+ coccus, is spherical in shape and forms grape-like clusters. The average size of a single cell ranges between 0.5 and 1.5 μm. Finally, B. subtilis is a G+ rod-shaped (1.5–3.0 μm) aerobic microorganism, which naturally occupied soil and plants. As a consequence of various size and morphology, the selected bacteria N. subflava, S. carnosus, S. maltophilia, B. subtilis occupy a significantly different surface area of a substrate: 0.64; 0.79; 1.43, and 1.48 μm2, respectively.
Zeta-potentials of the bacterial strains
The interaction between bacteria and nanoparticles is mediated by electrostatic/dispersive forces between the bacteria and NP surface functional groups (Hwang et al. 2012). Therefore, sensible information about an average surface charge, crucial for the electrostatic interaction between nanoparticles and biological moieties, can be gained from the zeta potential values (Kłodzińska et al. 2010). Therefore, this parameter was measured for all the bacterial strains and AuNPs used in this study. The measured values of the ζ-potential of the bacteria suspended in deionized water at physiological pH value of 6.5 ± 0.1 are in the range from – 41 mV (B. subtilis) to − 37 mV (S. carnosus), whereas significantly higher values were determined for N. subflava (− 30 mV) and S. maltophilia (− 26 mV). The obtained results are in line with the reported zeta-potential for various bacterial strains, which changes from − 3 to − 50 mV (Ahimou et al. 2007; Nguyen et al. 2011; Hua et al. 2017). Since the actual value of ζ-potential strongly depends on the physicochemical conditions of the suspension, all the measurements were standardized with respect to the electrolyte type and concentration. Due to the different chemical nature of the G+ and G− bacterial walls (see Fig. 1), the accumulated surfaces charges (both negative) are substantially different with the lower values obtained for the G+ strains. The observed differences in the ζ-potential have a strong impact on the bacterial adhesive properties, as discussed below.
Characteristics of the gold nanoparticles
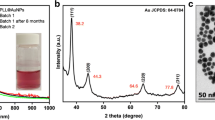
In order to evaluate the adhesive properties of the investigated bacterial strains, monodisperse AuNPs were used. As described elsewhere (Hwang et al. 2012), size and surface charge of NPs are two main factors playing a critical role in their adsorption on the bacterial surface. As presented in Fig. 3, the average size of the synthesized AuNPs, assessed from the NTA measurements, was 50 nm (± 0.7 nm), and their average concentration was equal to 7.5 × 1012/mL. For more in-depth insight into the size distribution and morphology, TEM observations were performed, and the results are presented in the insets of Fig. 3.
Insets a and b (Fig. 3) clearly illustrate a rhombicuboctahedral morphology typically observed for Au nanoparticles (Louis and Pluchery 2012) and reveal a smaller diameter (~ 30 nm), when compared with the results obtained from the NTA measurements. Indeed, the NTA-determined size represents the hydrodynamic diameter of the particle, always of the higher value. It should be noticed, however, that both the complementary methods revealed the narrow size distribution of the synthesized Au nanoparticles, as described elsewhere (Bastús et al. 2011), with the characteristic value of the ζ-potential equal to − 7 mV (± 0.6 mV).
Interaction between bacterial walls and nanoparticles
In order to quantify the number of gold nanoparticles adsorbed on various bacterial strains, SEM microphotographs of the investigated bacteria after incubation in the AuNP suspension were analyzed. Figure 4 shows representative examples of the bacterial cells decorated with the gold nanoparticles.
