Abstract
CC chemokine receptor type 7 (CCR7) and its ligands has been implicated in the occurrence and progression of NSCLC. Previous studies have revealed that the diagnostic value of CCR7/CCL19 axis in lung tumorigenesis remains controversial. The present study evaluates the relationship between the mRNA expression of CCR7/CCL19 axis and selected regulatory miRNAs in NSCLC patients. It analyzes the expression level of CCR7 mRNA and its ligand in tumor tissue in relation to expression level of two miRNAs: miR let-7a and miR-335, as transcriptional regulators of study genes. Twenty-seven patients (n = 27) were enrolled. The expression of the studied genes and miRNAs was evaluated by qPCR. Tumour tissue fragments, adjacent macroscopically-unchanged lung tissue (control) and patient serum were used as biological material for study. Elevated expression of CCR7 and CCL19 mRNA was observed in patients with metastasis to lymph nodes. We noticed upregulated miR-335 expression and downregulated miR let-7a expression in patient serum with regard to AJCC tumor staging. Higher miR-335 expression and lower miR let-7a expression level was observed in patients with metastasis to lymph node. The presence of changes observed in the expression level of miR-335 and miR let-7a in the serum of NSCLC patients in relation to lymph node metastases and tumor stage may serve as a non-invasive molecular biomarker of tumor progression; however, this observation requires further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is a considerable global problem. It is the most commonly-diagnosed form of cancer in men and is the leading cause of cancer-related death of most patients with NSCLC diagnoses.
The molecular mechanisms responsible for the development and progression of NSCLC are not fully understand for the formation of tumor metastasis; however, it is known that Epithelial Mesenchymal Transition (ETM) and chemotaxis play pivotal roles in lung cancer metastasis. Cancer cell migration and metastasis are regulated by the chemokine system and their receptors. It was recently recognized that the interaction between C–C motif chemokine ligand 19 (CCL19) and G protein-coupled CC chemokine receptor 7 (CCR7) plays a key role in the activation of native T cells, B cells and dendritic cells, and the migration of these cells within the lymphoid organs [1]. A number of recent studies suggest that the CCL19/CCR7 axis may also be utilized to promote local tumor growth and metastasis [2, 3]. It has been shown that CCR7 expression level increases in many type of cancers, including lung cancer, and that the expression of CCR7 significantly correlates with lymph node metastases [2].
In addition, many recent studies have examined the roles of microRNAs (miRNAs) in cell development, differentiation and migration as key factors in the regulation of human carcinogenesis. MiRNAs are small, non-coding, single-stranded RNAs that can regulate about 30% of human genes, including these responsible for tumorigenesis and metastasis [4]. Therefore, it is proposed that miRNAs may act as potential molecular biomarkers for lung cancer, among others [5].
The present study aims to evaluate changes in the expression level of the CCR7 and CCL19 genes and miR let-7a and miR-335, two of the miRNAs known to regulate the expression of this genes according database MirTarBased. This miRNAs may act as non-invasive tools for identifying a risk for metastasis to lymph nodes assessed in serum collected from patients with NSCLC. It also analyses the differences in CCR7, CCL19, miR let-7a and miR-335 expression with the clinical features of patients with various stages of cancer according to the AJCC/TNM staging system.
Future prognosis of the disease occurrence and treatment outcomes should be based on combination of AJCC/TNM staging system and molecular tumor profiling. This approach may allow more individualization of treatment algorithms.
Materials and methods
The study was approved by the Bioethics Committee of the Medical University of Lodz, Poland (resolution No. KE/149/18). Written informed consent to participate was obtained from each patient.
Characterization of patients
The study cohort consists of 27 patients with diagnosed NSCLC, 11 women and 16 men, aged 51 to 81 (mean age 66.96 ± 7.95 years). Lung resection (pulmonectomy or lobectomy) in patients was performed at the Department of Thoracic Surgery, General and Oncologic Surgery, Military Medical Academy Memorial Teaching Hospital of The Medical University of Lodz—Central Veterans’ Hospital, Lodz, Poland during the years 2018–2019. The clinical features of the patients with diagnosed NSCLC are summarized in Table 1.
Tissue collection
Primary tumor tissue fragments from NSCLC patients were obtained from the center of the lesion (approximately 100 mg). The adjacent macroscopically-unchanged tissue (approximately 100 mg taken at a distance of 2 cm from the primary focus) was used as a control. The lung tissue was collected in fixRNA buffer (EURx, Gdansk, Poland), homogenized, frozen at − 80 °C and stored until use at the Department of Biomedicine and Genetics, Medical University of Lodz. None of the patients were treated with chemo- or radiotherapy.
