Abstract
Introduction
Epigenetic modifications have been reported to play an important role in regulating gene expression and these modifications become critical when they have a role in controlling another important layer of epigenetic regulation namely microRNAs. In the present study, we have identified the microRNAs that may be regulated by promoter DNA methylation and histone acetylation in Human papilloma virus-positive head and neck squamous cell carcinoma.
Methodology
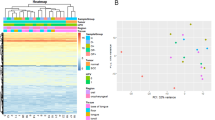
HPV-negative cell line (UPCI:SCC-116) and HPV-16 +ve cell line (UPCI:SCC-090) were treated with methylation inhibitor (5-aza-2′-deoxycytidine, AZA) and acetylation inhibitor (Trichostatin-A, TSA), followed by micro-array analysis. The differentially expressed miRNAs were validated in control (n = 10), HPV-16 +ve (n = 30), and HPV −ve (n = 30) HNC, TCGA (n = 529) tissue samples, and two HPV −ve (SCC116 and Hacat) and two HPV +ve (SCC090 and SiHa) cell lines. Methylation-specific PCR (MSP) and chromatin immunoprecipitation assay (CHIP) were performed to validate their regulation. In silico and in vitro analyses of identified miRNAs were done to study putative pathways they target and their possible role in carcinogenesis.
Results
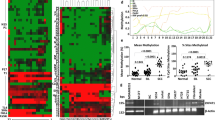
Among 10 miRNAs specifically up-regulated in microarray analysis of AZA-treated SCC090 cells, we observed significantly decreased expression of hsa-miR-181c-5p, hsa-miR-132-5p, hsa-miR-658 in HPV +ve HNC cohort, TCGA tissue samples, and cell lines as compared to their HPV −ve counterpart, and their promoter region also possesses CpG islands. MSP and analysis of TCGA data (MethHC) revealed increased frequency of methylation at the promoter of hsa-miR-132-5p that is negatively correlated with its expression. In TSA-treated SCC090 cells, out of 7 miRNAs, two namely Hsa-miR-129-2-3p and Hsa-miR-449a were found to be up-regulated as compared to HPV −ve cells. However, the levels of enrichment by anti-acetyl-H3 and anti-acetyl-H4 were significantly low in cell lines compared to respective controls and both were up-regulated in HPV +ve compared to HPV −ve TCGA tissue samples. In silico analysis revealed hsa-miR-132-5p targeted canonical β-catenin/wnt pathway and modulation of down-stream genes of the pathway was observed on over-expression/inhibition of hsa-miR-132-5p.
Conclusion
This study suggests the role of epigenetic modifications in regulating expression of miRNAs in HPV +ve HNSCC.





Similar content being viewed by others
References
zur Hausen H (2002) Papillomaviruses and cancer: from basic studies to clinical application. Nat Rev Cancer 2:342–350. https://doi.org/10.1038/nrc798
Bouvard V, Baan R, Straif K, Grosse Y, Secretan B, El Ghissassi F, Benbrahim-Tallaa L, Guha N, Freeman C, Galichet L, Cogliano V (2009) A review of human carcinogens—part B: biological agents. Lancet Oncol 10:321–322
Chang SS, Jiang WW, Smith I, Poeta LM, Begum S, Glazer C, Shan S, Westra W, Sidransky D, Califano JA (2008) MicroRNA alterations in head and neck squamous cell carcinoma. Int J Cancer 123:2791–2797. https://doi.org/10.1002/ijc.23831
Lajer CB, Garnaes E, Friis-Hansen L, Norrild B, Therkildsen MH, Glud M, Rossing M, Lajer H, Svane D, Skotte L, Specht L, Buchwald C, Nielsen FC (2012) The role of miRNAs in human papilloma virus (HPV)-associated cancers: bridging between HPV-related head and neck cancer and cervical cancer. Br J Cancer 106:1526–1534. https://doi.org/10.1038/bjc.2012.109
Wald AI, Hoskins EE, Wells SI, Ferris RL, Khan SA (2011) Alteration of microRNA profiles in squamous cell carcinoma of the head and neck cell lines by human papillomavirus. Head Neck 33:504–512. https://doi.org/10.1002/hed.21475