It can be easily noticed that the investigated bacteria exhibit dramatically different affinity to gold nanoparticles attachment on their surfaces. Taking into account that the NP suspension and the conditions of all the incubation experiments (temperature, time, and pH) were kept the same, it is clear that the various bacterial strains captured the substantially different amount of gold nanoparticles. Notably, the G+ bacteria (S. carnosus and B. subtilis) adhered much more AuNPs when compared with the G− ones (S. maltophilia and N. subflava). These observations were corroborated by more detailed TEM investigations of the bacteria coated with AuNPs. In Fig. 5, the cross sections (~ 60-nm-thick lamellas) of single resin-embedded cells of the studied bacteria decorated with AuNPs are presented. They illustrate the somehow different shape of the bacterial cells than observed in SEM images, but it should be taken into account that the cells were cross-sectioned at random orientations. The SEM and TEM microscopic observations revealed that the nanoparticles do not penetrate into the bacterial cells. This can be explained by relatively big nanoparticles size (30 nm) compared with the pores in the cell wall (4–16 nm) (Turner et al. 2013), and short time of incubation (15 min).
In accordance with the model of the bacterial walls presented in Fig. 1, for the G− species (S. maltophilia and N. subflava), significantly thinner cell wall structure of about 20 nm is observed, whereas for the G+ (S. carnosus and B. subtilis) the walls are in the range of 30–50 nm. Such a feature is characteristic for the G+ strains due to the thick peptidoglycan layer situated above the cell membrane. In general, the TEM images confirmed the SEM observations revealing the various numbers of NPs adhered on the bacterial cells. Whereas for the S. maltophilia, N. subflava, and S. carnosus, the adhered nanoparticles are well separated by a distance from several to tens of their dimensions; in the case of B. subtilis, a pronounced load of nanoparticles leads to the surface agglomeration of the excessive gold nanoparticles.
The microscopic observations of the adhered AuNPs to the bacterial cell walls were quantified as a number of nanoparticles covering 1 μm2 of the projected bacterial surface area. The results are summarized in Fig. 6 (grey bars).
The highest number of the NPs attached to the cell wall surface was observed for B. subtilis (103 NPs) and the lowest for S. maltophilia (60 NPs). For N. subflava and S. carnosus, the intermediate values of ~ 1.5 × 102 NPs and ~ 4.0 × 102 were observed, respectively. In the same figure, the experimental values of the ζ-potential for the investigated bacterial strains are also shown (red plot). The trend in the amount of AuNPs adhered to the bacterial cell surface B. subtilis >> S. carnosus > N. subflava > S. maltophilia correlates well with the values of ζ-potential: the higher the negative charge is, the higher the adhesion extends. It may be thus concluded that the ζ-potential plays the role of a simple, concise descriptor of the adsorption process in the bacteria–nanoparticle systems.
The bacteria and nanoparticles being suspended in the aqueous solution can be treated as interacting colloidal particles, and their behavior can be accounted for in terms of the DLVO model as a reasonable approximation. The potential interaction energy profiles between bacteria and nanoparticles were calculated based on the Derjaguin–Landau–Verwey–Overbeek (DLVO) theory (Israelachvili 2011; Hwang et al. 2012; Ohshima 2014). The total interaction energy (Vtot = VEL + VLW) results from electrostatic repulsive interactions (VEL) and attractive London-van der Waals forces (VLW). Based on previous studies (Hwang et al. 2012), the model for two interacting spherical particles with the radii a1 (bacteria) and a2 (nanoparticle) separated by the distance d can be approximated as
where A is the Hamaker constant calculated as described elsewhere (Ohshima 2014) (1.72 × 10−20 J); ε is expressed as the product of the relative permittivity of the medium εr, which is 80 for water at 20 °C, and the permittivity of a vacuum ε0 (8.854 × 10−12 C2/Jm); and ζ1 and ζ2 are the zeta potentials of bacteria and nanoparticles, respectively, whereas κ is the inverse Debye−Hückel length (729 nm for pH = 6.5) calculated according to (Israelachvili 2011). The Hamaker constant for this system, where the bacterial cells are interacting with nanoparticles in water, was calculated following the formula (Ohshima 2014):
where A132 is the Hamaker constant for the interaction between bacteria (A1 = 5.2 × 10−20 J) (Farahat et al. 2009) and gold nanoparticles (A2 = 45.3 × 10−20 J) (Israelachvili 2011) separated by a water medium (A3 = 3.7 × 10−20 J) (Israelachvili 2011). The value of Hamaker constant for bacterial cells was adapted from a similar research focused on the interaction of various bacterial strains with surfaces. Additionally, we checked several other Hamaker values (Liu et al. 2007; Farahat et al. 2009; Harimawan et al. 2013; Yoshihara et al. 2015) within the range of 10−23–10−20 J and found out that both the trends in critical radius and the primary energy minima are not sensitive to A1, at least in the realistic range for bacteria. Thus, in our opinion, the DLVO model is helpful in interpreting our experimental findings.
