Abstract
Jatropha curcas L. is found in all tropical regions and has garnered lot of attention for its potential as a source of biodiesel. As J. curcas is a plant that is still in the process of being domesticated, interest in improving its agronomic traits has increased in an attempt to select more productive varieties, aiming at sustainable utilization of this plant for biodiesel production. Therefore, the study of genetic diversity in different accessions of J. curcas in Brazil constitutes a necessary first step in genetic programs designed to improve this species. In this study we have used ISSR markers to assess the genetic variability of 332 accessions from eight states in Brazil that produce J. curcas seeds for commercialization. Seven ISSR primers amplified a total of 21,253 bands, of which 19,472 bands (91%) showed polymorphism. Among the polymorphic bands 275 rare bands were identified (present in fewer than 15% of the accessions). Polymorphic information content (PIC), marker index (MI) and resolving power (RP) averaged 0.26, 17.86 and 19.87 per primer, respectively, showing the high efficiency and reliability of the markers used. ISSR markers analyses as number of polymorphic loci, genetic diversity and accession relationships through UPGMA-phenogram and MDS showed that Brazilian accessions are closely related but have a higher level of genetic diversity than accessions from other countries, and the accessions from Natal (RN) are the most diverse, having high value as a source of genetic diversity for breeding programs of J. curcas in the world.







Similar content being viewed by others
References
Fairless D (2007) The little shrub that could—maybe. Nature 449:652–655
Achten WMJ, Verchot L, Franken YJ, Mathijs E, Singh VP, Aerts R, Muys B (2008) Jatropha bio-diesel production and use. Biomass Bioenergy 1–13. doi: 10.1016/j.biombioe.2008.03.003
Becker K, Makkar HPS (2008) Jatropha curcas: a potencial source for tomorrow′s oil and biodiesel. Lipid Technol 20:104–109. doi:10.1002/lite.200800023
Basha SD, Francis G, Makkar HPS, Becker K, Sujatha M (2009) A comparative study of biochemical traits and molecular markers for assessment of genetic relationships between Jatropha curcas L. germplasm from different countries. Plant Sci 176:812–823. doi:10.1016/j.plantsci.2009.03.008
Heller J (1996) Physic nut. Jatropha curcas L. Promoting the conservation and use of underutilized and neglected crops. Insitute of Plant Genetics and Crop Plant Research, Gatersleben, Germany and International Plant Genetic Resources Institute, Rome, Italy, 1996. http://www.bio-nica.info/biblioteca/Heller1996Jatropha.pdf. Accessed 14 Mar 2008
Henning RK (2009) The Jatropha system. An integrated approach of rural development. http://www.jatropha.de/documents/TheJatrophaBook-2009.pdf. Accessed 16 Dec 2009
Kumar A, Sharma S (2008) An evaluation of multipurpose oil seed crop for industrial uses (Jatropha curcas L.): a review. Ind Crops Prod 28:1–10. doi: 10.1016/j.indcrop.2008.01.001
Makkar HPS, Aderibigbe AO, Becker K (1998) Comparative evaluation of non-toxic and toxic varieties of Jatropha curcas for chemical composition, digestibility, protein degradability and toxic factors. Food Chem 62:207–215
Jongschaap REE, Corré WJ, Bindraban PS, Brandenburg WA (2007) Claims and facts on Jatropha curcas L. Global Jatropha curcas evaluation, breeding and propagation programme. http://www.ifad.org/events/jatropha/breeding/claims.pdf. Accessed 14 Mar 2008
Arruda FP, Beltrão NEM, Andrade AP, Pereira WE, Severino LS (2004) Cultivo de pinhão manso (Jatropha curcas L.) como alternativa para o semi-árido nordestino. Revista Brasileira de Oleaginosas e Fibrosas 8:789–799
Ranade SA, Srivastava AP, Rana TS, Srivastava J, Tul R (2008) Easy assessment of diversity in Jatropha curcas L. plants using two simple-primer amplification (SPAR) methods. Biomass Bioenergy 32:533–540. doi:10.1016/j.biombioe.2007.11.006
Gupta S, Srivastava M, Mishra GP, Naik PK, Chauhan RS, Tiwari SK, Kumar M, Singh R (2008) Analogy of ISSR and RAPD markers for comparative analysis of genetic diversity among different Jatropha curcas genotypes. Afr J Biotechnol 7:4230–4243
Ram SG, Parthiban KT, Kumar RS, Thiruvengadam V, Paramathma M (2008) Genetic diversity among Jatropha species as revealed by RAPD markers. Genet Resour Crop Evol. doi:10.1007/s10722-007-9285-7
Pamidiamarri DVNS, Singh S, Mastan SG, Patel J, Reddy MP (2009) Molecular characterization and identification of markers for toxic and non-toxic varieties of Jatropha curcas L. using RAPD, AFLP and SSR markers. Mol Biol Rep 36:1357–1364. doi:10.1007/s11033-008-9320-6
