Abstract
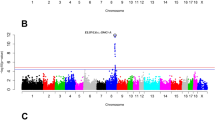
Cathepsin K (CTSK) was selected as a candidate gene for fat deposition in pigs because recently, in human and mouse, it was shown that this lysosomal proteinase is an obesity marker. A single nucleotide polymorphism (SNP) was identified in intron 4 of the porcine CTSK gene (g.15G>A; FM209043). Allele frequencies of this polymorphism were analysed in seven pig breeds. Radiation hybrid mapping confirmed the localization of CTSK to porcine chromosome 4, close to the FAT1 QTL region. Three populations of pigs (one Italian Large White and two Italian Duroc groups of pigs) were selected for association analysis. In the Italian Large White breed the g.15G>A SNP was not informative. Association analysis including all Italian Duroc pigs showed that the CTSK marker was associated with back fat thickness and lean cuts (P < 0.01), and average daily gain and feed:gain ratio (P < 0.05) estimated breeding values.
Similar content being viewed by others
References
Kim KS, Larsen N, Short T, Plastow G, Rothschild MF (2000) A missense variant of the porcine melanocortin-4 receptor (MC4R) gene is associated with fatness, growth, and feed intake traits. Mamm Genome 11:131–135. doi:10.1007/s003350010025
Van Laere AS, Nguyen M, Braunschweig M, Nezer C, Collette C, Moreau L, Archibald AL, Haley CS, Buys N, Tally M, Andersson G, Georges M, Andersson L (2003) A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 425:832–836. doi:10.1038/nature02064
Fontanesi L, Scotti E, Buttazzoni L, Davoli R, Russo V (2009) The porcine fat mass and obesity associated (FTO) gene is associated with fat deposition in Italian Duroc pigs. Anim Genet 40:90–93. doi:10.1111/j.1365-2052.2008.01777.x
Turk V, Turk B, Turk D (2001) Lysosomal cysteine proteases: facts and opportunities. EMBO J 20:4629–4633. doi:10.1093/emboj/20.17.4629
Brix K, Dunkhorst A, Mayer K, Jordans S (2008) Cysteine cathepsins: cellular roadmap to different functions. Biochimie 90:194–207. doi:10.1016/j.biochi.2007.07.024
Drake FH, Dodds RA, James IE, Connor JR, Debouck C, Richardson S, Lee-Rykaczewski E, Coleman L, Rieman D, Barthlow R, Hastings G, Gowen M (1996) Cathepsin K, but not cathepsins B, L, or S, is abundantly expressed in human osteoclasts. J Biol Chem 271:12511–12516. doi:10.1074/jbc.271.21.12511
Saftig P, Hunziker E, Wehmeyer O, Jones S, Boyde A, Rommerskirch W, Moritz JD, Schu P, von Figura K (1998) Impaired osteoclastic bone resorption leads to osteopetrosis in cathepsin-K-deficient mice. Proc Natl Acad Sci USA 95:13453–13458. doi:10.1073/pnas.95.23.13453
Asagiri M, Hirai T, Kunigami T, Kamano S, Gober HJ, Okamoto K, Nishikawa K, Latz E, Golenbock DT, Aoki K, Ohya K, Imai Y, Morishita Y, Miyazono K, Kato S, Saftig P, Takayanagi H (2008) Cathepsin K-dependent toll-like receptor 9 signaling revealed in experimental arthritis. Science 319:624–627. doi:10.1126/science.1150110
Chiellini C, Costa M, Novelli SE, Amri EZ, Benzi L, Bertacca A, Cohen P, Del Prato S, Friedman JM, Maffei M (2003) Identification of cathepsin K as a novel marker of adiposity in white adipose tissue. J Cell Physiol 195:309–321. doi:10.1002/jcp.10253
Xiao Y, Junfeng H, Tianhong L, Lu W, Shulin C, Yu Z, Xiaohua L, Weixia J, Sheng Z, Yanyun G, Guo L, Min L (2006) Cathepsin K in adipocyte differentiation and its potential role in the pathogenesis of obesity. J Clin Endocrinol Metab 91:4520–4527. doi:10.1210/jc.2005-2486
Funicello M, Novelli M, Ragni M, Vottari T, Cocuzza C, Soriano-Lopez J, Chiellini C, Boschi F, Marzola P, Masiello P, Saftig P, Santini F, St-Jacques R, Desmarais S, Morin N, Mancini J, Percival MD, Pinchera A, Maffei M (2007) Cathepsin K null mice show reduced adiposity during the rapid accumulation of fat stores. PLoS One 2:e683. doi:10.1371/journal.pone.0000683
Moller M, Berg F, Riquet J, Pomp D, Archibald A, Anderson S, Feve K, Zhang Y, Rothschild M, Milan D, Andersson L, Tuggle CK (2004) High-resolution comparative mapping of pig chromosome 4, emphasizing the FAT1 region. Mamm Genome 15:717–731. doi:10.1007/s00335-004-2366-4
Andersson L, Haley CS, Ellegren H, Knott SA, Johansson M, Andersson K, Andersson-Eklund L, Edfor-Lilja I, Fredholm M, Hansson I, Håkansson J, Lundström K (1994) Genetic mapping of quantitative trait loci for growth and fatness in pigs. Science 263:1771–1774. doi:10.1126/science.8134840
