Abstract
Background
Splice-disrupt genomic variants are one of the causes of cancer-causing errors in gene expression. Little is known about splice-disrupt genomic variants.
Methods and results
Here, pattern of splice-disrupt variants was investigated using 21,842,764 genomic variants in different types of prostate cancer. A particular attention was paid to genomic locations of splice-disrupt variants on target genes. HLA-A in prostate cancer, MSR1 in familial prostate cancer, and EGFR in both castration-resistant prostate cancer and metastatic castration-resistant had the highest allele frequencies of splice-disrupt variations. Some splice-disrupt variants, located on coding sequences of NCOR2, PTPRC, and CRP, were solely present in the advanced metastatic castration-resistant prostate cancer. High-risk splice-disrupt variants were identified based on computationally calculated Polymorphism Phenotyping (PolyPhen), Sorting Intolerant From Tolerant (SIFT), and Genomic Evolutionary Rate Profiling (GERP) + + scores as well as the recorded clinical significance in dbSNP database of NCBI. Functional annotation of damaging splice-disrupt variants highlighted important cancer-associated functions, including endocrine resistance, lipid metabolic process, steroid metabolic process, regulation of mitotic cell cycle, and regulation of metabolic process. This is the first study that profiles the splice-disrupt genomic variants and their target genes in prostate cancer. Literature mining based variant analysis highlighted the importance of rs1800716 variant, located on the CYP2D6 gene, involved in a range of important functions, such as RNA spicing, drug interaction, death, and urotoxicity.
Conclusions
This is the first study that profiles the splice-disrupt genomic variants and their target genes in different types of prostate cancer. Unravelling alternative splicing opens a new avenue towards the establishment of new diagnostic and prognostic markers for prostate cancer progression and metastasis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alternative RNA splicing is an emerging topic in molecular and clinical oncology [1, 2]. Alternative splicing is the key mechanism to generate many mRNA transcripts from the relatively low number of human genes, which can lead to the assembly of different protein isoforms with distinct functions. This structural modification of gene transcripts and their encoded proteins is considered a vital process that increases diversity of protein functions to generate the complex cellular proteome [3, 4]. The outcome of alternative splicing can result in a complete loss of function or the acquisition of new functions [3, 4]. In humans, it is estimated that up to 94% of genes undergo alternative splicing, resulting in more than 100,000 transcripts [5,6,7]. Accumulating evidence highlights the importance of study of gene and protein in parallel with alternative splice variants [8].
Dysregulation of post-transcriptional regulation can result in defective proteins or transcripts without causing genetic diseases [9]. Alternative pre-mRNA splicing leads to distinct products of gene expression in normal development and disease. Here, we observed higher variation in GERP++ score splice-disrupt variants of progressive prostate cancer compared to the non-progressive one. Antagonistic splice variants of genes involved in differentiation, apoptosis, invasion and metastasis often exist in a delicate equilibrium that is found to be perturbed in tumors.
Precise pre-mRNA splicing is vital for correct protein translation. Precise pre-mRNA splicing is related to the presence of consensus “cis” sequences, identifying exon-intron boundaries and regulatory sequences rby splicing machinery [10]. Point mutations may occur in both introns and exons disrupting existing splice sites or splicing regulatory sequences, generating new transcripts, even pathogenic ones [10]. Splice-disrupt genomic mutations can also be a source of cancer-causing errors in gene expression resulting in cancer-specific alternative splicing [11]. It has been demonstrated that nearly half of all active alternative splicing events are altered in ovarian and breast tumour cells compared to normal tissue [4]. Cancer can occur irrespective of changes in expression of a gene or protein, but rather because of aberrant splice variants that are linked to cancer progression and/or drug resistance and is compensated by the decreased expression of other splice variants originating from that same gene.
Within different types of cancer-associated genomic variants, splice-disrupt genomic variants are the less studied, particularly in prostate cancer. There is an ample evidence of the discovered point mutations that prevent appropriate splicing by disruption of exonic splicing enhancers [12]. Point mutation in the splicing acceptor or donor site can lead to the production of an altered mature mRNA and can result in intron retention, exon skipping, or alternative 3′ and 5′ splicing site [13]. Even, missense mutations or silent substitutions that do not alter protein function can influence on pre-mRNA splicing and have undiscovered pathological functions [12, 14]. In multiple sclerosis, splice-disrupt variants have been identified on 27 immune-related and myelin genes [15]. Splice-disrupt variants on NBAS, SLC16A1, RHD, PNPLA2 have been accounted as genetic basis of recurrent liver failure, ketoacidosis, variant D phenotype, progressive severe myopathy, respectively [16,17,18,19]. Pathogenic consequences of splice-disrupt variants in CHEK2 genes in many cancers has been documented in individuals carrying a single pathogenic splice-disrupt variant [13].