The tissue samples were subjected to postoperative histopathological evaluation and classified according to AJCC and TNM staging system (pTNM) [6]. The histopathological and clinical assessment of biological materials is shown in Table 2.
Serum collection
Serum from all (n = 27) study patients with diagnosed NSCLC was collected before the surgery. Serum from 21 patients (volunteers) was obtained as a control. All samples of serum was stored at − 20 °C before use.
RNA isolation from tissue and serum, qualitative and quantitative RNA evaluation
Isolation of total RNA from tissue homogenates was performed using the mirVana™ miRNA Isolation Kit (Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocol. Exosomes from the serum were extracted using the Total Exosome Isolation Reagent. Total Exosome RNA was extracted from exosomes & Protein Isolation Kit (Applied Biosystems, USA), according to the manufacturer’s instructions.
Qualitative and quantitative assessment of the RNA was performed by spectrophotometric method (260/280 nm) using an Eppendorf BioPhotometerTM Plus apparatus (Eppendorf, Hamburg, Germany).
Evaluation of relative gene expression (RQ)
Reverse transcription (RT) for genes was performed using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, USA) in a Personal Thermocycler (Eppendorf, Germany), according to the manufacturer’s instructions.
The relative expression analysis for study genes was performed in 7900HT Fast Real-Time PCR System (Applied Biosystems, Carlsbad, CA, USA) using TaqMan assay for the genes CCR7 (Hs01013469_m1) CCL19 (Hs00171149_m1) and ACTB (Hs99999903_m1) used as an endogenous control.
The relative expression level of the study genes was evaluated by the delta–delta CT method (TaqMan Relative Quantification Assay software, Applied Biosystems). RNA from normal lung tissue (Human Lung Total RNA, Ambion®, USA) was used as calibrator.
Evaluation of relative miRNA expression level (RQ)
The reverse transcription (RT) for miRNA was carried out using a TaqMan® MicroRNA Reverse Transcription Kit (Applied Biosystems, Carlsbad, CA) according to the manufacturer’s protocol. RT reaction was performed in a Personal Thermocycler (Eppendorf, Germany) For miRNAs we used following assays: miR-335 (UCAAGAGCAAUAACGAAAAAUGU) and miR let-7a (UGAGGUAGUAGGUUGUAUAGUU). Global normalization was used to assess the relative RQ value of the studied miRNAs. The median CT of those assays is used as the normalizer, on a per sample basis. The level of calibrator expression (RNA isolated from healthy patient) was regarded as RQ = 1.
Statistical analysis
The RQ values for the study genes/miRNAs were presented as median values. The Mann–Whitney U test and Kruskal–Willis test were performed. The accepted level of statistical significance was estimated at p < 0.05. All statistical analyses were performed using Statistica for Windows 10.0 software.
Results and discussion
Despite progress in the diagnosis of lung cancer, the mortality rate of patients still remains high, with a rate of 26% being associated with cancer of the lung and bronchus [7]. Therefore, it is important to identify new diagnostic methods based on molecular tools and clarify the molecular background of carcinogenesis.
Chemokines are key factors in immunological homeostasis [8, 9] but also serve as oncogenic activators and/or inhibitors of tumor suppressor gene pathways, and their activities often result in epithelial mesenchymal-transition (EMT), tumor growth, metastasis [10]. However, the mechanisms behind this relationship appear to be complex and poorly understood.
It has been recognized that in a similar way to its two key ligands, CCL19 and CCL21, CCR7 may influence tumor cell adherence to the endothelium, blood vessel extravasation, angiogenesis, proliferation and immune response evasion [11, 12]. Lately, it has been documented that lung cancer cells possess CCR7 on their surface [2], and CCR7 mRNA overexpression in lung tumors has been associated with cancer progression [13].
As so far only a few studies have been performed in this area [2, 13], the present study assessed the expression of CCL19/CCR7 mRNA in non-small cell lung cancer tissue samples.