For calculations, the spherical shapes of bacteria with an equivalent radius of 400, 440, 460, and 520 nm were assumed for N. subflava, S. carnosus, S. maltophilia, and B. subtilis, respectively. The latter value was assessed as an average radius of a model spherical cell corresponding to the actual cylindrical/spherical bacterial shapes with the same volume. The resultant DLVO energy profiles as a function of a separation distance for the tested bacterial strains interacting with AuNPs are presented in Fig. 7.
The calculated DLVO energy profiles exhibit rather small activation barriers of 1.45–1.55 kT at 500–800 nm (inset a) and the sufficiently deep primary energy minima of 120–170 kT (inset b) for the adsorption process to be effective. It is also worth to underline that the depth of the observed primary minima for each of the bacterial strain is in line with the amount of the adhered nanoparticles observed in the microscopic measurements (see Figs. 4, 5, and 6).
In Fig. 8, a reasonably good linear correlation of the number of gold nanoparticles as a function of a critical radius (rcrit) to the third power is shown. The critical radius is defined by a formula: rcrit3 = R3 − a13, where R is designated by a location of energy maxima in DLVO profile for each bacteria of radius a1 (see upper inset in Fig. 7). The value of rcrit3 gauges the critical volume around the bacterial cell, where the attraction forces dominate. It may be interpreted in terms that the observed differences in the attachment efficiency for various strains result from the availability of the nanoparticles in the critical volume of the suspension surrounding the bacterial cells.
It is worth noting that the obtained results have more general relevance and practical applications. Indeed, owing to a non-specific bacteria–nanoparticle interaction governed by the ζ-potential essentially, the established relationships can be readily extended for other systems. Due to the remarkable adhesion properties of B. subtilis, we can explain why these specific microorganisms are predestined as a template to agglomerate Au nanoparticles into organized long ribbons, as shown elsewhere (He et al. 2006). Understanding the bacteria–nanoparticles interactions are especially important for designing of biocompatible and stable biosensors. The subject is intensively investigated nowadays, and there are several examples illustrating the applications of AuNPs in colorimetric biosensors array (see, e.g., Verma et al. 2016). The presented results provide the basis for future advancement of a fast, simple, label-free method of bacteria detection in clinical diagnostics and environmental monitoring (Li et al. 2015). Besides the biosensors, the effective adhesion of AuNPs to bacterial walls can also help in a thorough biochemical characterization of bacteria via SERS method, as discussed elsewhere (Lin et al. 2014; Zhou et al. 2015).
Conclusions
In this paper, the adhesion efficiency of gold nanoparticles (30 nm) for Gram-positive (B. subtilis, S. carnosus) and Gram-negative (N. subflava, S. maltophilia) bacterial strains was examined. It was found out that among the investigated strains the attachment efficiency decreases in the order B. subtilis >> S. carnosus > N. subflava > S. maltophilia. A direct correlation between the number of the attached nanoparticles and the ζ-potential of the bacterial strains was established, and the results were accounted for within the DLVO theory. The small activation barriers for the adhesion process (1.45–1.55 kT), together with rather deep primary energy minima (120–170 kT) are favorable for the effective adsorption process. The linear correlation between the nanoparticles adhered to the bacterial cells surface and the size of the critical volume around the bacterial cells where the attraction forces dominate implies the dramatic differences in the attachment efficiency result from the availability of the nanoparticles in the critical volume of the surrounding suspensions.