Our knowledge about abnormal splice-disrupt genomic variants in prostate cancer, and their prospective contribution to cancer progression (castration-resistant prostate cancer and metastatic castration-resistant prostate cancer) is limited. Furthermore, their relative abundance and their potential to use splice-disrupt in cancer diagnosis and treatment has been largely neglected [20].
Whole genome sequencing projects, particularly 1000 Genomes Project, have resulted in bulk identification of human variations, including splice-disrupt variants [21]. The identified variants progressively deposit in major public repositories of genomic variants, noticeably dbSNP, the NCBI database of genetic variation, to be employed by the researchers and clinicians worldwide [22]. The deposited data is a great resource to shed light on the possible involvement of splicing events in human diseases [2, 20].
For the first time, we mined a large dataset of genomic variants in different types of prostate cancer, identifing the splice-disrupt variants and their target genes. Furthermore, we investigated the association between splice-disrupt variants and advanced types of prostate cancer. Finally, computational systems biology was applied to investigate the functional consequences of the discovered splice-disrupt variants.
Methods
Identification of splice-disrupt variants in different types of prostate cancer and their genomic locations
The National Center for Biotechnology Information (NCBI) database of genetic variation (dbSNP database) [22] was used as the main resource for gathering of variants. Variants were retrieved for common and advanced types of prostate cancer (biological associations), including prostate cancer (PC), castration-resistant prostate cancer (CRPC), familial prostate cancer (FPC), and metastatic castration-resistant prostate cancer (MCRPC). SQL-based Pathway Studio Web tool (Elsevier) was used for navigating and downloading variants from dbSNP, as previously described [20].
Translational impact of variants including missense, splice-disrupt, coding sequences insertion or deletion (CDs indel), nonsense, misstart, and non-stop were retrieved and variants were filtered for splice-disrupt variants using SQL Table of Pathway Studio tool. As a result, splice-disrupt variants were identified for different types of prostate cancer (PC, CRPC, FPC, and MCRPC).
Characterising of splice-disrupt variants
Variant (allele) frequency
Allele (variant) frequency stands for a gene variant (an allele) in a specific locus in a population, commonly presented as percentage/fraction. Minor allele frequency explains the frequency where the second most common allele occurs in a population; for example, allele frequency of 1% mean that the variant happens in 1% of population. We used 1000 Genomes Project [21] to extract the minor allele frequency using Pathway Studio tool. No cutoff was used for minor allele frequency of splice-disrupt variants in this study. This allowed us to investigate the contribution of less frequent (1–5% minor allele frequency) and rare variants (< 1% minor allele frequency) to different types of prostate cancer.
Genomic locations
Genomic locations of variants were identified based on dbSNP, and variants were assigned to the following locations: CDs (coding sequences), 3′UTR (untranslated region), 5′UTR, intergenic, and intronic variants.
Scoring of splice-disrupt variants and identifying of high-risk splice-disrupt variants
High-risk splice-disrupt variants were identified using dbNSFP [23,24,25], a database of human non-synonymous SNVs and their functional predictions as well as dbSNP. Scoring of splice-disrupt variants was performed using computationally calculated Polymorphism Phenotyping (PolyPhen), Sorting Intolerant From Tolerant (SIFT), and Genomic Evolutionary Rate Profiling (GERP)++ scores from dbNSFP as well as the recorded clinical significance in dbSNP database of NCBI.
SIFT score
SIFT algorithm is developed to predict the effect of coding variants on protein function based on sequence homology and the physical properties of amino acids. SIFT is considered as a standard tool for characterizing missense variation [26, 27]. SIFT score is an important computational functional measurement predicting whether an amino acid substitution is deleterious [28]. SIFT values were calculated for splice-disrupt variants located on CDs. We categorized the variants based on the cutoff of 0.05 as: (1) Tolerable (SIFT score > 0.05) and (2) Damaging (SIFT score ≤ 0.05).