Our findings confirm the presence of CCR7 mRNA expression in tumor tissue. CCR7 mRNA was found to be upregulated (RQ > 1) in 96% cancer cells and all macroscopically-unchanged lung tissue surrounding the center of the tumor, with the higher expression of CCR7 mRNA in control tissue (median RQ values: 9.135 and 6.816 respectively) (p > 0.05). However, other studies have confirmed upregulation of CCR7 mRNA in both cancer tissue and macroscopically-unchanged lung tissue, but with significantly higher CCR7 mRNA expression observed in the cancer cells [13]. This observation may be related to the molecular heterogeneity of the cancer cells and may be associated with presence of early molecular changes of CCR7 mRNA expression in the morphologically-unchanged lung tissue surrounding the tumor. This area is recognized as the molecular margin of tumor [14]. It has been suggested that molecular changes observed in this area might be followed by a future molecular transformation into a malignant tumor. Thus, the presence of a molecular tumor margin—as a negative histological margin—confirmed the need to employ histological and molecular validation in diagnostics.
This is the first study to examine CCR7 mRNA expression in NSCLC patients in relation to smoking history, which revealed higher expression level in patients with a smoking status ≥ 40PYs (see Table 3). Previous studies focused on exposure of patient with chronic obstructive pulmonary disease to cigarette smoke have suggested that cigarette smoke may have an influence on CCR7 expression [15].
Regarding AJCC cancer stage, the highest expression of CCR7 mRNA was observed in patients with stage III cancer and this elevated expression level positively correlated with the presence of histologically confirmed lymph node metastasis: higher expression was noted in patients with N1 + N2 feature (see Table 4). It is consistent with hypothesis that the activation of CCR7 contributes to the increased potential for lymphatic metastasis of tumor and recurrence after surgery [3, 16]. Furthermore, the highest CCR7 mRNA expression was observed in patients with T3 (see Table 4). It is possible that CCR7 mRNA expression in patient with NSCLC, may be considered as diagnostic molecular biomarker with poor prognostic value. However, in study of others, it has been found that a better prognosis was noticed for patients with diagnosed adenocarcinoma who demonstrated higher CCR7 mRNA expression [17]. Interestingly, while no statistically significant differences in CCR7 mRNA expression were found between histopathological type (SCC vs. AC) in the present study, higher CCR7 mRNA expression was found in the SCC group (see Table 4). This observation is consistent with those of Liu et al. [13] who also report higher CCR7 mRNA expression in SCC histological type; however, this observation is in contrast to those of other authors who consider that AC histological type is more aggressive [18].
Currently it is known that CCL19 and CCL21 are able to control the chemokine system and influence the local tumor microenvironment, tumor growth and distant metastasis [19, 20].
Our study revealed downregulated (RQ < 1) expression of CCL19 mRNA in both NSCLC tissue (67%) and macroscopically-unchanged lung tissue (85%); however, significantly higher CCL19 mRNA expression was observed in the tumor tissue (median RQ values: 0.700 and 0.179 respectively, p = 0.011). Our observation was consistent with those of Liu et al. [13]. Our study regarding NSCLC histopathological subtypes, revealed insignificantly higher expression of CCL19 mRNA in SCC compared to AC (see Table 4); however, Liu et al. reported significant higher expression of CCL19 mRNA in the AC subtype than the SCC.
Additionally, recent studies have found high expression of CCL19 mRNA in tumor tissue to be a good prognostic factor in NSCLC (specially in adenocarcinoma) [17]. The study, conducted in a lung adenocarcinoma model, showed that high expression of CCL19 mRNA promotes IFN-γ-dependent antitumor responses [21].
In the present study no statistically significant differences were observed in CCL19 mRNA expression among patients divided according to AJCC or the pTNM staging system. However, the greatest expression was noticed in a patient with stage III cancer according to AJCC and in a case of T3 according to pTNM (see Table 4, Fig. 1). Liu et al. demonstrated that the expression of CCL19 mRNA was positively correlated with tumor staging but negatively with lymph node metastasis [13], but our findings indicate higher expression of CCL19 mRNA in patients with lymph node metastasis (see Table 4).
Emerging data highlights the role of miRNAs in a variety of biological processes within the tumor cell, including proliferation, differentiation, invasion and tumor cell migration [22]. Abnormal expression (up- or downregulation) of different classes of miRNAs has been documented in lung cancer in relation to various cancer-related genes [23, 24]. Many of the studied miRNAs have been confirmed as diagnostic and/or prognostic biomarkers in NSCLC development and progression [25, 26].
The present study, as few as before, examined the expression of miR let-7a and miR-335 in exosomes from the serum of study patients. Both were found to regulate the studied genes: CCR7 and/or CCL19 according the MirTarBase program.