Lajer CB, Nielsen FC, Friis-Hansen L, Norrild B, Borup R, Garnaes E, Rossing M, Specht L, Therkildsen MH, Nauntofte B, Dabelsteen S, von Buchwald C (2011) Different miRNA signatures of oral and pharyngeal squamous cell carcinomas: a prospective translational study. Br J Cancer 104:830–840. https://doi.org/10.1038/bjc.2011.29
Jimenez-Wences H, Peralta-Zaragoza O, Fernandez-Tilapa G (2014) Human papilloma virus, DNA methylation and microRNA expression in cervical cancer (Review). Oncol Rep 31:2467–2476. https://doi.org/10.3892/or.2014.3142
Wilting SM, Verlaat W, Jaspers A, Makazaji NA, Agami R, Meijer CJ, Snijders PJ, Steenbergen RD (2013) Methylation-mediated transcriptional repression of microRNAs during cervical carcinogenesis. Epigenetics 8:220–228. https://doi.org/10.4161/epi.23605
Wilting SM, Miok V, Jaspers A, Boon D, Sorgard H, Lando M, Snoek BC, van Wieringen WN, Meijer CJ, Lyng H, Snijders PJ, Steenbergen RD (2016) Aberrant methylation-mediated silencing of microRNAs contributes to HPV-induced anchorage independence. Oncotarget 7:43805–43819. https://doi.org/10.18632/oncotarget.9698
Au Yeung CL, Tsang WP, Tsang TY, Co NN, Yau PL, Kwok TT (2010) HPV-16 E6 upregulation of DNMT1 through repression of tumor suppressor p53. Oncol Rep 24:1599–1604
Botezatu A, Goia-Rusanu CD, Iancu IV, Huica I, Plesa A, Socolov D, Ungureanu C, Anton G (2011) Quantitative analysis of the relationship between microRNA124a, -34b and – 203 gene methylation and cervical oncogenesis. Mol Med Rep 4:121–128. https://doi.org/10.3892/mmr.2010.394
Melar-New M, Laimins LA (2010) Human papillomaviruses modulate expression of microRNA 203 upon epithelial differentiation to control levels of p63 proteins. J Virol 84:5212–5221. https://doi.org/10.1128/JVI.00078-10
Nuovo GJ, Wu X, Volinia S, Yan F, di Leva G, Chin N, Nicol AF, Jiang J, Otterson G, Schmittgen TD, Croce C (2010) Strong inverse correlation between microRNA-125b and human papillomavirus DNA in productive infection. Diagn Mol Pathol 19:135–143. https://doi.org/10.1097/PDM.0b013e3181c4daaa
Jung HM, Phillips BL, Chan EK (2014) miR-375 activates p21 and suppresses telomerase activity by coordinately regulating HPV E6/E7, E6AP, CIP2A, and 14-3-3zeta. Mol Cancer 13:80. https://doi.org/10.1186/1476-4598-13-80
Sannigrahi MK, Sharma R, Singh V, Panda NK, Rattan V, Khullar M (2017) Role of host miRNA Hsa-miR-139-3p in HPV-16-induced carcinomas. Clin Cancer Res 23:3884–3895. https://doi.org/10.1158/1078-0432.CCR-16-2936
White JS, Weissfeld JL, Ragin CC, Rossie KM, Martin CL, Shuster M, Ishwad CS, Law JC, Myers EN, Johnson JT, Gollin SM (2007) The influence of clinical and demographic risk factors on the establishment of head and neck squamous cell carcinoma cell lines. Oral Oncol 43:701–712. https://doi.org/10.1016/j.oraloncology.2006.09.001
Chakrabarti O, Veeraraghavalu K, Tergaonkar V, Liu Y, Androphy EJ, Stanley MA, Krishna S (2004) Human papillomavirus type 16 E6 amino acid 83 variants enhance E6-mediated MAPK signaling and differentially regulate tumorigenesis by notch signaling and oncogenic Ras. J Virol 78:5934–5945. https://doi.org/10.1128/JVI.78.11.5934-5945.2004
Chacon-Cortes D, Haupt LM, Lea RA, Griffiths LR (2012) Comparison of genomic DNA extraction techniques from whole blood samples: a time, cost and quality evaluation study. Mol Biol Rep 39:5961–5966. https://doi.org/10.1007/s11033-011-1408-8
Sannigrahi MK, Singh V, Sharma R, Panda NK, Radotra BD, Khullar M (2016) Detection of active human papilloma virus-16 in head and neck cancers of Asian North Indian patients. Oral Dis 22:62–68. https://doi.org/10.1111/odi.12382
