Abstract
Stochastic models that incorporate birth, death and immigration (also called birth–death and innovation models) are ubiquitous and applicable to many problems such as quantifying species sizes in ecological populations, describing gene family sizes, modeling lymphocyte evolution in the body. Many of these applications involve the immigration of new species into the system. We consider the full high-dimensional stochastic process associated with multispecies birth–death–immigration and present a number of exact and asymptotic results at steady state. We further include random mutations or interactions through a carrying capacity and find the statistics of the total number of individuals, the total number of species, the species size distribution, and various diversity indices. Our results include a rigorous analysis of the behavior of these systems in the fast immigration limit which shows that of the different diversity indices, the species richness is best able to distinguish different types of birth–death–immigration models. We also find that detailed balance is preserved in the simple noninteracting birth–death–immigration model and the birth–death–immigration model with carrying capacity implemented through death. Surprisingly, when carrying capacity is implemented through the birth rate, detailed balance is violated.





Similar content being viewed by others
References
Allen, L.J.S.: An Introduction to Stochastic Processes with Applications to Biology. Taylor and Francis, Boca Raton (2010)
Bansaye, V., Méléard, S.: Birth and death processes. In: Stochastic Models for Structured Populations, Mathematical Biosciences Institute Lecture Series, pp. 7–17. Springer, Cham (2015)
Baxter, G.J., Blythe, R.A., McKane, A.J.: Exact solution of the multi-allelic diffusion model. Math. Biosci. 209, 124–170 (2007)
Bell, G.: Neutral macroecology. Science 293(5539), 2413–2418 (2001)
Billingsley, P.: Probability and Measure, 4th edn. Wiley, Hoboken (2012)
Bulmer, M.G.: On fitting the Poisson lognormal distribution to species-abundance data. Biometrics 30(1), 101–110 (1974)
Chiu, C.-H., Wang, Y.-T., Walther, B.A., Chao, A.: An improved nonparametric lower bound of species richness via a modified Good–Turing frequency formula. Biometrics 70, 671–682 (2014)
Chou, T., D’Orsogna, M.R.: Coarsening and accelerated equilibration in mass-conserving heterogeneous nucleation. Phys. Rev. E 84, 011608 (2011)
Colwell, R.K., Coddington, J.A.: Estimating terrestrial biodiversity through extrapolation. Philos. Trans. R. Soc. B 345, 101–118 (1994)
Desponds, J., Mora, T., Walczak, A.M.: Fluctuating fitness shapes the clone-size distribution of immune repertoires. Proc. Natl. Acad. Sci. USA 113, 274–279 (2016)
Dessalles, R., Fromion, V., Robert, P.: A stochastic analysis of autoregulation of gene expression. J. Math. Biol. 75, 1–31 (2017)
D’Orsogna, M.R., Lakatos, G., Chou, T.: Stochastic self-assembly of incommensurate clusters. J. Chem. Phys. 136, 084110 (2012)
D’Orsogna, M.R., Zhao, B., Berenji, B., Chou, T.: Combinatoric analysis of heterogeneous stochastic self-assembly. J. Chem. Phys. 137, 121918 (2013)
Fisher, R.A., Corbet, A.S., Williams, C.B.: The relation between the number of species and the number of individuals in a random sample of an animal population. J. Anim. Ecol. 12, 42–58 (1943)
Gibbs, J.P., Martin, W.T.: Urbanization, technology, and the division of labor: international patterns. Am. Sociol. Rev. 27, 667–677 (1962)
Goyal, S., Kim, S., Chen, I.S.Y., Chou, T.: Mechanisms of blood homeostasis: lineage tracking and a neutral model of cell populations in rhesus macaques. BMC Biol. 13(1), 85 (2015)
Grimmett, G., Stirzaker, D.: Probability and Random Processes. Oxford University Press, Oxford (2001)
Hubbell, S.: The Unified Neutral Theory of Biodiversity and Biogeography (MPB-32) (Monographs in Population Biology). Princeton University Press, Princeton (2001)
Hurlbert, S.H.: The nonconcept of species diversity: A critique and alternative parameters. Ecology 52, 577–586 (1971)
Jost, L.: Entropy and diversity. Oikos 113, 363–375 (2006)
Karev, G.P., Wolf, Y.I., Rzhetsky, A.Y., Berezovskaya, F.S., Koonin, Eugene V.: Birth and death of protein domains: a simple model of evolution explains power law behavior. BMC Evolut. Biol. 2, 18 (2002)