PolyPhen score
PolyPhen algorithm, like SIFT, use sequence homology of related proteins to evaluate whether an amino acid substitution can be deleterious to protein function based on the degree of conservation of the affected base throughout evolution. SIFT relies solely on sequence homology [29] while PolyPhen employs annotated UniProt entries to evaluate whether the amino acid substitution happens within an important structural or functional site of the protein, such as active or binding sites, and residues involved in disulphide bond formation [30, 31]. PolyPhen scores were classified in 3 levels as: (1) Benign (PolyPhen score ≤ 0.452), (2) Possibly Damaging (0.452 < PolyPhen score < 0.957), and Probably Damaging (PolyPhen score ≥ 0.957).
GERP++ conservation score
GERP is an evolutionary conservation score which have a good correspondence with clinical significance and pathogenicity level. GERP++ demonstrates the constrained elements in multiple alignments by quantifying substitution deficits. These deficits identify substitutions that would have happened if the element were neutral DNA, but did not happen as the element has been experienced functional constraint [32]. Low values of GERP++ score stand for low level of conservation and high values for high level of conservation.
Clinical significance
Pathogenic status of splice-disrupt variants was obtained from dbSNP database. dbSNP is the main global reference and repository of single nucleotide variations [22, 33, 34]. Assertions of clinical significance for alleles of human sequence variations are provided by the submitter at the time of submission of variant to NCBI, as: non-pathogenic, probable pathogenic, pathogenic, drug response, histocompatibility, untested, and unknown. Pathogenetic status is also supported by associated publications (citations) in dbSNP.
Functional annotation of splice-disrupt variants
Genes targeted by splice-disrupt variants in each type of prostate cancer (PC, CRPC, FPC, and MCRPC) were used as input for Gene ontology (GO) enrichment analysis in three terms of biological process (BP), molecular function (MF), and cellular component (CC). To this end, the following tools were employed: (1) Comparative GO [35, 36], and (2) STRING [37].
Literature-mining network of splice-disrupt variants, their annotated genes, and interactions with different types of prostate cancer
Literature mining more was used as a validation of in silico discovered splice-disrupt variants, to to investigate which of the computationally selected splice-disrupt variants have evidence of involvement in different types of prostate cancer.
Literature mining was performed using Pathway Studio Mammal database (Elsevier) [38], Version 12.4.0.3, as recently described [39]. Database contains functional relationships and pathways of mammalian proteins, covering human, mice, and rat. The database is enriched with protein-drug interaction and protein-disease interaction databases, called ChemEffect and DiseaseFx [40], respectively. The database is compiled using Medscan technology [41], a natural language processing engine, by text mining of over 24,000,000 PubMed abstracts and over 3,500,000 Elsevier and 3rd party full-text papers. The database is enriched with variation databases dbSNP v145 and dbNSFP v2.9, providing the opportunity to discover and visualise the relationships between genomic variants, genes, clinical parameters, diseases, and chemicals (small molecules).
Statistical analysis
Analysis of Variance (ANOVA) followed by Mean comparison using Tukey test was performed to compare the allele frequency of GERP++ conservation score between different types of prostate cancer. Two-sample proportion test was used to compare the clinical significance (occurrence of pathogenic status) between types of prostate cancer. Leven’s test was used to compare variance of allele frequency, and GERP++ between types of prostate cancer. Analysis was performed in MINITAB 18 (https://www.minitab.com). Graphs were visualized using GraphPad Prisim 7 (https://www.graphpad.com/).
Results
Mined splice-disrupt variants in prostate cancer (PC), castration-resistant prostate cancer (CRPC), familial prostate cancer (FPC), and metastatic castration-resistant prostate cancer (MCRPC)
Variant analysis resulted in identification of 854, 24, 112, and 35 splice-disrupt variants in PC, FPC, CRPC, and MCRPC. As presented in Supplementary 1, HLA-A in PC, MSR1 in FPC, and EGFR in both CRPC and MCRPC had the highest allele frequencies of splice-disrupt variations. Supplementary 1 presents list of variants in different types of prostate cancer based on whole genome sequencing.
Genomic locations of splice-disrupt variants
Genomic locations of splice-disrupt variants are presented at Fig. 1. As it can be inferred from Fig. 1, there is a remarkable difference in genomic locations of splice-disrupt variants in different types of prostate cancer. Splice-disrupt variants in MCRPC are only located on CDs and Intron. MCRPC has the highest percentage of CDs variants (16.21%) compared to the other types of prostate cancer. FPC has remarkably high enrichment of 5′UTR variants (7.4%). CRPC has the highest occurrence of 3′UTR splice-disrupt variants (6.5%) demonstrating the possible involvement of variants affecting the expression level and possible involvement of 3′UTR.