MiR let-7a is a member of the let-7 miRNA family, which is believed to act as a tumor suppressor by inhibiting the expression of tumor-promoting genes [24]. Our study revealed a downregulation of miR let-7a expression in 78% samples from NSCLC patients (serum exosomes) and lower expression in the patient group compared to controls (median RQ values: 0.820 and 0.908 respectively, p > 0.05). Our results are consistent with previous observations, which also note reduced miR let-7a expression in the blood of NSCLC patients, as well as in tumor cells [27, 28].
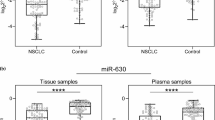
Our findings reveal a negative correlation between miR let-7a expression and cancer stage according to AJCC staging, and a positive correlation with tumor size (see Table 5). Additionally, a significant downregulation of miR let-7a expression were observed in patients with metastasis to lymph nodes compared to those without (Table 5, Fig. 2a). In contrast, Song et al. note a negative correlation between miR let-7a expression and tumor size (TNM staging system), however, this was found in laryngeal carcinoma. In addition, they also report lower expression of miR let-7a in patients with metastasis to lymph nodes [29].
Interestingly, our present findings indicate lower miR let-7a expression in serum in younger patients (≤ 65 age) and in smokers with smoking status ≥ 40 Pys (see Table 6). Previous studies indicated downregulation of miR let-7a in human serum or sputum from smokers [30, 31]. Another study has reported that miR let-7a downregulation could be reversible in smoking-related disease but irreversible in cancer [32].
The expression of miR-335 has been shown to be aberrant in tumor tissue from various types of cancer, and to be associated with cell proliferation and inflammation [33, 34]. Our present findings indicate upregulation of miR-335 (RQ > 1) in 78% NSCLC patients. The expression of miR-335 was found to be elevated in the serum of NSCLC patients compared to control group (median RQ values: 1.220 and 1.102 respectively, p > 0.05,): This may be the first assessment of miR-335 expression performed in the blood—as a marker in a liquid biopsy—in patients with NSCLC. However, Tang et al. report lower expression level of miR-335-5p in lung cancer tissues compared to noncancerous tissues [34]. In another study increased expression of miR-335, associated with hyper proliferation and progression of transformation, has also been documented in lung cancer [35].
Regarding the biological features of study patients, higher miR-335 expression was observed in female subjects and people aged < 65 (see Table 6). In smokers with history ≥ 40 PYs, higher expression of miR-335 was also noticed (see Table 6). Tang et al. also did not observe statistically significant differences in miR-335 expression level with regard to patient smoking status [34].
Increased miR-335 expression was observed with regard to AJCC staging; in addition, significant differences in miR-335 expression were found in the serum of patients with NSCLC, depending on presence of metastases. Higher miR-335 expression was observed in patients with lymph node metastases than in those without (see Table 5, Fig. 2b). In contrast, Tang et al. report significantly lower miR-335 expression in patients with metastases [34]. These conflicting results suggest that the expression of miR-335 may have prognostic value in patients with NSCLC, but further research should be carried out.
It has been suggested that the expression of CCR7 mRNA and its receptor ligands may be subjected to functional regulation by miRNA. The present study, assessing the relationship between the expression of CCR7/CCL19 mRNA and miR let-7a and miR-335. No reports have so far examined the level of CCR7/CCL19 expression in relation to miR let-7a or miR-335 in lung cancer, although similar studies carried out in breast cancer have observed a negative correlation between CCR7 expression and miR let-7a [36].
To conclude, the variations observed in the expression level of miR-335 and miR let-7a in serum of patients with NSCLC with regard to lymph node metastases and tumor size may potentially serve as non-invasive molecular biomarkers of tumor progression; however, this observation requires further investigation on larger group of samples.
