Abstract
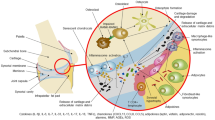
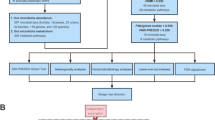
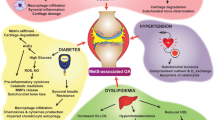
Inflammation is associated with glenohumeral arthritis and rotator cuff tendon tears. Epigenetically, miRNAs tightly regulate various genes involved in the inflammatory response. Alterations in the expression profile of miRNAs and the elucidation of their target genes with respect to the pathophysiology could improve the understanding of their regulatory role and therapeutic potential. Here, we screened key miRNAs that mediate inflammation and linked with JAK2/STAT3 pathway with respect to the coincidence of glenohumeral arthritis in patients suffering from rotator cuff injury (RCI). Human resected long head of the biceps tendons were examined for miRNA profile from two groups of patients: Group 1 included the patients with glenohumeral arthritis and massive rotator cuff tears and the Group 2 patients did not have arthritis or rotator cuff tears. The miRNA profiling revealed that 235 miRNAs were highly altered (fold change less than −3 and greater than +2 were considered). Data from the NetworkAnalyst program revealed the involvement and interaction between 3,430 different genes associated with inflammation out of which 284 genes were associated with JAK2/STAT3 pathway and interconnect 120 different pathways of inflammation. Around 1,500 miRNAs were found to play regulatory role associated with these genes of inflammatory responses and 77 miRNAs were found to regulate more than 10 genes. Among them, 25 genes with less than tenfold change were taken to consideration which altogether constitute for the regulation of 102 genes. Targeting these miRNAs and the underlying regulatory mechanisms may advance our knowledge to develop promising therapies in the management of shoulder tendon pathology.




Similar content being viewed by others
Abbreviations
- DAPI:
-
4′,6-Diamidino-2-phenylindole
- ECM:
-
Extracellular matrix
- JAK:
-
Janus-activated kinase
- miRNA:
-
MicroRNA
- PBS:
-
Phosphate buffered saline
- RCI:
-
Rotator cuff injury
- RIN:
-
RNA integrity number
- STAT:
-
Signal transducers and activators of transcription
References
Thankam FG, Dilisio MF, Agrawal DK (2016) Immunobiological factors aggravating the fatty infiltration on tendons and muscles in rotator cuff lesions. Mol Cell Biochem. doi:10.1007/s11010-016-2710-5
Pandey V, Jaap Willems W (2015) Rotator cuff tear: a detailed update. Asia-Pac J Sports Med Arthrosc Rehabil Technol 2:1–14. doi:10.1016/j.asmart.2014.11.003
Itoi E (2013) Rotator cuff tear: physical examination and conservative treatment. J Orthop Sci 18:197–204. doi:10.1007/s00776-012-0345-2
Thankam FG, Dilisio MF, Dougherty KA et al (2016) Triggering receptor expressed on myeloid cells and 5′adenosine monophosphate-activated protein kinase in the inflammatory response: a potential therapeutic target. Expert Rev Clin Immunol. doi:10.1080/1744666X.2016.1196138
Brooks P (2003) Inflammation as an important feature of osteoarthritis. Bull World Health Organ 81:689–690
Macaulay AA, Greiwe RM, Bigliani LU (2010) Rotator cuff deficient arthritis of the glenohumeral joint. Clin Orthop Surg 2:196. doi:10.4055/cios.2010.2.4.196
Smith AM (2005) Rotator cuff repair in patients with rheumatoid arthritis. J Bone Jt Surg Am 87:1782. doi:10.2106/JBJS.D.02452
Kaplan MH (2013) STAT signaling in inflammation. JAK-STAT 2:e24198. doi:10.4161/jkst.24198
Murray PJ (2007) The JAK-STAT signaling pathway: input and output integration. J Immunol 178:2623–2629. doi:10.4049/jimmunol.178.5.2623
Harrison DA (2012) The JAK/STAT pathway. Cold Spring Harb Perspect Biol 4:a011205. doi:10.1101/cshperspect.a011205
Roy S, Benz F, Luedde T, Roderburg C (2015) The role of miRNAs in the regulation of inflammatory processes during hepatofibrogenesis. Hepatobiliary Surg Nutr 4:24–33. doi:10.3978/j.issn.2304-3881.2015.01.05
Essandoh K, Li Y, Huo J, Fan G-C (2016) MiRNA-mediated macrophage polarization and its potential role in the regulation of inflammatory response. Shock 46:122–131. doi:10.1097/SHK.0000000000000604
Baumjohann D, Ansel KM (2013) MicroRNA-mediated regulation of T helper cell differentiation and plasticity. Nat Rev Immunol 13:666–678. doi:10.1038/nri3494