References
Ahimou F, Boonaert CJP, Adriaensen Y, Jacques P, Thonart P, Paquot M, Rouxhet PG (2007) XPS analysis of chemical functions at the surface of Bacillus subtilis. J Colloid Interface Sci 309:49–55. https://doi.org/10.1016/j.jcis.2007.01.055
Aruoja V, Pokhrel S, Sihtmäe M, Mortimer M, Mädler L, Kahru A (2015) Toxicity of 12 metal-based nanoparticles to algae, bacteria and protozoa. Environ Sci Nano 2:630–644. https://doi.org/10.1039/c5en00057b
Bar-Ilan O, Albrecht RM, Fako VE, Furgeson DY (2009) Toxicity assessments of multisized gold and silver nanoparticles in zebrafish embryos. Small 5:1897–1910. https://doi.org/10.1002/smll.200801716
Barnes RJ, Molina R, Xu J, Dobson PJ, Thompson IP (2013) Comparison of TiO2 and ZnO nanoparticles for photocatalytic degradation of methylene blue and the correlated inactivation of gram-positive and gram-negative bacteria. J Nanopart Res 15:1432. https://doi.org/10.1007/s11051-013-1432-9
Bastús NG, Comenge J, Puntes V (2011) Kinetically controlled seeded growth synthesis of citrate-stabilized gold nanoparticles of up to 200 nm. Langmuir 27:11098–11105. https://doi.org/10.1021/la201938u
Bozich JS, Lohse SE, Torelli MD, Murphy CJ, Hamers RJ, Klaper RD (2014) Surface chemistry, charge and ligand type impact the toxicity of gold nanoparticles to Daphnia magna. Environ Sci Nano 1:260–270. https://doi.org/10.1039/c4en00006d
Bozorgmehr A, Yazdanparast R, Mollasalehi H (2016) Non-crosslinking gold nanoprobe-LAMP for simple, colorimetric, and specific detection of Salmonella typhi. J Nanopart Res 18. https://doi.org/10.1007/s11051-016-3657-x
Braslavsky I, Dolev MB, Bernheim R et al (2016) Putting life on ice: bacteria that bind to frozen water. J R Soc Interface 13:1–11. https://doi.org/10.1098/rsif.2016.0210
Bresee J, Maier KE, Boncella AE, Melander C, Feldheim DL (2011) Growth inhibition of staphylococcus aureus by mixed monolayer gold nanoparticles. Small 7:2027–2031. https://doi.org/10.1002/smll.201100420
Calatayud MP, Sanz B, Raffa V, Riggio C, Ibarra MR, Goya GF (2014) The effect of surface charge of functionalized Fe3O4 nanoparticles on protein adsorption and cell uptake. Biomaterials 35:6389–6399. https://doi.org/10.1016/j.biomaterials.2014.04.009
Carrière M (2012) What about toxicity and ecotoxicity of gold nanoparticles? In: Louis C, Pluchery O (eds) Gold nanoparticles for physics. Chemistry And Biology. Imperial College Press, London, pp 333–353
Chissoe WF, Vezey EL, Skvarla JJ (1994) Hexamethyldisilazane as a drying agent for pollen scanning electron microscopy. Biotech Histochem 69:192–198. https://doi.org/10.3109/10520299409106286
Chwalibog A, Sawosz E, Hotowy A et al (2010) Visualization of interaction between inorganic nanoparticles and bacteria or fungi. Int J Nanomedicine 5:1085–1094. https://doi.org/10.2147/IJN.S13532
Clifton LA, Skoda MWA, Daulton EL, Hughes AV, le Brun AP, Lakey JH, Holt SA (2013) Asymmetric phospholipid: lipopolysaccharide bilayers; a gram-negative bacterial outer membrane mimic. J R Soc Interface 10:20130810. https://doi.org/10.1098/rsif.2013.0810
Deplanche K, Merroun ML, Casadesus M, Tran DT, Mikheenko IP, Bennett JA, Zhu J, Jones IP, Attard GA, Wood J, Selenska-Pobell S, Macaskie LE (2012) Microbial synthesis of core/shell gold/palladium nanoparticles for applications in green chemistry. J R Soc Interface 9:1705–1712. https://doi.org/10.1098/rsif.2012.0003