Identifying the high-risk splice-disrupt variants based on PolyPhen, SIFT, and GERP++ scores as well as clinical significance and their associated mechanisms
High-risk splice-disrupt variants were selected based on low SIFT score, high PolyPhen score and high GERP++ score as well as reported pathogenic clinical significance (retrieved from dbSNP database) (Supplementary 2–5). In more advanced type of prostate cancer (MCRPC), splice-disrupt variants, located on CDs of NCOR2, PTPRC, and CRP, are the high-risk variants. In CRPC, variants located on CDs of INSRR, MAEA, ESR1, TACC2, RB1, CRP as well as Intron based variants on BRCA1 are the high-risk ones. BRCA1, MLH1, MSR1, CYP1A1, CHEK2, and ELAC2 received the highest GERP++ score in FPC. INSRR, MDC1, WWOX, MAEA, FKBP5, TNFRSF10C, FAM13C, ESR1, CYP27A1, and BRCA1 are the top variants in PC. Figure 2 compares the overall GERP score in different types of PC where the highest score belongs to FPC documenting that this type of cancer has more simple genetic background, less affected by environmental conditions.
Highly pathogenic splice-disrupt genomic variants and their corresponding genes in different types of prostate cancer (PC, CRPC, FPC, and MCRPC are presented in Fig. 3. Clinical significance of human sequence variations was obtained from dbSNP. BRCA1 in all types of prostate cancer was the target of pathogenic splice-disrupt variants. Interestingly, CYP2D6 genes harbors splice-disrupt variants with high allele frequency.
Intersection of splice-disrupt variants in different types of prostate cancer
Common and specific splice-disrupt genomic variants between different types of prostate cancer (PC, CRPC, FPC, and MCRPC) is presented in Fig. 4. Splice-disrupt variants on BRCA1, AKR1C3, and KLK3 is observed in all types of prostate cancer. Splice-disrupt variants on CTSF and PTPRC is specific to MCRPC and may contribute to prostate cancer progression. FDFT1 solely happens in FPC. Splice-disrupt variants on NR5A2, HTRA2, AKR1C, ZWINT, and MST1 are linked to CRPC.
Overlapping splice-disrupt genomic variants and their target genes between different types of prostate cancer. A Splice-disrupt variants that are shared between different types of prostate cancer or are unique to a particular type of prostate cancer (specific splice-disrupt variants). B Genes that harbor splice-disrupt genomic variants in different types of prostate cancer. PC prostate cancer, CRPC castration-resistant prostate cancer, FPC familial prostate cancer, MCRPC metastatic castration-resistant prostate cancer. Total number of splice-disrupt genomic variants was 884 in PC, 24 in FPC, 112 in CRPC, and 35 in MCRPC
Progressive type of prostate cancer has higher diversity (variance) of GERP++ score in CDs
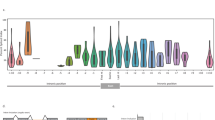
Variance analysis of CDs-located splice-disrupt variants showed that advanced type of prostate cancer, MCRPC, has remarkable higher variation in GERP++ score (Fig. 5). High diversity of GERP++ score in MCRPC demonstrates the more complex nature and diverse genomic background of MCRPC and highlights the importance of splice-disrupt variants in progressive prostate cancer.
Comparison of variation in GERP++ score of CDs (coding sequences)-located splice-disrupt variants in common and advanced types of prostate cancer. PC, CRPC, and MCRPC. Leven’s test was used for statistical comparison. FPC had only one splice-disrupt variants on CD. Consequently, it was not possible to calculate variance for FPC
Functional annotation of damaging splice-disrupt variants
Supplementary 6 presents functional annotation of high-risk splice-disrupt variants in different types of prostate cancer by enrichment analysis using GO database as well as KEGG pathways. According to KEGG pathway analysis, damaging splice-disrupt variants are significantly (p < 0.01) involved in Endocrine resistance and different types of cancer.
Lipid metabolic process, steroid metabolic process, regulation of mitotic cell cycle, negative regulation of metabolic process, negative regulation of signal transduction, and response to lipid are the key Biological Processes enriched by damaging splice-disrupt variants. One of the significant functions were negative regulation of production of miRNAs involved in gene silencing which variants on ESR1 are NCOR1 are involved in this function that can explain the link between UTR damaging variants and microRNA regulation.
