Abstract
An efficient vector, designated as pCAGX, was designed for direct cloning and enhanced expression of PCR-amplified ORFs in mammalian cells. It relied on the well-known TA-cloning principle, and utilized the CMV enhancer/chicken β-actin/rabbit β-globin (CAG) hybrid promoter instead of the classical CMV promoter to drive more efficient transgene expression in wider host cells. The specially designed cassette under CAG hybrid promoter contained two tandemly arrayed XcmI sites which were spaced by an additional EcoRV site. For direct cloning and expressing PCR-amplified ORFs, the T-vector was prepared by further digesting the EcoRV-linearized pCAGX with XcmI to produce T tails on both 3′-ends, which could efficiently minimize the non-recombinant background of T-vector and eliminate the necessity of selective marker genes such as LacZ that allowed blue/white screening. Various PCR fragments in length were prepared to verify the cloning efficiency by ligation with this vector, and GFP gene expression under control of the CAG hybrid promoter in different host cells was assayed by flow cytometry. The results indicated that this vector was higher efficient, especially suitable for cloning and expressing a number of interesting ORFs in parallel, and higher-level transgene expression in different mammalian cells was obtained than the reported vectors using the CMV promoter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Expression analyses is often employed to identify the genes of interest which were deduced by a computer program and cDNA analysis from sequence information of sequenced genomes. The PCR fragments of interesting genes were generally cloned into a cloning vector of Escherichia coli, usually T-vector, and then subcloned into an expression vector [1–3]. But this traditional strategy had several disadvantages as well. First, it was relatively time and labor consuming because of two steps of subcloning. Second, since the subcloning was based upon enzymatic digestion of the insert and the vector, attention must be paid on the compatibility of restriction enzymes, especially a longer open reading frames (ORFs) sequence containing more restriction sites. These disadvantages were even more notable sometimes when expression of large numbers of ORFs was required. Thus, an appropriate vector for direct cloning and immediate expression of PCR-amplified ORFs is significant.
The commercial “Gateway” system based on the site-specific recombination is amenable to high-throughput cloning, but it depends on more expensive bacteriophage λ integrase and requires the specifically designed PCR primers with 16–29 bp tail sequences [4, 5]. The vaccinia DNA topoisomerase I-based “TOPO” cloning technology which can avoid the use of restriction endonucleases is rapid, however, the cleavage of vectors and subsequent religation with PCR fragments need the expensive topoisomerase, and the commercial “TOPO” cloning kits were also extremely expensive [6–9]. For these strategies, they either suffer from high costs using the expensive enzymes, or demand specifically designed primers for amplification of ORFs and ligation with vectors. Moreover, the classical CMV (Human Cytomegalovirus) promoter adopted in these systems is not efficient enough for consistent and higher-level transgene expression, and its host cells range is limited [10–14].
In this paper, a well-applicable TA-cloning strategy was chosen, and an efficient pCAGX vector for direct cloning and enhanced expression of PCR-amplified ORFs in mammalian cells was constructed and verified. The PCR-amplified ORFs can be directly cloned into the TA site of this vector with minimized nonrecombinant background, and more efficiently expressed under control of the CMV enhancer/chicken β-actin/rabbit β-globin (CAG) hybrid promoter which was indicated higher efficiency and wider host cells than the popular CMV promoter [10–14]. Results demonstrated that this vector was higher efficient and suitable for high-throughput cloning and functional analysis of interesting genes, and higher-level transgene expression was obtained than the reported vectors using CMV promoter. Also, the vector designated as pCAGX-T was a derivative of pCDNA3.1, and therefore shared the same advantage of the parent vector to select the stablely expressing cell lines by the neomycin resistant cassette.
Materials and methods
Strains, vectors and reagents
Escherichia coli JM109, the plasmids of pCDNA3.1 and pEGFP-C1 were preserved in Henan Provincial Key Laboratory of Animal Immunology (Zhengzhou, China). All the reagents or kits, such as LA Taq DNA Polymerase, PrimeSTAR DNA Polymerase, T4 DNA Ligase, Restriction Enzyme, Bacterial Plasmid DNA Extraction Kit Ver.2.0, Universal Genomic DNA Extraction Kit Ver.3.0, Agarose Gel DNA purification kit ver.2.0, pMD19-T vectors Kit and DNA Marker were purchased from TaKaRa Company (Dalian, China). XcmI restriction enzyme was obtained from BioLads Company (New England). Lipofectamine 2000 reagent was from Invitrogen Company. The primers used in this study were synthesized in TaKaRa Company (Table 1).