Karlin, S., McGregor, J.: The number of mutant forms maintained in a population. In: Proceedings of the Fifth Berkeley Symposium on Mathematics, Statistics and Probability, vol. 4, pp. 415–438 (1967)
Lambert, A.: Species abundance distributions in neutral models with immigration or mutation and general lifetimes. J. Math. Biol. 63, 57–72 (2011)
Laydon, D.J., Bangham, C.R.M., Asquith, B.: Estimating T-cell repertoire diversity: limitations of classical estimators and a new approach. Philos. Trans. R. Soc. B 370, 20140291 (2015)
Lythe, G., Callard, R.E., Hoare, R.L., Molina-París, C.: How many TCR clonotypes does a body maintain? J. Theor. Biol. 389, 214–224 (2016)
MacArthur, R.H., Wilson, E.O.: The Theory of Island Biogeography. Princeton University Press, Princeton (2016)
Miles, J.J., Douek, D.C., Price, D.A.: Bias in the \(\alpha \beta \) T-cell repertoire: implications for disease pathogenesis and vaccination. Immunol. Cell Biol. 89, 375–387 (2011)
Morris, E.K., Caruso, T., Buscot, F., Fischer, M., Hancock, Christine, Maier, Tanja S, Meiners, Torsten, Müller, Caroline, Obermaier, Elisabeth, Prati, Daniel, Socher, Stephanie A, Sonnemann, Ilja, Wäschke, Nicole, Wubet, Tesfaye, Wurst, Susanne, Rillig, Matthias C: Choosing and using diversity indices: insights for ecological applications from the German Biodiversity Exploratories. Ecol. Evol. 4(18), 3514–3524 (2014)
Palmer, M.W.: The estimation of species richness by extrapolation. Ecology 71, 1195–1198 (2003)
Preston, F.W.: The commonness, and rarity, of species. Ecology 29(3), 254–283 (1948)
Sala, C., Vitali, S., Giampieri, E., do Valle, I.F., Remondini, D., Garagnani, P., Bersanelli, M., Mosca, E., Milanesi, L., Castellani, G.: Stochastic neutral modelling of the Gut Microbiota’s relative species abundance from next generation sequencing data. BMC Bioinform. 17, S16 (2016)
Tan, J.T., Dudl, E., LeRoy, E., Murray, R., Sprent, Jonathan, Weinberg, Kenneth I., Surh, Charles D.: IL-7 is critical for homeostatic proliferation and survival of naïve T cells. Proc. Natl. Acad. Sci. USA 98(15), 8732–8737 (2001)
Travaré, S.: The genealogy of the birth, death, and immigration process. In: Feldman, M.W. (ed.) Mathematical Evolution Theory, pp. 41–56. Princeton University Press, Princeton (1989). ISBN 0-691-08502-1
Volkov, I., Banavar, J.R., Hubbell, S.P., Maritan, A.: Neutral theory and relative species abundance in ecology. Nature 424(6952), 1035–1037 (2003)
Acknowledgements
This work was supported in part by an INRA Contrat Jeune Scientifique Award (RD) and by the National Science Foundation through grants DMS-1814364 (TC) and DMS-1814090 (MD). The authors also thank Song Xu for clarifying discussions.
Author information
Authors and Affiliations
Corresponding author
Appendices
Mathematical Appendices
A: Simple Birth–Death–Immigration Models (sBDI)
1.1 A.1: Finite Number of Species
So far, we have assumed immigration events introduce completely new species to the system, regardless of the existing population structure. Within the context of island biodiversity, this assumption corresponds to the mainland hosting an unlimited number of species, so that individuals who emigrate to the island are always part of a new species. Mathematically, we are assuming that each species immigrates only once.
In this Appendix, we consider an alternative model where the number of mainland species Q is finite. In this case, the probability that a newly immigrated individual belongs to species i (with \(1\le i\le Q\)) is 1 / Q and the number of species in the island cannot exceed Q. As a consequence, the total number of species \(C \le Q\), and the number of species with k individuals \(c_{k}\le Q\) for all k.
The dynamics of the total number of individuals N remains unchanged with respect to the sBDI model, as the type of species immigrating from the mainland does not affect overall birth or death rates. Therefore, the distribution for P(N) remains identical to the one derived in Eq. (5) for the simple BDI model. We can now determine the distribution of \(\vec {c}\) in the alternative model using the same approach taken for the sBDI model. Transitions are given by