Donlan RM, Costerton JW (2002) Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 15:167–193. https://doi.org/10.1128/CMR.15.2.167
Donmez M, Yilmaz MD, Kilbas B (2017) Fluorescent detection of dipicolinic acid as a biomarker of bacterial spores using lanthanide-chelated gold nanoparticles. J Hazard Mater 324:593–598. https://doi.org/10.1016/j.jhazmat.2016.11.030
Durán N, Marcato PD, Durán M, Yadav A, Gade A, Rai M (2011) Mechanistic aspects in the biogenic synthesis of extracellular metal nanoparticles by peptides, bacteria, fungi, and plants. Appl Microbiol Biotechnol 90:1609–1624. https://doi.org/10.1007/s00253-011-3249-8
Dykman LA, Khlebtsov NG (2011) Gold nanoparticles in biology and medicine: recent advances and prospects. Acta Nat 3:34–55. https://doi.org/10.1039/c0cs00018c
Eliceiri K, Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to ImageJ: 25 years of image analysis HISTORICAL commentary NIH image to ImageJ : 25 years of image analysis. Nat Methods 9:671–675. https://doi.org/10.1038/nmeth.2089
Faisal M, Saquib Q, Alatar AA, Al-Khedhairy AA (2018) Cytotoxicity of nanoparticles. Phytotoxicity of Nanoparticles:1–407. https://doi.org/10.1007/978-3-319-76708-6
Farahat M, Hirajima T, Sasaki K, Doi K (2009) Adhesion of Escherichia coli onto quartz, hematite and corundum: extended DLVO theory and flotation behavior. Colloids Surf B 74:140–149. https://doi.org/10.1016/j.colsurfb.2009.07.009
Fischer ER, Hansen BT, Nair V et al (2012) Scanning electron microscopy. Curr Protoc Microbiol. https://doi.org/10.1002/9780471729259.mc02b02s25
Gatea F, Teodor ED, Seciu AM et al (2015) Antitumour, antimicrobial and catalytic activity of gold nanoparticles synthesized by different pH propolis extracts. J Nanoparticle Res 17. https://doi.org/10.1007/s11051-015-3127-x
Harimawan A, Zhong S, Lim CT, Ting YP (2013) Adhesion of B. subtilis spores and vegetative cells onto stainless steel - DLVO theories and AFM spectroscopy. J Colloid Interface Sci 405:233–241. https://doi.org/10.1016/j.jcis.2013.05.031
He Y, Yuan J, Su F, Xing X, Shi G (2006) Bacillus subtilis assisted assembly of gold nanoparticles into long conductive nodous ribbons. J Phys Chem B 110:17813–17818. https://doi.org/10.1021/jp063729o
Hua XW, Bao YW, Wang HY, Chen Z, Wu FG (2017) Bacteria-derived fluorescent carbon dots for microbial live/dead differentiation. Nanoscale 9:2150–2161. https://doi.org/10.1039/c6nr06558a
Hwang G, Ahn IS, Mhin BJ, Kim JY (2012) Adhesion of nano-sized particles to the surface of bacteria: mechanistic study with the extended DLVO theory. Colloids Surf B 97:138–144. https://doi.org/10.1016/j.colsurfb.2012.04.031
Israelachvili J (2011) Intermolecular and surface forces. Elsevier, New York
Judy JD, Bertsch PM (2014) Bioavailability, toxicity, and fate of manufactured nanomaterials in terrestrial ecosystems. Adv Agron 123:1–64. https://doi.org/10.1016/B978-0-12-420225-2.00001-7
Jung JH, Lee SY, Seo TS (2018) In vivo synthesis of nanocomposites using the recombinant Escherichia coli. Small 14:1–7. https://doi.org/10.1002/smll.201803133