Literature-mining based identification of splice-disrupt variants, their annotated genes, and interactions with different types of prostate cancer
Literature-mining based identification of variants, their annotated genes, and interactions with different types of prostate cancer is presented in Fig. 6. Mined references underpinning the network is presented in Supplementary 7. BRCA1 splice-disrupt variants are involved in different types of prostate cancer (Fig. 6). rs545982789 splice-disrupt on CHEK2 kinase is an important genetic change involved in FPC (Fig. 7). Literature mining based variant analysis highlighted the importance of rs1800716 variant, located on the CYP2D6 gene, involved in a range of important functions, such as RNA spicing, drug interaction, death, and urotoxicity.
Literature-mining based identification of splice-disrupt genomic variants, their annotated genes, and their interactions with different types of prostate cancer. A Literature mining based network of splice-disrupt variants and their target genes in PC. B Literature mining based network of splice-disrupt variants and their target genes in FPC. C Literature mining based network of splice-disrupt variants and their target genes in CRPC. D Literature mining based network of splice-disrupt variants and their target genes in MCRPC. Mined references underpinning the Literature-mining network is presented in Supplementary 7
rs1800716 splice-disrupt variant, located on the CYP2D6, is involved in a range of important functions, such as RNA spicing, drug interaction, death, and urotoxicity. A Interaction of rs1800716 with clinical parameters, disease, small molecules (chemical/drugs). B Gene interaction network of CYP2D6, the target gene of rs1800716 variant with the other genes. Mined references underpinning the literature-mining network of rs1800716 is presented in Supplementary 8
Discussion
Alternative splicing is one of the complexities of systems biology, particularly in cancer studies. Compared to the other type of splice variants, little is known about splice-disrupt genomic variants in cancer. Here, we developed a bioinformatic pipeline for extraction and detection of high-risk splice-disrupt genomic variants, by analysis of big data of deposited variants in prostate cancer, castration-resistant prostate cancer, familial prostate cancer, and metastatic castration-resistant prostate cancer. We showed that some splice-disrupt variants are solely present in the advanced metastatic castration-resistant prostate cancer. This is the first study that profiles the splice-disrupt genomic variants and their target genes in different types of prostate cancer. In final step, literature mining was used to uncover and visualise the relationships between splice-disrupt variants, genes, clinical parameters, diseases, and chemicals (small molecules), highlighting the importance of rs1800716 splice-disrupt variants, located on CYP2D6, in prostate cancer.
Noticeably, no cutoff was used for minor allele frequency of splice-disrupt variants in this study. This allowed us to investigate the contribution of less frequent (1–5%, minor allele frequency) and rare variants (< 1% minor allele frequency) to different types of prostate cancer. While GWAS commonly the cutoff of 5% for minor allele frequency, recent publications suggest the potential contributions of less variants to the risk of different diseases have been discussed [42, 43]. It is believed that most of the heterozygosity in the human genome comes from in variants with a minor allele frequency. The fact that majority of detected splice-disrupt genomic variants in this study were minor variants supports this statement (96.01% in PC, 95.83 in FPC, 94.64 in CRP, and 97.14% in MCRP, Supplementary 1).
Apoptosis and angiogenesis are the main cancer-associated processes, where alternative splicing plays a crucial role in their regulation [44]. Functional annotation of high-risk splice-disrupt variants showed that they are mainly involved in endocrine resistance, regulation of mitotic cell cycle, negative and response to lipid that are involved in apoptosis and angiogenesis. We observed the enrichment of negative regulation of production of miRNAs involved in gene silencing, where ESR1 are NCOR1 are involved in this function.
Metastasis is the cause of more than 90% of cancer-related deaths and is the most complex function of cancer cells [7]. Metastasis requires phenotypic plasticity that is centred around epithelial–mesenchymal transition (EMT) [45]. Alternative splicing of several genes is shown to be linked to EMT, such as RBFox2 and ESRP [46, 47]. Shapiro et al. identified the first alternative splicing signature for EMT and showed that the key drivers of EMT, such as cytoskeleton remodelling, regulation of cell–cell junction formation, and regulation of cell migration, all experience alternative splicing events [48]. Recently, it has been reported that overexpression of PTPRC is involved in cell adhesion, facilitating the tumour proliferation and lymph node metastasis in cervical cancer patients [49]. In another study, the inhibition of PTPRC reduced the rates of tumour growth and metastasis in vivo [50]. The role of PTPRC has also been noticed in colon cancer metastasis [51]. It has been suggested that PTPRC, as an adhesion molecule, is involved in the spread of the tumour and immortalisation of the tumour cells during malignancy [52]. In gastric cancer, it has been shown that CTSF is involved in the growth and apoptosis where CTSF knockdown promotes proliferation by inhibiting apoptosis [53]. It has been suggested in CTSF gene may function as a tumour suppressor with high potential therapeutic value [53]. To best of our knowledge, this is the first report of splice-disrupt variants on PTPRC and CTSF genes and their involvement in prostate cancer. We found that splice-disrupt variants on CTSF and PTPRC is specific to MCRPC and may contribute to prostate cancer progression.