Note that the birth process rate is effectively augmented by \(\alpha /Q\), due to the possibility of a new individual immigrating into an existing species. Conversely, the corresponding immigration rate for new species is decreased by \(\alpha C/Q\). Also note that the limit \(Q \rightarrow \infty \) reduces the current model to the original sBDI. Using detailed balanced equations, similarly as in the sBDI model, we can write \(P(\vec {c})\) as follows
One can verify that this distribution satisfies all the required transition equations. Yet, contrary to the sBDI model, it is more difficult to determine the distributions of C, \(c_{k}\) and \(n_{i}\) based on this formulation; in particular the factor \(Q!/\left( Q-C\right) !\) prevents us from applying the same mathematical procedure used in the sBDI case.
We can however take a different route, namely invoking neutrality and the independence of the system, to deduce the distributions of C and \(c_{k}\). Since each species behaves independently from all others, we can consider the number \(m_{i}\) of individuals in the \(i^\mathrm{th}\) species (with \(1\le i\le Q\)) independently from the rest. Note that \(m_{i}\) is a random variable that can be zero when there are no individuals of species i present in the system. The quantity \(m_i\) is the counterpart to \(n_{i}\) introduced for the sBDI model with the caveat that \(n_i\) represents the number of individuals of a species actually present on the island (i.e. \(P\left( n_{i}=0\right) =0\)). In the current model \(n_{i}\) can be expressed as a function of \(m_i\) via
describing the distribution of the \(i^\mathrm{th}\) species provided that at least one of its individuals is on the island. The random variable \(m_{i}\) follows a birth and death process: its birth rate is \(\alpha /Q+rm_{i}\) and its death rate is \(\mu m_{i}\). The \(\alpha /Q\) rate corresponds to immigration, the rate \(r m_{i}\) corresponds to actual reproduction. We already determined the steady state distribution of this process in Eq. (5), yielding a negative binomial distribution with parameters \(\alpha /(rQ)\) and \(r/\mu \) as follows
The \(P(n_i)\) distribution can be determined from \(P(m_i)\) expressed above, using Eq. (40)
Finally, the number of species \(c_{k}\) with k individuals and the total number of species C can be expressed as a function of \(m_{i}\) as follows
Since all \(m_{i}\) are i.i.d., the probability distributions of \(c_{k}\) and C are given by
which are binomial distributions of respective parameters Q and \(P\left( m_{i}=k\right) \) for \(c_{k}\), and Q and \(1-P\left( m_{i}=0\right) \) for C. Note that this approach does not allow us to determine the diversity indices H and S.
1.2 A.2: Convergences in the Large Immigration Regime
In this section, we will prove the convergence of
in the large immigration regime defined by \(\alpha = \widetilde{\alpha }\Omega \), \(\Omega \rightarrow \infty \).
Proposition 1
The scaled total number of individuals \(N/\Omega \) converges in distribution to the constant \(\widetilde{\alpha }/(\mu -r)\).
Proof
The definition of the convergence in distribution described in Eq. (3) is equivalent to the convergence of its moment generating function. One is left with showing that
(see for instance [5, Chapter 5]). Since \(N\sim \text {NegBinom}\left( \widetilde{\alpha }\Omega /r,r/\mu \right) \) for which the moment generating function is known, we have for any \(\xi <0\):
Upon taking the logarithm of the previous expression, we find
so

thus proving the proposition. \(\square \)
Proposition 2
The scaled total number of species \(C/\Omega \) converges in distribution to

Proof
The proof is similar to Proposition 1. \(\square \)
Proposition 3
For each \(k>0\), \(c_{k}/\Omega \) converges in distribution to

Proof
For any vector \(\vec {c}\) and \(k\ge 1\), we have that
Consider the moment generating function of the random variable \(c_{k}\). For any \(\xi <0\), we have
Since \(n_{i}\) are identical and independently distributed and independent of C, and since their distributions do not depend on the parameter \(\Omega \), it follows that
Since the probability distribution of \(n_{1}\) is known, we have
Note that for any real A,
Considering the exponential of this expression, we have
Finally, since we have already shown that \(C/\Omega \) converges in distribution (Proposition 2 above), we find
\(\square \)
Proposition 4
The Shannon’s Entropy H converges in distribution as

Proof
Using the definition of H,
where \(c_{k}/\Omega \) and \(N/\Omega \) converge in distribution to known constants, we find

\(\square \)
Proposition 5
The Simpson’s diversity index S converges in distribution as