Khelissa SO, Jama C, Abdallah M, Boukherroub R, Faille C, Chihib NE (2017) Correction to: Effect of incubation duration, growth temperature, and abiotic surface type on cell surface properties, adhesion and pathogenicity of biofilm-detached Staphylococcus aureus cells. AMB Express 7:216. https://doi.org/10.1186/s13568-017-0509-8
Kłodzińska E, Szumski M, Dziubakiewicz E, Hrynkiewicz K, Skwarek E, Janusz W, Buszewski B (2010) Effect of zeta potential value on bacterial behavior during electrophoretic separation. Electrophoresis 31:1590–1596. https://doi.org/10.1002/elps.200900559
Li X, Robinson SM, Gupta A, Saha K, Jiang Z, Moyano DF, Sahar A, Riley MA, Rotello VM (2014) Functional gold nanoparticles as potent antimicrobial agents against multi-drug-resistant bacteria. ACS Nano 8:10682–10686. https://doi.org/10.1021/nn5042625
Li D, Dong Y, Li B, Wu Y, Wang K, Zhang S (2015) Colorimetric sensor array with unmodified noble metal nanoparticles for naked-eye detection of proteins and bacteria. Analyst 140:7672–7677. https://doi.org/10.1039/c5an01267h
Lin HY, Huang CH, Hsieh WH, Liu LH, Lin YC, Chu CC, Wang ST, Kuo IT, Chau LK, Yang CY (2014) On-line SERS detection of single bacterium using novel SERS nanoprobes and a microfl uidic dielectrophoresis device. Small 10:4700–4710. https://doi.org/10.1002/smll.201401526
Liu XM, Sheng GP, Yu HQ (2007) DLVO approach to the flocculability of a photosynthetic H 2-producing bacterium, Rhodopseudomonas acidophila. Environ Sci Technol 41:4620–4625. https://doi.org/10.1021/es070107n
Loeb S, Li C, Kim JH (2018) Solar photothermal disinfection using broadband-light absorbing gold nanoparticles and carbon black. Environ Sci Technol 52:205–213. https://doi.org/10.1021/acs.est.7b04442
López-Lorente ÁI, Cárdenas S, González-Sánchez ZI (2019) Effect of synthesis, purification and growth determination methods on the antibacterial and antifungal activity of gold nanoparticles. Mater Sci Eng C 103:109805. https://doi.org/10.1016/j.msec.2019.109805
Louis C, Pluchery O (2012) Gold nanoparticles for physics, chemistry and biology. Imperial College Press, London
Lovering AL, Lin LYC, Sewell EW, Spreter T, Brown ED, Strynadka NCJ (2010) Structure of the bacterial teichoic acid polymerase TagF provides insights into membrane association and catalysis. Nat Struct Mol Biol 17:582–589. https://doi.org/10.1038/nsmb.1819
Malanovic N, Lohner K (2016) Gram-positive bacterial cell envelopes: the impact on the activity of antimicrobial peptides. Biochim Biophys Acta Biomembr 1858:936–946. https://doi.org/10.1016/j.bbamem.2015.11.004
McCracken C, Dutta PK, Waldman WJ (2016) Critical assessment of toxicological effects of ingested nanoparticles. Environ Sci Nano 3:256–282. https://doi.org/10.1039/c5en00242g
Miller SE, Bell CS, Mejias R et al (2016) Colistin-functionalized nanoparticles for the rapid capture of acinetobacter baumannii. J Biomed Nanotechnol 12:1806–1819. https://doi.org/10.1166/jbn.2016.2273
Mmola M, Roes-Hill M, Durrell K, Bolton J, Sibuyi N, Meyer M, Beukes D, Antunes E (2016) Enhanced antimicrobial and anticancer activity of silver and gold nanoparticles synthesised using Sargassum incisifolium aqueous extracts. Molecules 21:1633. https://doi.org/10.3390/molecules21121633