Cell culture
African green monkey kidney COS-7 cells, Porcine kidney epithelial PK15 cells, Baby hamster kidney BHK21 cells, Human hepatocellular carcinoma SMMC-7721 cells and Human cervical carcinoma HeLa cells were obtained from Henan Provincial Key Laboratory of Animal Immunology (Zhengzhou, China), and maintained in dulbecco’s modified eagle’s medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (FBS, Gibco). Chicken pre-B lymphocyte DT40 cells were from China Central Type Culture Collection (Shanghai, China), for the cultivation of DT40 cells, RPMI-1640 growth medium was supplemented with 10% FBS, 5% chicken serum (CHS, Gibco), 50 μM 2-mercaptoethanol.
Mouse immunoglobulin G (IgG) against chicken erythrocytes
Mouse IgG was purified from hyperimmune serum collected from a mouse that had been inoculated with chicken erythrocytes. IgG was precipitated with ammonium sulphate ((NH4)2SO4) and further purified by DEAE chromatography according to the reported method [15].
Preparation of chicken β-actin/rabbit β-globin (AG) hybrid promoter
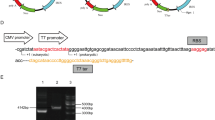
The genome DNA of DT40 cells was isolated using DNA Extraction Kit according to the manufacturer’s instruction, and a 1,236 bp fragment of the chicken β-actin promoter (GenBanK: E02199, 1–1,236 bp) was amplified with primer pair actin-F/actin-R. Then as shown in Fig. 1, the chicken β-actin promoter fragment was further amplified by PCR using primer pair actin-F/actin-R1, followed the subsequent PCR amplification with primer pair acin-F/actin-R2. All the previous PCRs were carried out with high fidelity PrimeSTAR DNA Polymerase. As a result, the rabbit β-globin intron (GenBanK: E03011, 1,237–1,339 bp) was fused with the chicken β-actin promoter, which not only resulted in the chicken β-actin/rabbit β-globin (AG) hybrid promoter but also created the annealing sites of universal M13-47 primer.
Construction of pCAGX vector
The chicken β-actin/rabbit β-globin (AG) hybrid promoter fragment and pCDNA3.1 were both digested by SnaBI and KpnI. Then the AG promoter was cloned into pCDNA3.1 and the vector pCDNA-AG was constructed. An XcmI intermediate cassette containing a 610 bp spacer DNA sequence between two tandemly XcmI sites was obtained from pUC19 by PCR amplification with primer pair medfrag-F/medfrag-R. After digested by BamHI and XhoI, the intermediate cassette was ligated into pCDNA-AG to construct pCAG-Mf. All the previous PCRs were carried out with high fidelity PrimeSTAR DNA Polymerase. After digested by EcoRV to remove the spacer DNA sequence, the EcoRV-digested pCAG-Mf was cyclized by self-ligation, resulting in a new construct termed pCAGX. The constructing procedure of pCAGX was shown in Fig. 2.
Schematic diagram of pCAGX and pCAGX-T construct. a Construction process of pCAGX-T. b Multiple cloning sites (MCS) region of pCAGX-T. Abbreviations are: AG, chicken β-actin/rabbit β-globin hybrid promoter; Mf, intermediate fragment; PCMV, human cytomegalovirus promoter; BGHpA, bovine growth hormone polyadenylation signal; PCAG, CMV enhancer/chicken β-actin/rabbit β-globin hybrid promoter; PSV40, Simian Virus 40 promoter; Neo r, neomycin resistant gene; SV40pA, Simian Virus 40 polyadenylation signal
Preparation of pCAGX-T for PCR fragments cloning
After cutting the pCAGX vectors with EcoRV, the freezed absolute alcohol was added into the digest to quickly precipitate the vectors DNA. Then the linearized pCAGX vectors were prepared. For T-vector, the linearized pCAGX was subsequently cleaved with XcmI at 37°C over night. Then the excessive restriction enzyme was removed by phenol-extraction. After being precipitated by freezed absolute alcohol and resuspended in deionised H2O, the released T-vector renamed pCAGX-T resulting from cleavage at both XcmI sites was prepared (Fig. 2). The resultant pCAGX-T was about 70 ng/μl finally.
