Abstract
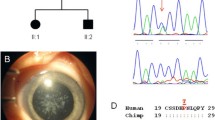
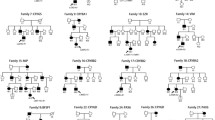
A bilaterally blind woman, with a three generation family history of autosomal dominant congenital cataracts, variably associated with iris colobomata and microcornea, sought preconception genetic consultation. Whole-exome sequencing was performed in three affected family members, one unaffected first degree relative, and one spouse. The sequence variant c.168C>G; p.(Tyr56∗) in CRYGD, previously reported as pathogenic, and a novel mutation c.809C>A; p.(Ser270Tyr) in MAF, were identified in two affected family members; the grandmother, and half-brother of the proband. The proband inherited only the MAF mutation, whereas her clinically unaffected sister had the CRYGD change. In silico analysis supported a pathogenic role of p.(Ser270Tyr) in MAF, which was absent from publicly available whole-exome datasets, and 1161 Czech individuals. The frequency of CRYGD p.(Tyr56∗) in the ExAC dataset was higher than the estimated incidence of congenital cataract in the general population. Our study highlights that patients with genetically heterogeneous conditions may exhibit rare variants in more than one disease-associated gene, warranting caution with data interpretation, and supporting parallel screening of all genes known to harbour pathogenic mutations for a given phenotype. The pathogenicity of sequence variants previously reported as cataract-causing may require re-assessment in light of recently released datasets of human genomic variation.

Similar content being viewed by others
Change history
10 October 2017
There was a spacing error in the initial online publication, and there were errors in the Acknowledgments section. The original article has been updated.
References
Lambert SR, Drack AV (1996) Infantile cataracts. Surv Ophthalmol 40:427–458
Reddy MA, Francis PJ, Berry V, Bhattacharya SS, Moore AT (2004) Molecular genetic basis of inherited cataract and associated phenotypes. Surv Ophthalmol 49:300–315. doi: 10.1016/j.survophthal.2004.02.013
Moore AT (2004) Understanding the moleculas genetics of congenital cataract may have wider implications for age related cataract. Br J Ophthalmol 88:2–3
Shiels A, Bennett TM, Hejtmancik JF (2010) MolVis 16:2007–2015
Ma AS, Grigg JR, Ho G et al (2016) Sporadic and familial congenital cataracts: mutational spectrum and new diagnoses using next-generation sequencing. Hum Mutat 37:371–384. doi: 10.1002/humu.22948
Churchill A, Graw J (2011) Clinical and experimental advances in congenital and paediatric cataracts. Philos Trans R Soc Lond B Biol Sci 366:1234–1249. doi: 10.1098/rstb.2010.0227
Hoehenwarter W, Klose J, Jungblut PR (2006) Eye lens proteomics. Amino Acids 30:369–389. doi: 10.1007/s00726-005-0283-9
Brakenhoff RH, Aarts HJ, Reek FH, Lubsen NH, Schoenmakers JG (1990) Human gamma-crystallin genes. A gene family on its way to extinction. J Mol Biol 216:519–532
Kmoch S, Brynda J, Asfaw B et al (2000) Link between a novel human gammad-crystallin allele and a unique cataract phenotype explained by protein crystallography. Hum Mol Genet 9:1779–1786
Stephan DA, Gillanders E, Vanderveen D et al (1999) Progressive juvenile-onset punctate cataracts caused by mutation of the gammad-crystallin gene. Proc Natl Acad Sci USA 96:1008–1012
Hejtmancik JF (2008) Congenital cataracts and their molecular genetics. Semin Cell Dev Biol 19:134–149. doi: 10.1016/j.semcdb.2007.10.003
Motohashi H, O’Connor T, Katsuoka F, Engel JD, Yamamoto M (2002) Integration and diversity of the regulatory network composed of Maf and CNC families of transcription factors. Gene 294:1–12
Kannan MB, Solovieva V, Blank V (2012) The small MAF transcription factors MAFF, MAFG and MAFK: current knowledge and perspectives. Biochim Biophys Acta 1823:1841–1846. doi: 10.1016/j.bbamcr.2012.06.012