Moriarty TF, Poulsson AHC, Rochford ETJ, Richards RG (2011) Bacterial adhesion and biomaterial surfaces. Compr Biomater 4:75–100. https://doi.org/10.1016/B978-0-08-055294-1.00007-6
Nangia Y, Wangoo N, Goyal N, Shekhawat G, Suri CR (2009) A novel bacterial isolate Stenotrophomonas maltophilia as living factory for synthesis of gold nanoparticles. Microb Cell Factories 8:1–7. https://doi.org/10.1186/1475-2859-8-39
Nath D, Banerjee P (2013) Green nanotechnology - a new hope for medical biology. Environ Toxicol Pharmacol 36:997–1014. https://doi.org/10.1016/j.etap.2013.09.002
Nguyen VT, Turner MS, Dykes GA (2011) Influence of cell surface hydrophobicity on attachment of Campylobacter to abiotic surfaces. Food Microbiol 28:942–950. https://doi.org/10.1016/j.fm.2011.01.004
Ohshima H (2014) Interaction of colloidal particles. In: Ohshima H, Makino K (eds) Colloid and interface science in pharmaceutical research and development. Elsevier, New York, pp 1–28
Pradeepa, Vidya SM, Mutalik S et al (2016) Preparation of gold nanoparticles by novel bacterial exopolysaccharide for antibiotic delivery. Life Sci 153:171–179. https://doi.org/10.1016/j.lfs.2016.04.022
Rai M, Yadav A, Gade A (2009) Silver nanoparticles as a new generation of antimicrobials. Biotechnol Adv 27:76–83. https://doi.org/10.1016/j.biotechadv.2008.09.002
Ramasamy M, Lee SS, Yi DK, Kim K (2014) Magnetic, optical gold nanorods for recyclable photothermal ablation of bacteria. J Mater Chem B 2:981–988. https://doi.org/10.1039/c3tb21310b
Ramasamy M, Lee JH, Lee J (2016) Potent antimicrobial and antibiofilm activities of bacteriogenically synthesized gold-silver nanoparticles against pathogenic bacteria and their physiochemical characterizations. J Biomater Appl 31:366–378. https://doi.org/10.1177/0885328216646910
Romero-Brey I, Bartenschlager R (2015) Viral infection at high magnification: 3D electron microscopy methods to analyze the architecture of infected cells. Viruses 7:6316–6345. https://doi.org/10.3390/v7122940
Saha K, Agasti SS, Kim C, Li X, Rotello VM (2012) Gold nanoparticles in chemical and biological sensing. Chem Rev 112:2739–2779. https://doi.org/10.1021/cr2001178
Sajjanar B, Kakodia B, Bisht D, Saxena S, Singh AK, Joshi V, Tiwari AK, Kumar S (2015) Peptide-activated gold nanoparticles for selective visual sensing of virus. J Nanopart Res 17:1–9. https://doi.org/10.1007/s11051-015-3043-0
Silhavy T, Kahne D, Walker S (2010) The bacterial cell envelope. Cold Spring Harb Perspect Biol 2:1–16. https://doi.org/10.1101/cshperspect.a000414
Spriano S, Sarath Chandra V, Cochis A, Uberti F, Rimondini L, Bertone E, Vitale A, Scolaro C, Ferrari M, Cirisano F, Gautier di Confiengo G, Ferraris S (2017) How do wettability, zeta potential and hydroxylation degree affect the biological response of biomaterials? Mater Sci Eng C 74:542–555. https://doi.org/10.1016/j.msec.2016.12.107
Srivastava SK, Yamada R, Ogino C, Kondo A (2013) Biogenic synthesis and characterization of gold nanoparticles by Escherichia coli K12 and its heterogeneous catalysis in degradation of 4-nitrophenol. Nanoscale Res Lett 8:70. https://doi.org/10.1186/1556-276X-8-70
The European Commission (2011) Commission recommendation of 18 October 2011 on the definition of nanomaterial. Off J Eur Union:38–40