Verifying TA-cloning efficiency of pCAGX-T
To test the cloning efficiency of pCAGX-T vectors, various PCR fragments were prepared for the ligation with the vectors. All the PCR fragments were shown in Table 2. These PCR fragments with 3′-A overhang were, respectively, ligated with the linearized pCAGX-T vectors. Vectors (0.03 pmol) and PCR fragments (0.18 pmol) were combined in 10 μl volume with 350 units T4 DNA ligase. After incubation at 16° for 4 h, the ligation transformed E. coli JM109. Total about 150 transformants for each PCR fragment were picked randomly from plates and tested by colony PCR assay with primer pair M13-47/BGHrev. Several samples of transformants in each assay were casually chosen to be further identified by plasmid restriction enzyme digestions and agarose gel. According to the results of PCR scanning, the comparison of the cloning efficiency was performed between pCAGX-T and pMD19-T (TaKaRa Company).
Cloning and expression of PCR-amplyfied GFP gene using pCAGX-T
To characterize the usefulness of pCAGX-T, PCR was performed using LA Taq DNA polymerase to amplify the 717 bp GFP (green fluorescent protein) gene (GenBanK: U55763, 613–1,329 bp) from the plasmids of pEGFP-C1, and the amplified GFP fragments were ligated with the linearized pCAGX-T vectors. Colony PCR was employed to identify the direction of the inserts using a forward primer GFP-F homologous to the insert and a reverse universal primer BGHrev homologous to the vector. The pCAGT-GFP construct harboring the forward oriented GFP coding sequence under control of the CAG hybrid promoter was transfected into COS-7, PK15, BHK21, SMMC-7721, and HeLa cells using a Lipofectamine 2000 reagent according to the manufacturer’s standard protocol. The pCDNA3.1-GFP construct containing GFP coding sequence driven by the CMV promoter was also transfected into the same cells as control. The expression of GFP was observed 36 h after transfection under a fluorescence microscope. Meanwhile, transfected cells were harvested to assay for GFP fluorescence by FACS Calibur (BD) and the strength of CAG and CMV promoters was compared more quantitatively.
Cloning and expression of mouse IgG Fc receptor II (FcγRII or CD32) utilizing pCAGX-T
Peripheral blood leucocytes (PBLs) were isolated from the peripheral blood of mouse by density centrifugation on lymphoprep, and RT–PCR was performed on the mRNA from PBLs using primers of moFc2b-F and moFc2b-R to amplify the FcγRII gene (Genbank: NM_010187, 69–950 bp). The resultant PCR fragments were directly cloned into the pCAGX-T vectors, and the clones containing plasmids with forward oriented inserts were quickly scanned via colony PCR using the combined primers moFc2b-F and BGHrev. The comparative cDNA constructs pCAGT-moFcRII and pCDNA3.1-moFcRII were transfected into COS-7 cells, respectively, as described previously. Thirty six hours post transfection, the FcγRII molecules expressed on the surface of transfected COS-7 cells were checked according to the immunoglobulin-binding assay reported before [16]. In brief, transfected COS-7 cells were incubated in DMEM medium without serum for 2 h and the chicken erythrocytes were sensitized with mouse IgG for 2 h at 4°C. The IgG-sensitized chicken erythrocytes were resuspended in serum-free medium and added to the mouse FcγRII-expressed COS-7 cell monolayer. After 45 min incubation at room temperature with occasional gentle agitation, non-adherent erythrocytes were washed off with phosphate buffered saline (PBS). The monolayer cells were fixed with methanol for 10 min, and stained with HRP-conjugated goat anti-mouse IgG antibody, washed and incubated with 3-amino-9-ethylcarbazole (AEC) (Sigma).
