Abstract
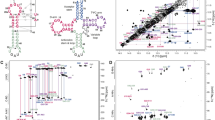
Three-dimensional atomic models of complexes between yeast tRNAPhe and 10- or 15-mer oligonucleotides complementary to the 3′-terminal tRNA sequence have been constructed using computer modeling. It has been found that rapidly formed primary complexes appear when an oligonucleotide binds to the coaxial acceptor and T stems of the tRNAPhe along the major groove, which results in the formation of a triplex. Long stems allow the formation of a sufficiently strong complex with the oligonucleotide, which delivers its 3′-terminal nucleotides to the vicinity of the T loop adjoining the stem. These nucleotides destabilize the loop structure and initiate conformational rearrangements involving local tRNAPhe destruction and formation of the final tRNAPhe-oligonucleotide complementary complex. The primary complex formation and the following tRNAPhe destruction constitute the “molecular wedge” mechanism. An effective antisence oligonucleotide should consist of three segments—(1) complex initiator, (2) primary complex stabilizer, and (3) loop destructor—and be complementary to the (free end)/loop-stem-loop tRNA structural element.
Similar content being viewed by others
REFERENCES
Branch A.D. 1998. A good antisense molecule is hard to find. Trends Biochem. Sci. 23, 45–50.
Mathews D.H., Burkard M.E., Frier S.M., Wyatt J.R. Turner D.H. 1999. Predicting oligonucleotide affinity to nucleic acid targets. RNA. 5, 1458–1469.
Petyuk V.A., Zenkova M.A., Giedge R., Vlasov V.V. 1999. Hybridization of antisense oligonucleotides with 3′ part of tRNAphe. FEBS Lett. 444, 217–221.
Petyuk V.A., Giedge R., Vlasov V.V., Zenkova M.A. 2000. Mechanism of oligonucleotide hybridization with the 3′-terminal region of yeast tRNAPhe. Mol. Biol. 234, 879–886.
Frenkel D., Smith B. 1996. Understanding Molecular Simulation. From Algorithms to Applications. Oxford: Academic Press.
Schlick T. 2000. Molecular Modeling and Simulations: An Interdisciplinary Guide. N.Y.: Springer.
AMBER Home page: http://www.amber.ucsf.edu/amber/index.html.
GROMOS96 Home page: http://www.igc.ethz.ch/gromos/.
Cornell W.D., Cieplak P., Bayly C.I., Gould I.R., Merz K.M., Ferguson D.M., Spellmeyer D.C., Fox T., Caldwell J.W., Kollman P.A. 1995. A second generation force field for simulation of proteins, nucleic acids and organic molecules. J. Am. Chem. Soc. 117, 5179–5197.
Lazaridis T., Karplus M. 1999. Effective energy function for protein in solution. Proteins. 35, 133–152.
Park B., Levitt M. 1996. Energy functions that discriminate X-ray and near native folds from well constructed decoys. J. Mol. Biol. 258, 367–392; http://dd.stanford.edu.
Zanger W. 1987. Principles of Structural Organization of Nucleic Acids [Russian translation]. Moscow: Mir.
Leontis N.B., Stombaugh J., Westhof E. 2002. The non-Watson-Crick base pairs and their associated isostericity matrices. Nucleic Acids Res. 30, 3497–3531.
Cantor C.R., Schimmel P.R. 1980. Biophysical Chemistry: III. The Behavior of Biological Macromolecules. San Francisco: W. H. Freeman.
Sugimoto N., Nakano S., Katoh M., Matsumura A., Nakamura H., Ohmichi T., Yoneyama M., Sasaki M. 1995. Thermodynamic parameters to predict stability of RNA/DNA hybrid duplexes. Biochemistry. 34, 11 211–11 216.
Alberti P., Arimondo P.B., Mergny J., Garestier T. Helene C., Sun J.-S. 2002. A directional-zipping mechanism for triple helix formation. Nucleic Acids Res. 30, 5407–5415.
Zurkin V.B., Raghunathan G., Ulyanov N.B., Jernigan R.L. 1994. A parallel DNA triplex as model for the intermediate in homologous recombination. J. Mol. Biol. 239, 181–200.
Shchyolkina A.K., Borisova O.F. 1997. Stabilizing and destabilizing effects of intercalators on DNA triplexes. FEBS Lett. 419, 27–31.
Gilson M.K., Given, J.A., Bush B.L., McCammon J.A. 1997. The statistical-thermodynamic basis for computation of binding affinities: A critical review. Biophys. J. 72, 1047–1069.
Author information
Authors and Affiliations
Additional information
__________
Translated from Molekulyarnaya Biologiya, Vol. 39, No. 5, 2005, pp. 887–895.
Original Russian Text Copyright © 2005 by Vorobjev.
Rights and permissions
About this article
Cite this article
Vorobjev, Y.N. Study of the Mechanism of Interaction of Oligonucleotides with the 3′-Terminal Region of tRNAPhe by Computer Modeling. Mol Biol 39, 777–784 (2005). https://doi.org/10.1007/s11008-005-0093-x
Received:
Issue Date:
DOI: https://doi.org/10.1007/s11008-005-0093-x