This is the first study that profiles the splice-disrupt genomic variants and their target genes in prostate cancer. Noticeably, we found an association between specific splice-disrupt variants with advanced prostate cancer. The major limitation of this study was the remarkable difference between number of splice-disrupt genomic in different types of prostate cancer. The variants in FPC, CRPC, and MCRPC are more detrimental and harder to detect compared to early stage of PC. Alternative splicing contributes to a range of phenotypic traits of tumours as they progress and undergo metastasis and is a potential target for gene therapy [7, 11]. Unravelling alternative splicing opens a new avenue towards the establishment of new diagnostic and prognostic markers for prostate cancer progression and metastasis [48], as well as the development of a new generation of anticancer therapeutics: Treatments that inhibit specific splice variants, rather than targeting genes.
References
Pajares MJ, Ezponda T, Catena R, Calvo A, Pio R, Montuenga LM (2007) Alternative splicing: an emerging topic in molecular and clinical oncology. Lancet Oncol 8:349–357
Baharlou Houreh M, Ghorbani Kalkhajeh P, Niazi A, Ebrahimi F, Ebrahimie E (2018) SpliceDetector: a software for detection of alternative splicing events in human and model organisms directly from transcript IDs. Sci Rep 8:5063
Stamm S, Ben-Ari S, Rafalska I et al (2005) Function of alternative splicing. Gene 344:1–20
Venables JP, Klinck R, Koh C et al (2009) Cancer-associated regulation of alternative splicing. Nature Struct Mol Biol 16:670
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ (2008) Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet 40:1413
Wang ET, Sandberg R, Luo S et al (2008) Alternative isoform regulation in human tissue transcriptomes. Nature 456:470
Oltean S, Bates DO (2013) Hallmarks of alternative splicing in cancer. Oncogene 33:5311
Ebrahimie E, Rahimirad S, Tahsili M, Mohammadi-Dehcheshmeh M (2021) Alternative RNA splicing in stem cells and cancer stem cells: importance of transcript-based expression analysis. World J Stem Cells 13:1394
Enrique L-P, Manuel D, Alberto G, María Victoria G-G (2016) Neurogenesis: regulation by alternative splicing and related posttranscriptional processes. Neuroscientist 23:466–477
Anna A, Monika G (2018) Splicing mutations in human genetic disorders: examples, detection, and confirmation. J Appl Genet 59:253–268
Wang L, Duke L, Zhang PS et al (2003) Alternative splicing disrupts a nuclear localization signal in spleen tyrosine kinase that is required for invasion suppression in breast cancer. Cancer Res 63:4724–4730
Lastella P, Resta N, Miccolis I, Quagliarella A, Guanti G, Stella A (2004) Site directed mutagenesis of hMLH1 exonic splicing enhancers does not correlate with splicing disruption. J Med Genet 41:e72–e72
Agiannitopoulos K, Papadopoulou E, Tsaousis GN et al (2019) Characterization of the c. 793-1G> A splicing variant in CHEK2 gene as pathogenic: a case report. BMC Med Genet 20:131
Gorlov IP, Gorlova OY, Frazier ML, Amos CI (2003) Missense mutations in hMLH1 and hMSH2 are associated with exonic splicing enhancers. Am J Hum Genet 73:1157–1161
Hecker M, Rüge A, Putscher E et al (2019) Aberrant expression of alternative splicing variants in multiple sclerosis—a systematic review. Autoimmun Rev. https://doi.org/10.1016/j.autrev.2019.05.010
Rius R, Riley LG, Guo Y et al (2019) Cryptic intronic NBAS variant reveals the genetic basis of recurrent liver failure in a child. Mol Genet Metab 126:77–82
Al-Khawaga S, AlRayahi J, Saraswathi S et al (2019) A Novel SLC16A1 mutation in an infant with ketoacidosis and neuroimaging assessment: expanding the clinical spectrum of MCT1 deficiency. Front Pead 7:299
Raud L, Ka C, Gourlaouen I et al (2019) Functional analysis of novel RHD variants: splicing disruption is likely to be a common mechanism of variant D phenotype. Transfusion 59:1367–1375