References
Förster R, Schubel A, Breitfeld D, Kremmer E, Renner-Müller I, Wolf E, Lipp M (1999) CCR1 coordinates the primary immune response by establishing functional microenvironments in secondary lymphoid organs. Cell 99(1):23–33
Takanami I (2003) Overexpression of CCR2 MRNA in nonsmall cell lung cancer: correlation with lymph node metastasis. Int J Cancer 105(2):186–189. https://doi.org/10.1002/ijc.11063
Ding Y, Shimada Y, Maeda M, Kawabe A, Kaganoi J, Komoto I, Hashimoto Y, Miyake M, Hashida H, Imamura M (2003) Association of CC chemokine receptor 7 with lymph node metastasis of esophageal squamous cell carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res 9(9):3406–3412
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5(7):522–531. https://doi.org/10.1038/nrg1379
Qin X, Xu H, Gong W, Deng W (2014) The tumor cytosol MiRNAs, fluid MiRNAs, and exosome MiRNAs in lung cancer. Front Oncol 4:357. https://doi.org/10.3389/fonc.2014.00357
Mirsadraee S, Oswal D, Alizadeh Y, Caulo A, van Beek EJ (2012) The 7th lung cancer TNM classification and staging system: review of the changes and implications. World J Radiol 4(4):128–134. https://doi.org/10.4329/wjr.v4.i4.128
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA 68(1):7–30. https://doi.org/10.3322/caac.21442
Sarvaiya PJ, Guo D, Ulasov I, Gabikian P, Lesniak MS (2013) Chemokines in tumor progression and metastasis. Oncotarget 4(12):2171–2185. https://doi.org/10.18632/oncotarget.1426
Cheng Z-H, Shi Y-X, Yuan M, Xiong D, Zheng J-H, Zhang Z-Y (2016) Chemokines and their receptors in lung cancer progression and metastasis. J Zhejiang Univ Sci B 17(5):342–351. https://doi.org/10.1631/jzus.B1500258
Mantovani A (2010) Molecular pathways linking inflammation and cancer. Curr Mol Med 10(4):369–373
Kakinuma T, Hwang ST (2006) Chemokines, chemokine receptors, and cancer metastasis. J Leukoc Biol 79(4):639–651. https://doi.org/10.1189/jlb.1105633
Zlotnik A, Burkhardt AM, Homey B (2011) Homeostatic chemokine receptors and organ-specific metastasis. Nat Rev Immunol 11(9):597–606. https://doi.org/10.1038/nri3049
Liu Y, Wu B-Q, Geng H, Xu M-L, Zhong H-H (2015) Association of chemokine and chemokine receptor expression with the invasion and metastasis of lung carcinoma. Oncol Lett 10(3):1315–1322. https://doi.org/10.3892/ol.2015.3402
Kalinowska-Herok M, Roś M, Widłak P (2013) Tumor molecular margins. Nowotw J Oncol 63(1):28–34
Liao SX, Ding T, Rao XM, Sun DS, Sun PP, Wang YJ, Fu DD, Liu XL, Ou-Yang Y (2015) Cigarette smoke affects dendritic cell maturation in the small airways of patients with chronic obstructive pulmonary disease. Mol Med Rep. 11(1):219–225. https://doi.org/10.3892/mmr.2014.2759
Suzuki K, Kachala SS, Kadota K, Shen R, Mo Q, Beer DG, Rusch VW, Travis WD, Adusumilli PS (2011) Prognostic immune markers in non-small cell lung cancer. Clin Cancer Res Off J Am Assoc Cancer Res 17(16):5247–5256. https://doi.org/10.1158/1078-0432.CCR-10-2805
Itakura M, Terashima Y, Shingyoji M, Yokoi S, Ohira M, Kageyama H, Matui Y, Yoshida Y, Ashinuma H, Moriya Y et al (2013) High CC chemokine receptor 7 expression improves postoperative prognosis of lung adenocarcinoma patients. Br J Cancer 109(5):1100–1108. https://doi.org/10.1038/bjc.2013.440
Bieniasz M, Bartkowiak J, Szemraj J (2008) Ekspresja genów kodujących białka morfogenetyczne kości, metaloproteinazy i inhibitory metaloproteinaz w nowotworach płuc i ich znaczenie rokownicze. Post Biochem 54(1):82–90
Keeley EC, Mehrad B, Strieter RM (2010) CXC chemokines in cancer angiogenesis and metastases. Adv Cancer Res 106:91–111. https://doi.org/10.1016/S0065-230X(10)06003-3
Singh R, Lillard JW, Singh S (2011) Chemokines: key players in cancer progression and metastasis. Front Biosci 3:1569–1582
Hillinger S, Yang S-C, Zhu L, Huang M, Duckett R, Atianzar K, Batra RK, Strieter RM, Dubinett SM, Sharma S (2003) EBV-induced molecule 1 ligand chemokine (ELC/CCL19) promotes IFN-gamma-dependent antitumor responses in a lung cancer model. J Immunol 171(12):6457–6465