Statistical analysis
SPSS for Windows (standard version 11.5) was used for statistical analysis. One-way anova was performed to compare differences between the new construct of pCAGX-T and the commercial vectors of PMD19-T and pCDNA3.1, with respect to TA-cloning and transgene expression efficiency. Summary statistics were expressed as the means ± the standard deviations (M ± SD), except as otherwise stated. In all statistical analysis, a P value of less than 0.05 was considered statistically significant, and all P values were two sided.
Results and discussion
Construction of pCAGX
In the pCAGX vector, the foreign gene expressing cassette consisted of CAG promoter (1–1,724 bp), MCS (1,725–1,843 bp) and BGH terminator (1,844–2,125 bp). Sequencing result showed that an ingenious XcmI cassette containing two tandemly arrayed XcmI sites spaced by an addition EcoRV site was exactly incorporated into the downstream MCS of the CAG promoter, resulting in a new construct named pCAGX (Fig. 2). After cutting pCAGX at both XcmI sites with XcmI, the released T-vectors termed pCAGX-T could be quickly prepared in large quality. The PCR-amplified ORFs can be directly cloned into pCAGX-T and the construct harboring the forward oriented ORFs under control of the hybrid CAG promoter can also be easily identified using a forward primer homologous to the insert and a reverse universal primer BGHrev homologous to the vector.
TA-cloning efficiency of pCAGX-T
Unlike the well-known pUC or pGEM vectors using white-blue screening to select positive clones [17, 18], there was a specially designed cassette containing an additional EcoRV site between the two tandemly arrayed XcmI sites, adopted in pCAGX to increase TA-cloning efficiency and eliminate the necessity of selective marker genes such as LacZ that allowed blue-white screening. For the preparation of pCAGX-T, It is important to perform EcoRV digestion of pCAGX prior to the digestion with XcmI. In this way, even if the vectors were partly digested by XcmI, the incompletely cleaved molecules would have been completely digested by the efficient enzyme EcoRV, and could be prevented from self-ligation because of no complementation of one blunt end and one cohesive end. Thus the self-circularization of T-vectors would be effectively minimized. Different PCR fragments in length 500, 1,000, 1,500, and 2,000 bp, were used to test the TA-cloning efficiency. Almost 95% of cloning efficiency was got when 500 bp of PCR fragments were verified. This efficiency was similar to the conventional blue-white selection (Table 3). When PCR fragments were as large as 1,000–2,000 bp, over 80.4% of clones still contained recombinant plasmids with inserts of the correct size (Table 3). These results demonstrated that pCAGX-T was also perfect T vector and its TA-cloning efficiency was obviously higher than pMD19-T vector in statistics. Also, the cloning efficiency of pCAGX-T was substantially increased when the combined use of XcmI and EcoRV instead of XcmI treatment alone (data not shown).
Cloning and expression of GFP by pCAGX-T
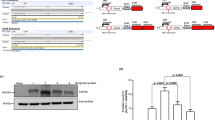
Two comparable construct pCDNA3.1-GFP and pCAGT-GFP in which GFP expression were under control of CMV and CAG hybrid promoter respectively, were used to check the transgene expression. After transfecting COS-7, PK15, BHK21, SMMC-7721, and HeLa cells, the percentage of GFP-expressing cells and level of GFP fluorescence were quantified by flow cytometry. Each of the vectors could generate GFP-expressing cells (Fig. 3), and similar transfection efficiencies were observed (Fig. 3c). But the mean fluorescence intensity of the GFP-positive cells produced by pCAGT-GFP was significantly higher than pCDNA3.1-GFP (Fig. 3d). So pCAGX-T using the hybrid CAG promoter not only had perfect transfection efficiencies but also presented higher-level transgene expression than the commercial pCDNA3.1 utilizing the classic CMV promoter.