Agarwal V, Bell GW, Nam JW, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife 4:e05005. https://doi.org/10.7554/eLife.05005
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, Lim B, Rigoutsos I (2006) A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell 126:1203–1217. https://doi.org/10.1016/j.cell.2006.07.031
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pages F, Trajanoski Z, Galon J (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Hsu JB, Chiu CM, Hsu SD, Huang WY, Chien CH, Lee TY, Huang HD (2011) miRTar: an integrated system for identifying miRNA-target interactions in human. BMC Bioinform 12:300. https://doi.org/10.1186/1471-2105-12-300
Sylvester PW (2011) Optimization of the tetrazolium dye (MTT) colorimetric assay for cellular growth and viability. Methods Mol Biol 716:157–168. https://doi.org/10.1007/978-1-61779-012-6_9
Pozarowski P, Darzynkiewicz Z (2004) Analysis of cell cycle by flow cytometry. Methods Mol Biol 281:301–311. https://doi.org/10.1385/1-59259-811-0:301
Liang CC, Park AY, Guan JL (2007) In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc 2:329–333. https://doi.org/10.1038/nprot.2007.30
Papillon-Cavanagh S, Lu C, Gayden T, Mikael LG, Bechet D, Karamboulas C, Ailles L, Karamchandani J, Marchione DM, Garcia BA, Weinreb I, Goldstein D, Lewis PW, Dancu OM, Dhaliwal S, Stecho W, Howlett CJ, Mymryk JS, Barrett JW, Nichols AC, Allis CD, Majewski J, Jabado N (2017) Impaired H3K36 methylation defines a subset of head and neck squamous cell carcinomas. Nat Genet 49:180–185. https://doi.org/10.1038/ng.3757
Huang WY, Hsu SD, Huang HY, Sun YM, Chou CH, Weng SL, Huang HD (2015) MethHC: a database of DNA methylation and gene expression in human cancer. Nucleic Acids Res 43:D856–D861. https://doi.org/10.1093/nar/gku1151
Backes C, Kehl T, Stockel D, Fehlmann T, Schneider L, Meese E, Lenhof HP, Keller A (2017) miRPathDB: a new dictionary on microRNAs and target pathways. Nucleic Acids Res 45:D90–D96. https://doi.org/10.1093/nar/gkw926
Khamas A, Ishikawa T, Shimokawa K, Mogushi K, Iida S, Ishiguro M, Mizushima H, Tanaka H, Uetake H, Sugihara K (2012) Screening for epigenetically masked genes in colorectal cancer using 5-aza-2′-deoxycytidine, microarray and gene expression profile. Cancer Genom Proteom 9:67–75
Schmelz K, Sattler N, Wagner M, Lubbert M, Dorken B, Tamm I (2005) Induction of gene expression by 5-aza-2′-deoxycytidine in acute myeloid leukemia (AML) and myelodysplastic syndrome (MDS) but not epithelial cells by DNA-methylation-dependent and -independent mechanisms. Leukemia 19:103–111. https://doi.org/10.1038/sj.leu.2403552
Liang G, Gonzales FA, Jones PA, Orntoft TF, Thykjaer T (2002) Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res 62:961–966
Basu B, Chakraborty J, Chandra A, Katarkar A, Baldevbhai JR, Dhar Chowdhury D, Ray JG, Chaudhuri K, Chatterjee R (2017) Genome-wide DNA methylation profile identified a unique set of differentially methylated immune genes in oral squamous cell carcinoma patients in India. Clin Epigenetics 9:13. https://doi.org/10.1186/s13148-017-0314-x
Sartor MA, Dolinoy DC, Jones TR, Colacino JA, Prince ME, Carey TE, Rozek LS (2011) Genome-wide methylation and expression differences in HPV(+) and HPV(-) squamous cell carcinoma cell lines are consistent with divergent mechanisms of carcinogenesis. Epigenetics 6:777–787
Liu Y, Li Y, Liu J, Wu Y, Zhu Q (2015) MicroRNA-132 inhibits cell growth and metastasis in osteosarcoma cell lines possibly by targeting Sox4. Int J Oncol 47:1672–1684. https://doi.org/10.3892/ijo.2015.3147
Liu K, Li X, Cao Y, Ge Y, Wang J, Shi B (2015) MiR-132 inhibits cell proliferation, invasion and migration of hepatocellular carcinoma by targeting PIK3R3. Int J Oncol 47:1585–1593. https://doi.org/10.3892/ijo.2015.3112
