Abstract
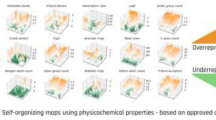
Self-Organizing Map (SOM) models were built to distinguish inhibitors of HMG-CoA reductase from its non-binding decoys. The molecules were represented by five global molecular descriptors and seven 2D property autocorrelation descriptors. Based on these molecular descriptors, 35 HMG-CoA reductase ligands and 1480 decoys were projected into a self-organizing network. In the map, the ligands and the decoys were well separated, where no neuron was occupied by a ligand and a decoy at the same time. Afterward, the discriminating power of the selected molecular descriptors was further validated by extending the datasets to 135 inhibitors. Finally, the SOM approach was subsequently used to identify active compounds in a virtual screening experiment by an external test set which included 32 HMG-CoA reductase inhibitors and 1103 decoys. In this study, 84.4% of the inhibitors (true positives) are retrieved with 15% contamination by non-hits (false positives). The SOM models obtained in this article exhibited powerful ability in virtual screening to find novel inhibitors for HMG-CoA reductase.
Similar content being viewed by others
References
Mukhtar RY, Reckless JP (2005) Statin-induced myositis: a commonly encountered or rare side effect? Curr Opin Lipidol 16: 640–647
Bauknecht H, Zell A, Bayer H, Levi P, Wagener M, Sadowski J, Gasteiger J (1996) Locating biologically active compoundsin medium-sized heterogeneous datasets by topological autocorrelation vectors: dopamine and benzodiazepine agonists. J Chem Inf Comput Sci 36: 1205–1213. doi:10.1021/ci960346m
Bleicher KH, Böhm H, Müller K, Alanine AI (2003) Hit and lead generation: beyond high-throughput screening. Nat Rev Drug Discov 2: 369–378. doi:10.1038/nrd1086
Schneider G (2002) Trends in virtual combinatorial library design. Curr Med Chem 9: 2095–2101. doi:10.2174/0929867023368755
Dessalew N, Bharatam PV (2007) Identification of potential glycogen kinase-3 inhibitors by structure based virtual screening. Biophys Chem 128: 165–175. doi:10.1016/j.bpc.2007.04.001
Gruneberg S, Stubbs MT, Klebe G (2002) Successful virtual screening for novel inhibitors of human carbonic anhydrase: Strategy and experimental confirmation. J Med Chem 45: 3588–3602. doi:10.1021/jm011112j
Powers RA, Morandi F, Shoichet BK (2002) Structure-based discovery of a novel, noncovalent inhibitor of AmpC beta-lactamase. Structure (Camb) 10: 1013–1023. doi:10.1016/S0969-2126(02)00799-2
Zhao L, Brinton RD (2005) Structure-based virtual screening for plant-based ERbeta-selective ligands as potential preventative therapy against age-related neurodegenerative diseases. J Med Chem 48: 3463–3466. doi:10.1021/jm0490538
Zhou Y, Peng H, Ji Q, Qi J, Zhu Z, Yang C (2006) Discovery of small molecule inhibitors of integrin alphavbeta3 through structure-based virtual screening. Bioorg Med Chem Lett 16: 5878–5882. doi:10.1016/j.bmcl.2006.08.061
Villoutreix BO, Eudes R, Miteva MA (2009) Structure-based virtual ligand screening: recent success stories. Comb Chem High Throughput Screen 12: 1000–1016. doi:10.2174/138620709789824682
Park H, Lee J, Lee S (2006) Critical assessment of the automated AutoDock as a new docking tool for virtual screening. Proteins 65: 549–554. doi:10.1002/prot.21183
Wang R, Lu Y, Wang S (2003) Comparative evaluation of 11 scoring functions for molecular docking. J Med Chem 46: 2287–2303. doi:10.1021/jm0203783
Kohonen T (1982) Self-organized formation of topologically correct feature maps. Biol Cybern 43: 59–69. doi:10.1007/BF00337288
Ivanenkov YA, Savchuk NP, Ekins S, Balakin KV (2009) Computational mapping tools for drug discovery. Drug Discov Today 14: 767–775. doi:10.1016/j.drudis.2009.05.016
Zupan J., Gasteiger J. (1999) Neural networks in chemistry and drug design, 2nd edn. Weinheim, Wiley-VCH
Schneider P, Tanrikulu Y, Schneider G (2009) Self-organizing maps in drug discovery: compound library design, scaffold-hopping, repurposing. Curr Med Chem 16: 258–266. doi:10.2174/092986709787002655
Yan AX (2006) Application of self-organizing maps in compounds pattern recognition and combinatorial library design. Comb Chem High throughput Screen 9: 473–480. doi:10.2174/138620706777698562
Wang Z, Yan AX, Yuan QP (2009) Classification of blood-brain barrier permeation by Kohonen’s Self-Organizing Neural Network (KohNN) and Support Vector Machine (SVM). QSAR Comb Sci 28: 989–994. doi:10.1002/qsar.200960008