Proof
By the definition of S (Eq. (2))
and since \(c_{k}/\Omega \) and \(N/\Omega \) converge in distribution to known constants, we find

One can then recognize the power series identity
and hence show that the second term vanishes as \(\Omega \rightarrow \infty \) and deduce the result  . \(\square \)
. \(\square \)
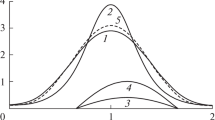
Distribution of the number of individuals in one species \(n_{i}\) under different parameter choices. Dots represent simulations for various values of \(u=\alpha /r\), \(v=r/\mu \), (\(r=1\)) and \(\epsilon \); solid lines depict logarithmic distributions with parameter \(r(1-\epsilon )/\mu \). As expected, the logarithmic distributions match the simulations \(n_{i}\), and the distributions of \(n_{i}\) do not depend on u
Accuracy of Shannon’s entropy and Simpson’s index (as defined in Eq. (25)). We plot the ratio of the estimates of Shannon’s entropy and Simpson’s index and their respective values measured via simulation for different \(u=\alpha /r\), \(v=r/\mu \) (by taking \(r=1\)), and different \(\epsilon \). The estimates become more accurate as \(\mathbb {E}\left[ N\right] \) increases: the error is below \(10\%\) for any parameters u, v, \(\epsilon \) such that \(\mathbb {E}\left[ N\right] \) is larger than 5
B: BDI Model with Mutation (BDIM)
1.1 B.1: Distribution of the Number of Individual in One Species
We propose an argument for a Log-series distribution of any species
when all species are independent of each other. There are several ways to interpret \(\pi _{k}\). First consider the explicit dynamics of each species. Denote by \(m_{q}(t)\) the number of individuals of species q at time t and define \(a_{q}\) as the time of arrival (by convention, we order the species such as \(a_{0}=0<a_{1}<a_{2}<\ldots \)) and \(d_{q}\) its “lifespan”, i.e. the species will be extinct at time \(a_{q}+d_{q}\) (see the example in Fig. 8a). Note that the index q indicates the order of arrival (and not the species identity index i used in the main article), and that the distribution of the times \(a_{q}\) is not specified and can be adapted to any rate of species creation (either by immigration or by mutation). The evolution of each species is independent of each other, and each of them defines an identically distributed birth–death process characterized by the following transitions
Due to the \(r<\mu \) assumption, this process will become extinct almost surely [2, Chapter 2] and the lifespan \(d_{q}\) of each species is finite (Figs. 6 and 7).
a A representative trajectory of three immigrated species. The q-th species is introduced at time \(a_{q}\) and extinguishes at time \(a_{q}+d_{q}\). b Construction of the process \(\overline{m}\) (defined in Eq. (43)) by stacking and concatenating the trajectories of each species \(\left( m_{q}\right) _{q\in \mathbb {N}}\)
In the main article, we interpreted \(\pi _{k}\) as the number of individuals in a given species at steady state, that is to say, we considered the \(T\rightarrow \infty \) limit
where \(J_{T}\) is the index of a randomly sampled species among those that exist at time T; i.e., \(J_{T}\) is uniformly chosen among all the species q such that \(a_{q}<T<a_{q}+d_{q}\).
However, there is another way to interpret \(\pi _{k}\). Consider all species that exist or have existed up to time T and then randomly select one of them, species \(I_{T}\). The number of individuals in species \(I_{T}\) at a randomly chosen time \(\tau _{I_{T}}\) between the introduction of the species (at time \(a_{I_{T}}\)) and the extinction (at time \(a_{I_{T}}+d_{I_{T}}\)) is denoted \(m_{I_{T}}\). In this picture, we can characterize \(\pi _{k}\) according to
The main difference between the two approaches is that, in the first case, we sample among the species that exist at a precise time T before taking \(T\rightarrow \infty \), while in the second case, we sample among all the species that existed before time T (before taking \(T\rightarrow \infty \)).
For a fixed time T, the last species introduced in the system is given by