Civil A, van Genesen ST, Lubsen NH (2002) c-Maf, the gammad-crystallin Maf-responsive element and growth factor regulation. Nucleic Acids Res 30:975–982
Jamieson RV, Munier F, Balmer A, Farrar N, Perveen R, Black GC (2003) Pulverulent cataract with variably associated microcornea and iris coloboma in a MAF mutation family. Br J Ophthalmol 87:411–412
Narumi Y, Nishina S, Tokimitsu M et al (2014) Identification of a novel missense mutation of MAF in a Japanese family with congenital cataract by whole exome sequencing: a clinical report and review of literature. Am J Med Genet A 164A:1272–1276. doi: 10.1002/ajmg.a.36433
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26:589–595. doi: 10.1093/bioinformatics/btp698
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. doi: 10.1093/bioinformatics/btp352
Shiels A, Bennett TM, Hejtmancik JF (2010) Cat-Map: putting cataract on the map. Mol Vis 16:2007–2015
Ng PC, Henikoff S (2001) Predicting deleterious amino acid substitutions. Genome Res 11:863–874. doi: 10.1101/gr.176601
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249. doi: 10.1038/nmeth0410-248
Li B, Krishnan VG, Mort ME et al (2009) Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25:2744–2750. doi: 10.1093/bioinformatics/btp528
Choi Y, Sims GE, Murphy S, Miller JR, Chan AP (2012) Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 7:e46688. doi: 10.1371/journal.pone.0046688
Schwarz JM, Rodelsperger C, Schuelke M, Seelow D (2010) MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 7:575–576. doi: 10.1038/nmeth0810-575
Calabrese R, Capriotti E, Fariselli P, Martelli PL, Casadio R (2009) Functional annotations improve the predictive score of human disease-related mutations in proteins. Hum Mutat 30:1237–1244. doi: 10.1002/humu.21047
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217. doi: 10.1006/jmbi.2000.4042
den Dunnen JT, Antonarakis SE (2000) Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat 15:7–12. doi: 10.1002/(SICI)1098-1004(200001)15:1
Santana A, Waiswol M, Arcieri ES, Cabral de Vasconcellos JP, Barbosa de Melo M (2009) Mutation analysis of CRYAA. CRYGC, and CRYGD associated with autosomal dominant congenital cataract in Brazilian families. Mol Vis 15:793–800
Vanita V, Guo G, Singh D, Ott CE, Robinson PN (2014) Differential effect of cataract-associated mutations in MAF on transactivation of MAF target genes. Mol Cell Biochem 396:137–145. doi: 10.1007/s11010-014-2150-z
Hansen L, Eiberg H, Rosenberg T (2007) Novel MAF mutation in a family with congenital cataract-microcornea syndrome. Mol Vis 13:2019–2022
Acknowledgements
This work was supported by Charles University institutional programs UNCE 204011 and PROGRES-Q26/LF1. Specific support was provided by Grants 15-28208A and 17-30500A from the Ministry of Health of the Czech Republic and LQ1604 NPU II from the Ministry of Education of the Czech Republic. We thank The National Center for Medical Genomics (LM2015091) for bioinformatical support with next-generation sequencing data analysis and for providing ethnically matched population genotype frequency data (project CZ.02.1.01/0.0/0.0/16_013/0001634). We thank Jana Kasakova for patients’ referrals and clinical assessment.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This article has been changed to correctly present the values in the title that uphold the article’s content.
A correction to this article is available online at https://doi.org/10.1007/s11033-017-4130-3.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dudakova, L., Stranecky, V., Ulmanova, O. et al. Segregation of a novel p.(Ser270Tyr) MAF mutation and p.(Tyr56∗) CRYGD variant in a family with dominantly inherited congenital cataracts. Mol Biol Rep 44, 435–440 (2017). https://doi.org/10.1007/s11033-017-4121-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-017-4121-4