He X, Jing Z, Cheng G (2014) MicroRNAs: new regulators of toll-like receptor signalling pathways. Biomed Res Int 2014:1–14. doi:10.1155/2014/945169
Amado T, Schmolka N, Metwally H et al (2015) Cross-regulation between cytokine and microRNA pathways in T cells: highlights-Review. Eur J Immunol 45:1584–1595. doi:10.1002/eji.201545487
Olarerin-George AO, Anton L, Hwang Y-C et al (2013) A functional genomics screen for microRNA regulators of NF-kappaB signaling. BMC Biol 11:19. doi:10.1186/1741-7007-11-19
Li H, Chen X, Guan L et al (2013) MiRNA-181a regulates adipogenesis by targeting tumor necrosis factor-Î ± (TNF-Î ±) in the Porcine model. PLoS ONE 8:e71568. doi:10.1371/journal.pone.0071568
Sun Y, Qin Z, Li Q et al (2016) MicroRNA-124 negatively regulates LPS-induced TNF-α production in mouse macrophages by decreasing protein stability. Acta Pharmacol Sin 37:889–897. doi:10.1038/aps.2016.16
Millar NL, Wei AQ, Molloy TJ et al (2009) Cytokines and apoptosis in supraspinatus tendinopathy. J Bone J 91:417–424. doi:10.1302/0301-620X.91B3.21652
Jin-fang Zhang JG, Kai-ming Chan GL (2015) A mini-review: microRNA in Tendon Injuries. J Stem Cell Res Ther. doi:10.4172/2157-7633.1000303
Gupta GK, Agrawal T, DelCore MG et al (2012) Vitamin D deficiency induces cardiac hypertrophy and inflammation in epicardial adipose tissue in hypercholesterolemic swine. Exp Mol Pathol 93:82–90. doi:10.1016/j.yexmp.2012.04.006
Fischer AH, Jacobson KA, Rose J, Zeller R (2008) Hematoxylin and eosin staining of tissue and cell sections. Cold Spring Harb Protoc. doi:10.1101/pdb.prot4986
Fong G, Backman LJ, Hart DA et al (2013) Substance P enhances collagen remodeling and MMP-3 expression by human tenocytes. J Orthop Res 31:91–98. doi:10.1002/jor.22191
Agrawal T, Gupta GK, Agrawal DK (2013) Vitamin D supplementation reduces airway hyperresponsiveness and allergic airway inflammation in a murine model. Clin Exp Allergy. doi:10.1111/cea.12102
Thankam FG, Dilisio MF, Dietz NE, Agrawal DK (2016) TREM-1, HMGB1 and RAGE in the shoulder tendon: dual mechanisms for inflammation based on the coincidence of glenohumeral arthritis. PLoS ONE 11:e0165492. doi:10.1371/journal.pone.0165492
Millar NL, Gilchrist DS, Akbar M et al (2015) MicroRNA29a regulates IL-33-mediated tissue remodelling in tendon disease. Nat Commun 6:6774. doi:10.1038/ncomms7774
Thankam FG, Boosani CS, Dilisio MF et al (2016) MicroRNAs associated with shoulder tendon matrisome disorganization in glenohumeral arthritis. PLoS ONE 11:e0168077. doi:10.1371/journal.pone.0168077
Xia J, Benner MJ, Hancock REW (2014) NetworkAnalyst—integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res 42:W167–W174. doi:10.1093/nar/gku443
Tedgui A (2006) Cytokines in atherosclerosis: pathogenic and regulatory pathways. Physiol Rev 86:515–581. doi:10.1152/physrev.00024.2005
Abate M, Gravare-Silbernagel K, Siljeholm C et al (2009) Pathogenesis of tendinopathies: inflammation or degeneration? Arthritis Res Ther 11:235. doi:10.1186/ar2723
Millar NL, Hueber AJ, Reilly JH et al (2010) Inflammation is present in early human tendinopathy. Am J Sports Med 38:2085–2091. doi:10.1177/0363546510372613
Dakin SG, Dudhia J, Smith RKW (2014) Resolving an inflammatory concept: the importance of inflammation and resolution in tendinopathy. Vet Immunol Immunopathol 158:121–127. doi:10.1016/j.vetimm.2014.01.007
Yang X, He G, Hao Y et al (2010) The role of the JAK2-STAT3 pathway in pro-inflammatory responses of EMF-stimulated N9 microglial cells. J Neuroinflammation 7:54. doi:10.1186/1742-2094-7-54
Gao W, McCormick J, Connolly M et al (2015) Hypoxia and STAT3 signalling interactions regulate pro-inflammatory pathways in rheumatoid arthritis. Ann Rheum Dis 74:1275–1283. doi:10.1136/annrheumdis-2013-204105
Yuan J, Zhang F, Niu R (2015) Multiple regulation pathways and pivotal biological functions of STAT3 in cancer. Sci Rep 5:17663. doi:10.1038/srep17663
Starczynowski DT, Kuchenbauer F, Argiropoulos B et al (2010) Identification of miR-145 and miR-146a as mediators of the 5q-syndrome phenotype. Nat Med 16:49–58. doi:10.1038/nm.2054
Jiang Y, Zhang M, He H et al (2013) MicroRNA/mRNA profiling and regulatory network of intracranial aneurysm. BMC Med Genomics. doi:10.1186/1755-8794-6-36