Tavian D, Maggi L, Mora M, Morandi L, Bragato C, Missaglia S (2019) A novel PNPLA2 mutation causing total loss of RNA and protein expression in two NLSDM siblings with early onset but slowly progressive severe myopathy. Genes Dis. https://doi.org/10.1016/j.gendis.2019.07.006
Alanazi IO, Al Shehri ZS, Ebrahimie E, Giahi H, Mohammadi-Dehcheshmeh M (2019) Non‐coding and coding genomic variants distinguish prostate cancer, castration‐resistant prostate cancer, familial prostate cancer, and metastatic castration‐resistant prostate cancer from each other. Mol Carcinog 58:862–874
Consortium GP (2015) A global reference for human genetic variation. Nature 526:68
Sherry ST, Ward M-H, Kholodov M et al (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29:308–311
Liu X, Jian X, Boerwinkle E (2013) dbNSFP v2. 0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum Mutat 34:E2393–E2402
Liu X, Jian X, Boerwinkle E (2011) dbNSFP: a lightweight database of human nonsynonymous SNPs and their functional predictions. Hum Mutat 32:894–899
Liu X, Li C, Mou C, Dong Y, Tu Y (2020) dbNSFP v4: a comprehensive database of transcript-specific functional predictions and annotations for human nonsynonymous and splice-site SNVs. Genome Med 12:1–8
Sim N-L, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC (2012) SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res 40:W452–W457
Chan PA, Duraisamy S, Miller PJ et al (2007) Interpreting missense variants: comparing computational methods in human disease genes CDKN2A, MLH1, MSH2, MECP2, and tyrosinase (TYR). Hum Mutat 28:683–693
Vaser R, Adusumalli S, Leng SN, Sikic M, Ng PC (2016) SIFT missense predictions for genomes. Nat Protoc 11:1
Flanagan SE, Patch A-M, Ellard S (2010) Using SIFT and PolyPhen to predict loss-of-function and gain-of-function mutations. Genetic Test Mol biomarkers 14:533–537
Ramensky V, Bork P, Sunyaev S (2002) Human non-synonymous SNPs: server and survey. Nucleic Acids Res 30:3894–3900
Adzhubei I, Jordan DM, Sunyaev SR (2013) Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Human Genet 76:7.20.21-27.20.41
Davydov EV, Goode DL, Sirota M, Cooper GM, Sidow A, Batzoglou S (2010) Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput Biol 6:e1001025
Sherry ST, Ward M, Sirotkin K (1999) dbSNP—database for single nucleotide polymorphisms and other classes of minor genetic variation. Genome Res 9:677–679
Bhagwat M (2010) Searching NCBI’s dbSNP database. Curr Protoc Bioinf 32:1.19.11-11.19.18
Fruzangohar M, Ebrahimie E, Ogunniyi AD, Mahdi LK, Paton JC, Adelson DL (2013) Comparative GO: a web application for comparative gene ontology and gene ontology-based gene selection in bacteria. PLoS ONE 8:e58759
Fruzangohar M, Ebrahimie E, Adelson DL (2017) A novel hypothesis-unbiased method for gene ontology enrichment based on transcriptome data. PLoS ONE 12:e58759
Szklarczyk D, Gable AL, Lyon D et al (2018) STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613
Nikitin A, Egorov S, Daraselia N, Mazo I (2003) Pathway studio—the analysis and navigation of molecular networks. Bioinformatics 19:2155–2157
Mohammadi-Dehcheshmeh M, Moghbeli SM, Rahimirad S, Alanazi IO, Shehri ZSA, Ebrahimie E (2021) A transcription regulatory sequence in the 5′ untranslated region of SARS-CoV-2 is vital for virus replication with an altered evolutionary pattern against human inhibitory microRNAs. Cells 10:319
Yuryev A, Kotelnikova E, Daraselia N (2009) Ariadne’s chemeffect and pathway studio knowledge base. Expert Opin Drug Discov 4:1307–1318
Novichkova S, Egorov S, Daraselia N (2003) MedScan, a natural language processing engine for MEDLINE abstracts. Bioinformatics 19:1699–1706
Edenberg HJ, Foroud T (2014) Genetics of alcoholism. Handbook of clinical neurology. Elsevier, Amsterdam, pp 561–571
Bearden CE, Zandi P, Freimer NB (2016) Molecular architecture and neurobiology of bipolar disorder. Genomics, circuits, and pathways in clinical neuropsychiatry. Elsevier, Amsterdam, pp 467–486