All species that exist or have existed before time T are in the set \(\left\{ 0,\ldots ,Q_{T}\right\} \). Note that since \(a_{q}\) are increasing in q, \(\lim _{T\rightarrow \infty } Q_{T}=\infty \). As per Eq. (42), we have to sample one species among the set \(\left\{ 0,\ldots ,Q_{T}\right\} \). One key point is that the random selection is not uniform: there is a higher chance of selecting species with longer lifespans. If \(I_{T}\) is the index of the randomly chosen species, we can write
The first term \(\varvec{I}\left( q\le Q_{T}\right) \) ensures that the species q exists before time T while the second term proportionally weights the probability of sampling according to their lifespans. Conditioned on species \(I_{T}\) having been sampled, we then randomly chose a time \(\tau _{I_{T}}\) uniformly distributed between \(a_{I_{T}}\) and \(a_{I_{T}}+d_{I_{T}}\).
Proposition 6
The limiting distribution becomes
Proof
By summing over all possible species q, we can write
Next, consider the process \(\overline{m}(s)\) defined as
with
The process \(\overline{m}\) is simply the stacking of all the processes \(m_{q}\) in the sense that the process \(\overline{m}(t)\) for t between \(\overline{d}_{q}\) and \(\overline{d}_{q+1}\) will be equal to the process \(m_{q}(s)\) for \(s=t-\overline{d}_{q}+a_{q}\) between \(a_{q}\) and its extinction time \(a_{q}+d_{q}\) (see the example on Fig. 8b). With this stacked process,
By ergodicity of the process \(\overline{m}\), we have
Finally, we have to determine the steady state of the process \(\overline{m}\). Since the transitions of the process \(\overline{m}\) are a simple birth–death process
we have that its equilibrium distribution is a logarithmic series distribution with parameter \(p\equiv r(1-\epsilon )/\mu \) (by imposing equations of detailed balance). \(\square \)
1.2 B.2: Moments of C
The third relation of Eq. (1) yields the following expression for the moment generating function of N:
for any \(\xi <0\). Since all the \(\left( n_{i}\right) _{i\le C}\) are identical and independently distributed and independent of C, we have
Equation (20) shows that the distribution over \(n_{1}\) is a log-series distribution with parameter \(p=r\left( 1-\epsilon \right) \). By redefining the variable \(\xi '\) such that \(e^{\xi '}:=\log \left( 1-pe^{\xi }\right) /\log \left( 1-p\right) \) and eliminating \(\xi \) for \(\xi '\), Eq. (45) becomes an expression for the moment generating function of C,
By differentiating this expression, we can determine the second moment of C:
which yields the expression for \(\text {var}\left[ C\right] \) in Eq. (22).
C: BDI Model with Carrying Capacity (BDICC)
1.1 C.1: Steady State Distribution of \(\vec {c}\)
To determine \(P(\vec {c})\), the probability of occurrence of the species-count state \(\vec {c}\), first consider a finite \(K=\mathop {\text {argmax}}\limits _{i}(c_{i}>0)\). As explained in the main text, if the system is reversible, one instance of Eq. (27) is
Recursively unwinding this relationship, we find
After applying Eq. (28), we have by recursion
and
Since the state \(\vec {c}=\vec {0}\) uniquely corresponds to the state \(N=0\) and the above expression holds for K arbitrarily large, it follows that
One can verify that this steady-state distribution satisfies the detailed balanced conditions connecting all pairs of states:
1.2 C.2: Convergence of \(N/\Omega \)
Theorem 7
The random variable \(N/\Omega \) converges in probability to the real \(n^{*}\) which is the only solution of the fixed point Eq. (36).
To prove this Theorem, first define
The function f defines the steady-state constraint on \(n=N/\Omega \) given by Eq. (36) where \(x=n^{*}\) is the only real solution to \(f(x)=1\). With these definitions, the probability distribution over N can be expressed as
Now, consider the following lemma:
Lemma 8
The function f is strictly decreasing and there exists a \(\Omega ^{*}\) for which \(\,\forall \Omega \ge \Omega ^{*}\), \(\left( f_{k}\right) _{k\ge 1}\) is a decreasing sequence.
Proof
The decrease of the function f is a direct implication of the increase of \(\widetilde{\mu }\). For, \(\left( f_{k}\right) _{k\ge 1}\) we have
which is positive for large enough \(\Omega \). Since \(\widetilde{\mu }\) is increasing,
\(\square \)
To prove Theorem 7, we have to show that \(\forall \delta >0\),