Wang Z, Yan AX, Yuan QP, Gasteiger J (2008) Explorations into modeling human oral bioavailability. Eur J Med Chem 43: 2442–2452. doi:10.1016/j.ejmech.2008.05.017
Yan AX, Wang Z, Cai ZY (2008) Prediction of human intestinal absorption by GA feature selection and support vector machine regression. Int J Mol Sci 9: 1961–1976. doi:10.3390/ijms9101961
Kaiser D, Terfloth L, Kopp S, Schulz J, de Laet R, Chiba P, Ecker GF, Gasteiger J (2007) Self-organizing maps for identification of new inhibitors of P-glycoprotein. J Med Chem 50: 1698–1702. doi:10.1021/jm060604z
Huang N, Shoichet BK, Irwin JJ (2006) Benchmarking sets for molecular docking. J Med Chem 49: 6789–6801. doi:10.1021/jm0608356
Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK (2007) BindingDB: a webaccessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res 35: D198–D201. doi:10.1093/nar/gkl999
Bush MR, Kaufman MD, Kennedy RM, Larsen SD, Trivedi BK, Song YT, Hutchings RH, Poel TJ (2005) Preparation of imidazole-based HMG-CoA reductase inhibitors. WO 2005079790
Cheng XM, Hutchings RH (2007) Preparation of imidazoles as HMG-CoA reductase inhibitors. WO 2007042910
Griffin J, Lanza G, Yu J (2007) Preparation of imidazoles as inhibitors of p38 MAP kinase and/or HMG - CoA reductase. WO 2007051065
Kennedy RM, Park WKC, Roth BD, Song YT, Trivedi BK (2005) Preparation of 7-(1-pyrrolyl)-3,5-dihydroxyheptanoic acid derivatives as HMG-CoA reductase inhibitors. US Patent 2005043364
Zhuge B, Fang HY, Yu H, Rao ZM, Shen W, Song J, Zhuge J (2008) Bioconversion of lovastatin to a novel statin by Amycolatopsis sp. Appl Microbiol Biotechnol 79: 209–216. doi:10.1007/s00253-008-1430-5
Scifinder, version 2007.1. Chemical Abstracts Service: Columbus, OH, 2007; RN 58-08-2. (Accessed 10 Oct 2010)
PubChem Bioassay AID1066. http://pubchem.ncbi.nlm.nih.gov/assay/assay.cgi?aid=1066
ADRIANA.Code, version 2.2, Molecular Networks GmbH, Erlangen, Germany. http://www.molecular-networks.com. (Accessed April 2010)
Ertl P, Rohde B, Selzer P (2000) Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug tansport properties. J Med Chem 43: 3714–3717. doi:10.1021/jm000942e
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23: 3–25. doi:10.1016/S0169-409X(96)00423-1
Miller KJ (1990) Additivity methods in molecular polarizability. J Am Chem Soc 112: 8533–8542. doi:10.1021/ja00179a044
Petitjean M (1992) Applications of the radius-diameter diagram to the classification of topological and geometrical shapes of chemical compounds. J Chem Inf Comput Sci 32: 331–337. doi:10.1021/ci00008a012
Volkenstein MV (1963) Configurational statistics of polymeric chains. Wiley, New York
Tanford C (1961) Physical chemistry of macromolecules. Wiley, New York
Moreau G, Broto P (1980) The autocorrelation of a topological structure: a new molecular descriptor. Nouv J Chim 4: 359–360
Wagener M, Sadowski J, Gasteiger J (1995) Autocorrelation of molecular surface properties for modeling corticosteroid binding globulin and cytosolic ah receptor activity by neural networks. J Am Chem Soc 117: 7769–7775. doi:10.1021/ja00134a023
Gasteiger J, Marsili M (1978) A new model for calculating atomic charges in molecules. Tetrahedron Lett 34: 3181–3184. doi:10.1016/S0040-4039(01)94977-9
Gasteiger J, Marsili M (1980) Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron 36: 3219–3228. doi:10.1016/0040-4020(80)80168-2
Gasteiger J, Hutchings MG (1984) Quantitative models of gas-phase proton transfer reaction involving alcohols, ethers and their thio analogs. correlation analyses based on residual electronegativity and effective polarizability. J Am Chem Soc 106: 6489–6495. doi:10.1021/ja00334a006
Kohonen T. (1989) Self-organization and associative memory. Springer, Berlin
SONNIA can be obtained from Molecular Networks GmbH, Erlangen, Germany. http://www.molecular-networks.com. (Accessed April 2010)
Rodgers JL, Nicewander WA (1988) Thirteen ways to look at the correlation coefficient. Am Stat 42: 59–66. doi:10.2307/2685263
Gasteiger J, Teckentrup A, Terfloth L, Spycher S (2003) Neural networks as data mining tools in drug design. J Phys Org Chem 16: 232–245. doi:10.1002/poc.597
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
The Below is the Electronic Supplementary Material.
Rights and permissions
About this article
Cite this article
Wang, Z., Yan, A. Discriminating of HMG-CoA reductase inhibitors and decoys using self-organizing maps. Mol Divers 15, 655–663 (2011). https://doi.org/10.1007/s11030-010-9288-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-010-9288-8