Wu R-Q, Zhang D-F, Tu E et al (2014) The mucosal immune system in the oral cavity—an orchestra of T cell diversity. Int J Oral Sci 6:125–132. doi:10.1038/ijos.2014.48
Luo X, Ranade K, Talker R et al (2013) microRNA-mediated regulation of innate immune response in rheumatic diseases. Arthritis Res Ther 15:210. doi:10.1186/ar4194
Kolenda T, Przybyła W, Teresiak A et al (2014) The mystery of let-7d—a small RNA with great power. Współczesna Onkol 5:293–301. doi:10.5114/wo.2014.44467
Kutty RK, Nagineni CN, Samuel W et al (2013) Differential regulation of microRNA-146a and microRNA-146b-5p in human retinal pigment epithelial cells by interleukin-1β, tumor necrosis factor-α, and interferon-γ. Mol Vis 19:737–750
Ye E-A, Steinle JJ (2016) miR-146a attenuates inflammatory pathways mediated by TLR4/NF- κ B and TNF α to protect primary human retinal microvascular endothelial cells grown in high glucose. Mediat Inflamm 2016:1–9. doi:10.1155/2016/3958453
Xing Y, Fu J, Yang H et al (2015) MicroRNA expression profiles and target prediction in neonatal Wistar rat lungs during the development of bronchopulmonary dysplasia. Int J Mol Med. doi:10.3892/ijmm.2015.2347
Punga T, Le Panse R, Andersson M et al (2014) Circulating miRNAs in myasthenia gravis: miR-150-5p as a new potential biomarker. Ann Clin Transl Neurol 1:49–58. doi:10.1002/acn3.24
Sang W, Wang Y, Zhang C et al (2016) MiR-150 impairs inflammatory cytokine production by targeting ARRB-2 after blocking CD28/B7 costimulatory pathway. Immunol Lett 172:1–10. doi:10.1016/j.imlet.2015.11.001
Xie W, Li M, Xu N et al (2013) miR-181a regulates inflammation responses in monocytes and macrophages. PLoS ONE 8:e58639. doi:10.1371/journal.pone.0058639
Reynoso R, Laufer N, Hackl M et al (2014) MicroRNAs differentially present in the plasma of HIV elite controllers reduce HIV infection in vitro. Sci Rep. doi:10.1038/srep05915
Tonevitsky AG, Maltseva DV, Abbasi A et al (2013) Dynamically regulated miRNA-mRNA networks revealed by exercise. BMC Physiol 13:9. doi:10.1186/1472-6793-13-9
Kaukoniemi KM, Rauhala HE, Scaravilli M et al (2015) Epigenetically altered miR-193b targets cyclin D1 in prostate cancer. Cancer Med 4:1417–1425. doi:10.1002/cam4.486
Takulla NF, Wolff MS, Schenkel AB (2012) Caries management by risk assessment. N Y State Dent J 78:41–45
Yan J-K, Wang W-Q, Ma H-L, Wu J-Y (2012) Sulfation and enhanced antioxidant capacity of an exopolysaccharide produced by the medicinal fungus Cordyceps sinensis. Molecules 18:167–177. doi:10.3390/molecules18010167
Li J, Wan Y, Guo Q et al (2010) Altered microRNA expression profile with miR-146a upregulation in CD4+ T cells from patients with rheumatoid arthritis. Arthritis Res Ther 12:R81. doi:10.1186/ar3006
Zhai Y, Wang M, Zhang Q, Wang J (2015) MicroRNA-498 is downregulated in non-small cell lung cancer and correlates with tumor progression. J Cancer Res Ther 11:107. doi:10.4103/0973-1482.163859
Jin H, Li C, Ge H et al (2013) Circulating microRNA: a novel potential biomarker for early diagnosis of intracranial aneurysm rupture a case control study. J Transl Med 11:296. doi:10.1186/1479-5876-11-296
Sánchez-Jiménez C, Carrascoso I, Barrero J, Izquierdo JM (2013) Identification of a set of miRNAs differentially expressed in transiently TIA-depleted HeLa cells by genome-wide profiling. BMC Mol Biol 14:4. doi:10.1186/1471-2199-14-4
Xu K, Liang X, Shen K et al (2012) miR-297 modulates multidrug resistance in human colorectal carcinoma by down-regulating MRP-2. Biochem J 446:291–300. doi:10.1042/BJ20120386
Acknowledgements
This work was supported by research grants R01 HL112597, R01 HL116042, and R01 HL120659 to DK Agrawal from the National Heart, Lung and Blood Institute, National Institutes of Health, USA, and Creighton University LB692 Grant to MF Dilisio from the State of Nebraska. The content of this original research article is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or the State of Nebraska.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thankam, F.G., Boosani, C.S., Dilisio, M.F. et al. MicroRNAs associated with inflammation in shoulder tendinopathy and glenohumeral arthritis. Mol Cell Biochem 437, 81–97 (2018). https://doi.org/10.1007/s11010-017-3097-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-017-3097-7