Liu X, Yan S, Pei C, Cui Y (2015) Decreased microRNA-132 and its function in human non-small cell lung cancer. Mol Med Rep 11:3601–3608. https://doi.org/10.3892/mmr.2015.3222
Formosa A, Lena AM, Markert EK, Cortelli S, Miano R, Mauriello A, Croce N, Vandesompele J, Mestdagh P, Finazzi-Agro E, Levine AJ, Melino G, Bernardini S, Candi E (2013) DNA methylation silences miR-132 in prostate cancer. Oncogene 32:127–134. https://doi.org/10.1038/onc.2012.14
Zhang S, Hao J, Xie F, Hu X, Liu C, Tong J, Zhou J, Wu J, Shao C (2011) Downregulation of miR-132 by promoter methylation contributes to pancreatic cancer development. Carcinogenesis 32:1183–1189. https://doi.org/10.1093/carcin/bgr105
Iwai S, Yonekawa A, Harada C, Hamada M, Katagiri W, Nakazawa M, Yura Y (2010) Involvement of the Wnt-beta-catenin pathway in invasion and migration of oral squamous carcinoma cells. Int J Oncol 37:1095–1103
Chang HW, Roh JL, Jeong EJ, Lee SW, Kim SW, Choi SH, Park SK, Kim SY (2008) Wnt signaling controls radiosensitivity via cyclooxygenase-2-mediated Ku expression in head and neck cancer. Int J Cancer 122:100–107. https://doi.org/10.1002/ijc.23069
Bello JO, Nieva LO, Paredes AC, Gonzalez AM, Zavaleta LR, Lizano M (2015) Regulation of the Wnt/beta-catenin signaling pathway by human papillomavirus E6 and E7 oncoproteins. Viruses 7:4734–4755. https://doi.org/10.3390/v7082842
Uren A, Fallen S, Yuan H, Usubutun A, Kucukali T, Schlegel R, Toretsky JA (2005) Activation of the canonical Wnt pathway during genital keratinocyte transformation: a model for cervical cancer progression. Cancer Res 65:6199–6206. https://doi.org/10.1158/0008-5472.CAN-05-0455
Bulut G, Fallen S, Beauchamp EM, Drebing LE, Sun J, Berry DL, Kallakury B, Crum CP, Toretsky JA, Schlegel R, Uren A (2011) Beta-catenin accelerates human papilloma virus type-16 mediated cervical carcinogenesis in transgenic mice. PLoS ONE 6:e27243. https://doi.org/10.1371/journal.pone.0027243
Rampias T, Boutati E, Pectasides E, Sasaki C, Kountourakis P, Weinberger P, Psyrri A (2010) Activation of Wnt signaling pathway by human papillomavirus E6 and E7 oncogenes in HPV16-positive oropharyngeal squamous carcinoma cells. Mol Cancer Res 8:433–443. https://doi.org/10.1158/1541-7786.MCR-09-0345
Acknowledgements
This study was carried out at Genetics of Complex disorder laboratory, PGIMER, Chandigarh, India. UPCI:SCC-116 and UPCI:SCC-090 cell lines were kind gifts from Dr S Gollin (University of Pittsburgh, PA, USA). The authors are grateful to all individuals who participated in this study and the laboratory staff members for their skillful technical help. The work has been presented in various national and international conferences. Malay K Sannigrahi thankfully acknowledges research fellowship from Indian Council of Medical Research, New Delhi, India. Dr. Rajni Sharma thankfully acknowledges fellowship from DBT, India (BT/Bio-CARe/08/837/2010–2011). We sincerely thank and greatly appreciate the two anonymous reviewers for their comments and suggestions for improving the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interest
We declare no conflict of interest.
Ethical approval
Approved by Institutional Ethics Committee (IEC), Post Graduate Institute of Medical Education and Research (PGIMER), Chandigarh, India.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sannigrahi, M.K., Sharma, R., Singh, V. et al. DNA methylation regulated microRNAs in HPV-16-induced head and neck squamous cell carcinoma (HNSCC). Mol Cell Biochem 448, 321–333 (2018). https://doi.org/10.1007/s11010-018-3336-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-018-3336-6