that is to say, we have to show that


The proofs of convergence for both limits above are very similar so we will focus on the proof of Eq. (48). To simplify notation, we define \(a_{\Omega ,\delta } \equiv \left\lceil \Omega \left( n^{*}+\delta \right) \right\rceil \), (where \(\left\lceil \cdot \right\rceil \) is the ceiling function). Since the distribution of N is known, we have
Thus, it is enough to show

in order to prove the convergence of Eq. (48).
Proposition 9
In the \(\Omega \rightarrow \infty \) limit, the following equivalence holds
Proof
We first decompose the sum according to
The second term of the decomposition can be rewritten as
Since  , it follows that
, it follows that
As f is a strictly decreasing function (cf. Lemma 8), and since \(n^{*}\) is the only point where \(f(n^{*})=1\), it follows that \(f\left( n^{*}+\delta \right) <1\). Therefore, the sum over n converges, and we have
\(\square \)
With the previous Proposition, it is enough to prove that the ratio
diverges to infinity in order to prove the convergence of Eq. (48).
Proposition 10
The sum

diverges.
Proof
Since \(\left( f_{k}\right) _{k\ge 1}\) is decreasing for large \(\Omega \) (cf. Lemma 8), we have
for sufficiently large \(\Omega \). Therefore,
Since

for large enough \(\Omega \) and since f is decreasing, we have that \(f_{a_{\Omega ,\delta }-1}<1-\eta \) for \(\eta \) small enough. Therefore, we conclude the divergence

and proof of the proposition. \(\square \)
With this Proposition, we have proven the convergence of Eq. (48). The convergence of Eq. (49) can be proved using exactly the same methods by considering \(b_{\Omega ,\delta }= \left\lfloor \Omega \left( \delta +n^{*}\right) \right\rfloor \) instead of \(a_{\Omega ,\delta }\).
1.3 C.3: Convergence of \(C/\Omega \)
Theorem 11
The scaled total number of species \(C/\Omega \) converges in distribution to

in which \(n^{*}\) is the only real solution of the fixed point Eq. (36).
Proof
One has to prove that

with
First note that
Since \(N/\Omega \) converges in probability to \(n^{*}\),
Since the function \(\log \left( \frac{\widetilde{\alpha }e^{\xi /\Omega }+r \left( x-1\right) /\Omega }{\widetilde{\alpha }+r \left( x-1\right) /\Omega }\right) \) is decreasing in x, we can bound the sum with its lower and upper integral bounds
After rescaling \(y = (x-1)/\Omega \), the bounds can be expressed as
Upon taking \(\Omega \rightarrow \infty \) and expanding the above expression, we find that both bounds converge to
Thus, we find
\(\square \)
1.4 C.4: Convergence of \(n_{i}\)
Proposition 12
The marginal probability over each particle count \(n_{i}\) converges according to

Proof
The \(n_{i}\) values are identically distributed, so that for any \(i,j\le C\),
We can then compute the expectation
This expectation is over a product of two converging quantities:
where \(c_{k}/\Omega \) and \(C/\Omega \) converge in distribution to constants

We now apply the mapping theorem (see [5, Chapter 5]) to \(\mathbb {E}\left[ g\left( \frac{c_{k}}{\Omega },\frac{C}{\Omega }\right) \right] \) for any continuous function g to obtain

\(\square \)
1.5 C.5: Explicit Breakdown of Detailed Balance in the BDICC-bis Model with Birth-Mediated Carrying Capacity
Here, we consider a birth–death–immigration model with carrying capacity but contrary to the BDICC model presented in Fig. 1c, the carrying capacity is on the birth rate r(N), and the death rate \(\mu \) is a constant. By analogy with the BDICC analysis, we find a sufficient condition for a steady state to exist
The distribution P(N) of the total number of individuals is given by
where
All possible transitions of the BDICC-bis model are given by

If we assume detailed balance between pairs of states with maximum clone size K, we can recurse the relations
for \(2\le k\le K\) down to the states
to give
Using these chosen pairs of states to impose detailed balance, we find a unique distribution \(P(\vec {c})\). However, this form of \(P(\vec {c})\) will not obey detailed balance between all pairs of states. For example, balancing the transitions

would also require
However, using the \(P(\vec {c})\) from Eq. (50), we find
because generally, \(N\ne C\). Remarkably, the analogous exercise for the BDICC model where \(\mu = \mu (N)\) does satisfy detailed balance between all pairs of states and the \(P(\vec {c})\) we derived for the BDICC model, Eq. (29), is exact.
Rights and permissions
About this article
Cite this article
Dessalles, R., D’Orsogna, M. & Chou, T. Exact Steady-State Distributions of Multispecies Birth–Death–Immigration Processes: Effects of Mutations and Carrying Capacity on Diversity. J Stat Phys 173, 182–221 (2018). https://doi.org/10.1007/s10955-018-2128-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10955-018-2128-4