Ikbal, Boora KS, Dhillon RS (2010) Evaluation of genetic diversity of Jatropha curcas L. using RAPD markers. Indian J Biotechnol 9:50–57
Basha SD, Sujatha M (2007) Inter and intra-population variability of Jatropha curcas (L.) characterized by RAPD and ISSR markers and development of population-specific SCAR markers. Euphytica 156:375–386. doi:10.1007/s10681-007-9387-5
Kumar R S, Parthiban K T, Rao M G (2008) Molecular characterization of Jatropha genetic resources through inter-simple sequence repeat (ISSR) markers. Mol Biol Rep. doi:10.1007/s11033-008-9404-3
Tatikonda L, Wani SP, Kannan S, Beerelli N, Sreedevi TK, Hoisington DA, Devi P, Varshney RK (2009) AFLP-based molecular characterization of an elite germplasm collection of Jatropha curcas L., a biofuel plant. Plant Sci 176:505–513. doi:10.1016/j.plantsci.2009.01.006
Sun Q, Li L, Li Y, Wu G, Ge X (2008) SSR and AFLP markers reveal low genetic diversity in the biofuel plant Jatropha curcas in China. Crop Sci 48:1865–1871. doi:10.2135/cropsci208.02.0074
Varshney RK, Chabane K, Hendre PS, Aggarwal RK, Graner A (2007) Comparative assessment of EST-SSR, EST-SNP and AFLP markers for evaluation of genetic diversity and conservation of genetic resources using wild, cultivated and elite barleys. Plant Sci 173:638–649. doi:10.1016/j.plantsci.2007.08.010
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Roldan-Ruiz I, Dendauw J, Vanbockstaele E, Depicker A, De Loose M (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134
Doyle JJ, Doyle JL (1987) A rapid DNA isolation method for small quantities of fresh tissues. Phytochem Bull 19:11–15
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genetic 98:107–112
Bonin A, Ehrich D, Manel S (2007) Statistical analysis of amplified fragment length polymorphism data: a toolbox for molecular ecologists and evolutionists. Mol Ecol 16:3737–3758. doi:10.1111/j.1365-294X.2007.03435.x
Mohammadi SA, Prasanna BM (2003) Analysis of genetic diversity in crop plants—salient statistical tools and considerations. Crop Sci 43:1235–1248
Yeh FC, Yang R, Boyle T (1999) PopGene: Microsoft Window-based freeware for population genetic analysis, version 3.2, University of Alberta, Edmonton
Rohlf FJ (1993) NTSYS-pc (Numerical Taxonomy and Multivariate Analysis System), version 2.11. Exeter Software, Setauket
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval LG, Schneider S (2005) Arlequin ver. 3.1: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Patra B, Acharya L, Mukherjee AK, Panda MK, Panda MC (2008) Molecular characterization of ten cultivars of Canna lilies (Canna Linn.) using PCR based molecular markers (RAPDs and ISSRs). Int J Integr Biol 2:129–137
Thimmappaiah, Santosh WG, Shobha D, Melwyn GS (2009) Assessment of genetic diversity in cashew germplasm using RAPD and ISSR markers. Sci Hortic 120:411–417. doi:10.1016/j.scienta.2008.11.022
Gomes S, Martins-Lopes P, Lopes J, Guedes-Pinto H (2009) Assessing genetic diversity in Olea europaea L. using ISSR and SSR markers. Plant Mol Biol Rep 27:365–373. doi:10.1007/s11105-009-0106-3
Muthusamy S, Kanagarajan S, Ponnusamy S (2008) Efficiency of RAPD and ISSR markers system in accessing genetic variation of rice bean (Vigna umbellata) landraces. Electron J Biotechnol. doi:10.2225/vol11-issue3-fulltext-8
Fernandéz ME, Figueiras AM, Benito C (2002) The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet 104:845–851. doi:10.1007/s00122-001-0848-2
Acknowledgments
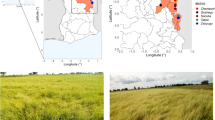
We would like to thank the farmers Nagashi Tominaga and Divino Nunes for donating J. curcas seeds grown in Janaúba (MG) and Rio Verde (GO) and Dr. Antônio Andrade of the Instituto de Pesquisas Jardim Botânico do Rio de Janeiro for his generous assistance with seed germination. We also thank Prof. Wellington Matos from the Universidade do Grande Rio for help with the map in Fig. 1. We are especially grateful to Prof. Martha Sorenson from the Universidade Federal do Rio de Janeiro for language editing. We are grateful to Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for financial support and fellowships.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Grativol, C., da Fonseca Lira-Medeiros, C., Hemerly, A.S. et al. High efficiency and reliability of inter-simple sequence repeats (ISSR) markers for evaluation of genetic diversity in Brazilian cultivated Jatropha curcas L. accessions. Mol Biol Rep 38, 4245–4256 (2011). https://doi.org/10.1007/s11033-010-0547-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0547-7