Turner RD, Hurd AF, Cadby A, Hobbs JK, Foster SJ (2013) Cell wall elongation mode in Gram-negative bacteria is determined by peptidoglycan architecture. Nat Commun 4:1496–1498. https://doi.org/10.1038/ncomms2503
Verma MS, Wei SC, Rogowski JL, Tsuji JM, Chen PZ, Lin CW, Jones L, Gu FX (2016) Interactions between bacterial surface and nanoparticles govern the performance of “chemical nose” biosensors. Biosens Bioelectron 83:115–125. https://doi.org/10.1016/j.bios.2016.04.024
von Moos N, Bowen P, Slaveykova VI (2014) Bioavailability of inorganic nanoparticles to planktonic bacteria and aquatic microalgae in freshwater. Environ Sci Nano 1:214. https://doi.org/10.1039/c3en00054k
Wadhwani SA, Shedbalkar UU, Singh R, Chopade BA (2016) Biogenic selenium nanoparticles: current status and future prospects. Appl Microbiol Biotechnol 100:2555–2566. https://doi.org/10.1007/s00253-016-7300-7
Wilson WW, Wade MM, Holman SC, Champlin FR (2001) Status of methods for assessing bacterial cell surface charge properties based on zeta potential measurements. J Microbiol Methods 43:153–164. https://doi.org/10.1016/S0167-7012(00)00224-4
Xing X, Ma W, Zhao X, Wang J, Yao L, Jiang X, Wu Z (2018) Interaction between surface charge-modified gold nanoparticles and phospholipid membranes. Langmuir 34:12583–12589. https://doi.org/10.1021/acs.langmuir.8b01700
Yang C, Bromma K, Di Ciano-Oliveira C et al (2018) Gold nanoparticle mediated combined cancer therapy. Cancer Nanotechnol:9. https://doi.org/10.1186/s12645-018-0039-3
Yoshihara A, Nobuhira N, Narahara H, Toyoda S, Tokumoto H, Konishi Y, Nomura T (2015) Estimation of the adhesive force distribution for the flagellar adhesion of Escherichia coli on a glass surface. Colloids Surf B Biointerfaces 131:67–72. https://doi.org/10.1016/j.colsurfb.2015.04.038
Young KD (2006) The selective value of bacterial shape. Microbiol Mol Biol Rev 70:660–703. https://doi.org/10.1128/MMBR.00001-06
Zhang Y, Shareena Dasari TP, Deng H, Yu H (2015) Antimicrobial activity of gold nanoparticles and ionic gold. J Environ Sci Health C Environ Carcinog Ecotoxicol Rev 33:286–327. https://doi.org/10.1080/10590501.2015.1055161
Zhou W, Gao X, Liu D, Chen X (2015) Gold nanoparticles for in vitro diagnostics. Chem Rev 115:10575–10636. https://doi.org/10.1021/acs.chemrev.5b00100
Zhu Y, Sun Q, Liu Y, Ma T, Su L, Liu S, Shi X, Han D, Liang F (2018) Decorating gold nanostars with multiwalled carbon nanotubes for photothermal therapy. R Soc Open Sci:5. https://doi.org/10.1098/rsos.180159
Acknowledgments
W. Pajerski acknowledges the fellowship with the project no. POWR.03.02.00-00-I013/16.
The SEM imaging was performed in the Laboratory of Field Emission Scanning Electron Microscopy and Microanalysis at the Institute of Geological Sciences, Jagiellonian University, Poland.
Funding
The study was financed by the Polish National Science Centre project awarded by decision number DEC-2016/21/B/ST8/00398.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Pajerski, W., Ochonska, D., Brzychczy-Wloch, M. et al. Attachment efficiency of gold nanoparticles by Gram-positive and Gram-negative bacterial strains governed by surface charges. J Nanopart Res 21, 186 (2019). https://doi.org/10.1007/s11051-019-4617-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11051-019-4617-z