Malek M, Dekkers JC, Lee HK, Baas TJ, Rothschild MF (2001) A molecular genome scan analysis to identify chromosomal regions influencing economic traits in the pig. I. Growth and body composition. Mamm Genome 12:630–636. doi:10.1007/s003350020018
Milan D, Bidanel JP, Iannuccelli N, Riquet J, Amigues Y, Gruand J, Le Roy P, Renard C, Chevalet C (2002) Detection of quantitative trait loci for carcass composition traits in pigs. Genet Sel Evol 34:705–728. doi:10.1051/gse:2002026
Cepica S, Stratil A, Kopecny M, Blazkova P, Schröffel J Jr, Davoli R, Fontanesi L, Reiner G, Bartenschlager H, Moser G, Geldermann H (2003) Linkage and QTL mapping for Sus scrofa chromosome 4. J Anim Breed Genet 120(Suppl. 1):28–37. doi:10.1046/j.0931-2668.2003.00421.x
Russo V, Fontanesi L, Scotti E, Beretti F, Davoli R, Nanni Costa L, Virgili R, Buttazzoni L (2008) Single nucleotide polymorphisms in several porcine cathepsin genes are associated with growth, carcass and production traits in Italian Large White pigs. J Anim Sci 86:3300–3314. doi:10.2527/jas.2008-0920
Tepel C, Brömme D, Herzog V, Brix K (2000) Cathepsin K in thyroid epithelial cells: sequence, localization and possible function in extracellular proteolysis of thyroglobulin. J Cell Sci 113:4487–4498
Yerle M, Pinton P, Robic A, Alfonso A, Palvadeau Y, Delcros C, Hawken R, Alexander L, Beattie C, Schook L, Milan D, Gellin J (1998) Construction of a whole-genome radiation hybrid panel for high-resolution gene mapping in pigs. Cytogenet Cell Genet 82:182–188. doi:10.1159/000015095
Milan D, Hawken R, Cabau C, Leroux S, Genet C, Lahbib Y, Tosser G, Robic A, Hatey F, Alexander L, Beattie C, Schook L, Yerle M, Gellin J (2000) IMpRH server: an RH mapping server available on the Web. Bioinformatics 16:558–559. doi:10.1093/bioinformatics/16.6.558
Ramos AM, Glenn KL, Serenius TV, Stalder KJ, Rothschild MF (2008) Genetic markers for the production of US country hams. J Anim Breed Genet 125:248–257. doi:10.1111/j.1439-0388.2007.00710.x
Hamasima N, Mikawa A, Suzuki H, Suzuki K, Uenishi H, Awata T (2008) A new 4016-marker radiation hybrid map for porcine-human genome analysis. Mamm Genome 19:51–60. doi:10.1007/s00335-007-9081-x
Hawken RJ, Murtaugh J, Flickinger GH, Yerle M, Robic A, Milan D, Gellin J, Beattie CW, Schook LB, Alexander LJ (1999) A first-generation porcine whole-genome radiation hybrid map. Mamm Genome 10:824–830. doi:10.1007/s003359901097
Berg F, Stern S, Andersson K, Andersson L, Moller M (2006) Refined localization of the FAT1 quantitative trait locus on pig chromosome 4 by marker-assisted backcrossing. BMC Genet 7:7. doi:10.1186/1471-2156-7-17
Mercadé A, Pérez-Enciso M, Varona L, Alves E, Noguera JL, Sánchez A, Folch JM (2006) Adipocyte fatty-acid binding protein is closely associated to the porcine FAT1 locus on chromosome 4. J Anim Sci 84:2907–2913. doi:10.2527/jas.2005-663
Davoli R, Fontanesi L, Braglia S, Nisi I, Sctti E, Buttazzoni L, Russo V (2006) Investigation of SNPs in the ATP1A2, CA3 and DECR1 genes mapped to porcine chromosome 4: analysis in group of pigs divergent for meat production and quality traits. Ital J Anim Sci 5:249–263
Ojeda A, Rozas J, Folch JM, Pérez-Enciso M (2006) Unexpected high polymorphism at the FABP4 gene unveils a complex history for pig populations. Genetics 174:2119–2127. doi:10.1534/genetics.106.063057
Ojeda A, Estellé J, Folch JM, Pérez-Enciso M (2008) Nucleotide variability and linkage disequilibrium patterns at the porcine FABP5 gene. Anim Genet 39:468–473. doi:10.1111/j.1365-2052.2008.01752.x
Chavey C, Mari B, Monthouel MN, Bonnafous S, Anglard P, Van Obberghen E, Tartare-Deckert S (2003) Matrix metalloproteinases are differentially expressed in adipose tissue during obesity and modulate adipocyte differentiation. J Biol Chem 278:11888–11896. doi:10.1074/jbc.M209196200
Acknowledgements
We thank Dr. M. Yerle (INRA) for providing the INRA-Minnesota 7000 rads RH panel and ANAS for providing samples and data. This work was supported by University of Bologna (FAGenomicH project) and MiPAAF (Selmol project) funds.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fontanesi, L., Scotti, E., Buttazzoni, L. et al. A single nucleotide polymorphism in the porcine cathepsin K (CTSK) gene is associated with back fat thickness and production traits in Italian Duroc pigs. Mol Biol Rep 37, 491–495 (2010). https://doi.org/10.1007/s11033-009-9678-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-009-9678-0