Comparison of GFP expression in different cells transfected by pCDNA3.1-GFP and pCAGT-GFP construct, respectively. a GFP expression observed under fluorescence microscope. The respective left panels were observed under white light. Magnification, ×200. b Histograms of fluorescence-activated cell sorting (FACS) analysis for GFP expression in the different transfected cells. Thirty six hours post-transfection, the transfected cells were harvested to analyze the GFP expression by flow cytometry. Transfection experiments were conducted for each vector in triplicate. Red and black denoted vectors expressing GFP and empty vectors transfected controls, respectively. The M1 region define the GFP-positive cells. c Percentage of GFP-expressing cells transfected by different construct. Transfection efficiencies indicated by the percentage of GFP-expressing cells have no differences in statistics. #, P > 0.05. d Mean fluorescence intensity of GFP expression in the GFP-positive-only cell population. GFP expression driven by the CAG promoter seems significantly higher than the CMV promoter in mammalian cells. §, P < 0.031; §§, P < 0.016; *, P < 0.01; **, P < 0.005; ***, P < 0.002. Abbreviations: COS-7, African green monkey kidney cells; PK15, porcine kidney epithelial cells; BHK21, baby hamster kidney cells; SMMC-7721, human hepatocellular carcinoma cells; HeLa, human cervical carcinoma cells (For interpretation of color mentioned in this figure legend, the reader is referred to the web version of the article.)
Cloning and expression of mouse IgG FcγRII by pCAGX-T
Rosetting test was performed to confirm the expression of mouse IgG receptor FcγRII, results shew that COS-7 cells, respectively, transfected by pCAGT-moFcRII and pCDNA3.1-moFcRII construct could bind to the chicken erythrocytes sensitized with mouse anti-erythrocytes IgG, but the pCAGT-moFcRII transfected cells seemed more efficient to bind the sensitized chicken erythrocytes (Fig. 4a, c). Sensitized erythrocytes did not attach to the empty vectors transfected COS-7 cells (Fig. 4b, d), and non-sensitized red cells did not attach to the transfected COS-7 cells (data not shown). The binding assay further demonstrated the CAG hybrid promoter seemed more efficient to drive transgene expression than the classical CMV promoter.
Rosetting test of the binding of mouse IgG-sensitized chicken erythrocytes to COS-7 cells transfected by the mouse FcγRII cDNA construct. a COS-7 cells transfected with pCAGT-moFcγRII and incubated with erythrocytes specifically sensitized with mouse IgG. Extensive binding of erythrocytes to individual cells was evident. b COS-7 cells transfected with empty pCAGX and incubated with mouse IgG-sensitized erythrocytes. No binding of erythrocytes was evident. c COS-7 cells transfected with pCDNA3.1-moFcγRII and incubated with mouse IgG-sensitized erythrocytes. d COS-7 cells transfected with empty pCDNA3.1 and incubated with mouse IgG-sensitized erythrocytes. No binding of erythrocytes was evident
In the commercial expression cloning system such as the recombination-based “Gateway” system and topoisomerase-dependent “TOPO” technology, the expensive enzymes must be used, and the commercial kits of this kind were extremely expensive. Also, especially in the “Gateway” system, the primers were also needed to be specifically designed in that the specific tail sequences of PCR fragments were necessary for ligation with vectors [4–9]. Furthermore, the classical CMV promoter of these system is not efficient enough for transgene expression, especially consistent and higher-level expression, and its host cells range is more limited than the hybrid CAG promoter [10–14].
But in the pCAGX vector of this work, it didn’t depend on the expensive integrase and topoisomerase. Also, the specific tail sequences of primers were not demanded. The specially designed cassette contained two tandemly arrayed XcmI sites spaced by an additional EcoRV, which not only minimized the non-recombinant background of T-vector and eliminated the necessity of selective marker genes such as LacZ that allowed blue/white screening, but also made the quick ligation of PCR-amplified ORFs and vectors realized with the cheaper T4 DNA ligase. The clones containing forward oriented ORFs under the hybrid CAG promoter can be quickly scanned via colony PCR. Moreover, the stronger CAG hybrid promoter ensure consistent and higher-level transgene expression in wider host cells. Results also showed that the T-vector was higher efficient, especially suitable for high-throughput cloning and expression analysis of interesting genes in parallel, and higher-level transgene expression driven by the CAG hybrid promoter was obtained than the CMV promoter. Therefore, this vector designed for direct cloning and immediate expression is higher efficient and more convenient in manipulation than the reported vectors.