Mercatante DR, Bortner CD, Cidlowski JA, Kole R (2001) Modification of alternative splicing of Bcl-x Pre-mRNA in prostate and breast cancer cells: analysis of apoptosis and cell death. J Biol Chem 276:16411–16417
Kalluri R, Weinberg RA (2009) The basics of epithelial-mesenchymal transition. J Clin Investig 119:1420–1428
Venables JP, Brosseau J-P, Gadea G et al (2013) RBFOX2 is an important regulator of mesenchymal tissue-specific splicing in both normal and cancer tissues. Mol Cell Biol 33:396–405
Warzecha CC, Jiang P, Amirikian K et al (2010) An ESRP-regulated splicing programme is abrogated during the epithelial–mesenchymal transition. EMBO J 29:3286–3300
Shapiro IM, Cheng AW, Flytzanis NC et al (2011) An EMT–driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype. PLoS Genet 7:e1002218
Mitra T, Elangovan S (2021) Cervical cancer development, chemoresistance, and therapy: a snapshot of involvement of microRNA. Mol Cell Biochem 476:4363–4385
Perron M, Saragovi HU (2018) Inhibition of CD45 phosphatase activity induces cell cycle arrest and apoptosis of CD45+ lymphoid tumors ex vivo and in vivo. Mol Pharmacol 93:575–580
Chu S, Wang H, Yu M (2017) A putative molecular network associated with colon cancer metastasis constructed from microarray data. World J Surg Oncol 15:1–9
Egan P, Drain S, Conway C, Bjourson AJ, Alexander HD (2016) Towards stratified medicine in plasma cell myeloma. Int J Mol Sci 17:1760
Ji C, Zhao Y, Kou Y-W et al (2018) Cathepsin F knockdown induces proliferation and inhibits apoptosis in gastric cancer cells. Oncol Res 26:83
Acknowledgements
This research was supported by use of the Nectar Research Cloud, a collaborative Australian research platform supported by the Australian Research Data Commons (ARDC) and National Collaborative Research Infrastructure Strategy (NCRIS). Additionally, this work was supported by resources provided by the Pawsey Supercomputing Centre with funding from the Australian Government and the Government of Western Australia.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by MM-D. The first draft of the manuscript was written by EE and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
11033_2022_7257_MOESM1_ESM.xlsx
Supplementary file 1 (DOCX 98 KB) List of splice-disrupt variantsin different types of prostate cancer. Prostate cancer (PC), castration-resistant prostate cancer (CRPC), and metastatic castration-resistant prostate cancer (MCRPC)
11033_2022_7257_MOESM2_ESM.docx
Supplementary file 2 (DOCX 25 KB) High-risk splice-disrupt variants in prostate cancer, based on PolyPhen, SIFT, GERP++ scores and reported clinical significance in dbSNP
11033_2022_7257_MOESM3_ESM.docx
Supplementary file 3 (DOCX 23 KB) High-risk splice-disrupt variants in familial prostate cancer (FPC) based on PolyPhen, SIFT, and GERP++ scores as well as reported clinical significance
11033_2022_7257_MOESM4_ESM.docx
Supplementary file 4 (DOCX 23 KB) High-risk splice-disrupt variants in castration-resistant prostate cancer (CRPC) based on PolyPhen, SIFT, and GERP++ scores as well as reported clinical significance
11033_2022_7257_MOESM5_ESM.docx
Supplementary file 5 (DOCX 23 KB) High-risk splice-disrupt variants in metastatic castration-resistant prostate cancer (MCRPC) based on PolyPhen, SIFT, and GERP++ scores as well as reported clinical significance
11033_2022_7257_MOESM7_ESM.xlsx
Supplementary file 7 (XLSX 53 KB) Mined references underpinning literature-mining network of splice-disrupt variants, their annotated genes, and interactions with different types of prostate cancer. The network of relationships/interaction is visualised in Figure 6)
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Alanazi, I.O., Alamery, S.F., Ebrahimie, E. et al. Splice-disrupt genomic variants in prostate cancer. Mol Biol Rep 49, 4237–4246 (2022). https://doi.org/10.1007/s11033-022-07257-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07257-9