Świtlik WZ, Szemraj J (2017) Krążące MiRNA Jako Nieinwazyjne biomarkery diagnostyczne, prognostyczne oraz predykcyjne w Terapii Niedrobnokomórkwego Raka Płuca. Adv Hyg Exp Med 71:649–661
Lu J, Zhan Y, Feng J, Luo J, Fan S (2018) MicroRNAs associated with therapy of non-small cell lung cancer. Int J Biol Sci 14(4):390–397. https://doi.org/10.7150/ijbs.22243
Chin LJ, Ratner E, Leng S, Zhai R, Nallur S, Babar I, Muller R-U, Straka E, Su L, Burki EA et al (2008) A SNP in a let-7 MicroRNA complementary site in the KRAS 3′ untranslated region increases non-small cell lung cancer risk. Cancer Res 68(20):8535–8540. https://doi.org/10.1158/0008-5472.CAN-08-2129
Chen X, Hu Z, Wang W, Ba Y, Ma L, Zhang C, Wang C, Ren Z, Zhao Y, Wu S et al (2012) Identification of ten serum MicroRNAs from a genome-wide serum MicroRNA expression profile as novel noninvasive biomarkers for nonsmall cell lung cancer diagnosis. Int J Cancer 130(7):1620–1628. https://doi.org/10.1002/ijc.26177
Chen L, Jin H (2014) MicroRNAs as novel biomarkers in the diagnosis of non-small cell lung cancer: a meta-analysis based on 20 studies. Tumour Biol 35(9):9119–9129. https://doi.org/10.1007/s13277-014-2188-2
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, Harano T, Yatabe Y, Nagino M, Nimura Y et al (2004) Reduced expression of the Let-7 MicroRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64(11):3753–3756. https://doi.org/10.1158/0008-5472.CAN-04-0637
Jeong HC, Kim EK, Lee JH, Lee JM, Yoo HN, Kim JK (2011) Aberrant expression of Let-7a MiRNA in the blood of non-small cell lung cancer patients. Mol Med Rep 4(2):383–387. https://doi.org/10.3892/mmr.2011.430
Song F-C, Yang Y, Liu J-X (2016) Expression and significances of MiRNA Let-7 and HMGA2 in laryngeal carcinoma. Eur Rev Med Pharmacol Sci 20(21):4452–4458
Rizk SA, Metwally FM, Elfiky AM, Mahmoud AA, Badawi NA, Sharaf NE, Elhefnawi MM (2018) Down-regulation of circulating MicroRNA Let-7a in Egyptian smokers. J Genet Eng Biotechnol 16(1):53–56. https://doi.org/10.1016/j.jgeb.2018.02.003
Huang J, Wu J, Li Y, Li X, Yang T, Yang Q, Jiang Y (2014) Deregulation of serum MicroRNA expression is associated with cigarette smoking and lung cancer. Biomed Res Int 2014:364316. https://doi.org/10.1155/2014/364316
Izzotti A, Calin GA, Steele VE, Croce CM, De Flora S (2009) Relationships of MicroRNA expression in mouse lung with age and exposure to cigarette smoke and light. FASEB J 23(9):3243–3250. https://doi.org/10.1096/fj.09-135251
Wang Y, Wang N, Zeng X, Sun J, Wang G, Xu H, Zhao W (2017) MicroRNA-335 and its target Rock1 synergistically influence tumor progression and prognosis in osteosarcoma. Oncol Lett 13(5):3057–3065. https://doi.org/10.3892/ol.2017.5818
Tang H, Zhu J, Du W, Liu S, Zeng Y, Ding Z, Zhang Y, Wang X, Liu Z, Huang J (2018) CPNE1 is a target of MiR-335-5p and plays an important role in the pathogenesis of non-small cell lung cancer. J Exp Clin Cancer Res 37(1):131. https://doi.org/10.1186/s13046-018-0811-6
Scarola M, Schoeftner S, Schneider C, Benetti R (2010) MiR-335 directly targets Rb1 (PRb/P105) in a proximal connection to P53-dependent stress response. Cancer Res 70(17):6925–6933. https://doi.org/10.1158/0008-5472.CAN-10-0141
Kim S-J, Shin J-Y, Lee K-D, Bae Y-K, Sung KW, Nam SJ, Chun K-H (2012) MicroRNA Let-7a suppresses breast cancer cell migration and invasion through downregulation of C-C chemokine receptor type 7. Breast Cancer Res 14(1):R14. https://doi.org/10.1186/bcr3098
Funding
This study was supported by the Medical University of Lodz (Statute No. 503/1-013-02/503-11-003). The founder had no role in study design data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Baran, K., Kiszałkiewicz, J., Migdalska-Sęk, M. et al. An assessment of the relationship between the expression of CCR7/CCL19 axis and selected regulatory miRNAs in non-small cell lung cancer. Mol Biol Rep 46, 5389–5396 (2019). https://doi.org/10.1007/s11033-019-04993-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04993-3