References
Hurgin V, Novick D, Rubinstein M (2002) The promoter of IL-18 binding protein: activation by an IFN-gamma-induced complex of IFN regulatory factor 1 and CCAAT/enhancer binding protein beta. Proc Natl Acad Sci USA 99(26):16957–16962
Li W, Zhang WW, Yang MM et al (2008) Cloning of the thermostable cellulase gene from newly isolated Bacillus subtilis and its expression in Escherichia coli. Mol Biotechnol 40(2):195–201
You L, Weng H, Chen Z et al (2009) A novel vector for direct cloning PCR fragments by positive selection based on the lethal barnase. Mol Biol Rep 36(7):1793–1798
Cheo DL, Titus SA, Byrd DR et al (2004) Concerted assembly and cloning of multiple DNA segments using in vitro site-specific recombination: functional analysis of multi-segment expression clones. Genome Res 14(10B):2111–2120
Hartley JL, Temple GF, Brasch MA (2000) DNA cloning using in vitro site-specific recombination. Genome Res 10(11):1788–1795
Shuman S (1994) Novel approach to molecular cloning and polynucleotide synthesis using vaccinia DNA topoisomerase. J Biol Chem 269(51):32678–32684
Shuman S (1991) Recombination mediated by vaccinia virus DNA topoisomerase I in Escherichia coli is sequence specific. Proc Natl Acad Sci USA 88(22):10104–10108
Shuman S, Prescott J (1990) Specific DNA cleavage and binding by vaccinia virus DNA topoisomerase I. J Biol Chem 265(29):17826–17836
Stivers JT, Shuman S, Mildvan AS (1994) Vaccinia DNA topoisomerase I: single-turnover and steady-state kinetic analysis of the DNA strand cleavage and ligation reactions. Biochemistry 33(1):327–339
Alexopoulou AN, Couchman JR, Whiteford JR (2008) The CMV early enhancer/chicken beta actin (CAG) promoter can be used to drive transgene expression during the differentiation of murine embryonic stem cells into vascular progenitors. BMC Cell Biol 9:2
Chung S, Andersson T, Sonntag KC et al (2002) Analysis of different promoter systems for efficient transgene expression in mouse embryonic stem cell lines. Stem Cells 20(2):139–145
Garg S, Oran AE, Hon H et al (2004) The hybrid cytomegalovirus enhancer/chicken beta-actin promoter along with woodchuck hepatitis virus posttranscriptional regulatory element enhances the protective efficacy of DNA vaccines. J Immunol 173(1):550–558
Liew CG, Draper JS, Walsh J et al (2007) Transient and stable transgene expression in human embryonic stem cells. Stem Cells 25(6):1521–1528
Miyazaki J, Takaki S, Araki K et al (1989) Expression vector system based on the chicken beta-actin promoter directs efficient production of interleukin-5. Gene 79(2):269–277
Temponi M, Kageshita T, Perosa F et al (1989) Purification of murine IgG monoclonal antibodies by precipitation with caprylic acid: comparison with other methods of purification. Hybridoma 8(1):85–95
Zhang G, Young JR, Tregaskes CA et al (1995) Identification of a novel class of mammalian Fc gamma receptor. J Immunol 155(3):1534–1541
Jeung JU, Cho SK, Shim KS et al (2002) Construction of two pGEM-7Zf(+) phagemid T-tail vectors using AhdI-restriction endonuclease sites for direct cloning of PCR products. Plasmid 48(2):160–163
Ullmann A (1992) Complementation in beta-galactosidase: from protein structure to genetic engineering. Bioessays 14(3):201–205
Acknowledgments
This work was supported by the National Basic Research Program (2005CB523200) and National Key Technology R&D Program (2006BAD06A04-6). We are grateful to the doctor of Li-yun Zheng at Zhengzhou university for his scientific promotion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
You, Lm., Luo, J., Wang, Ap. et al. A hybrid promoter-containing vector for direct cloning and enhanced expression of PCR-amplified ORFs in mammalian cells. Mol Biol Rep 37, 2757–2765 (2010). https://doi.org/10.1007/s11033-009-9814-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-009-9814-x