Abstract
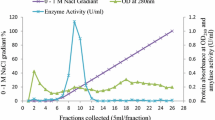
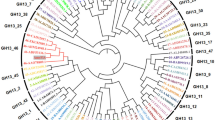
A fragment coding for a putative extracellular α-amylase, from the genomic library of the yeast Saccharomycopsis fibuligera KZ, has been subcloned into yeast expression vector pVT100L and sequenced. The nucleotide sequence revealed an ORF of 1,485 bp coding for a 494 amino acid residues long protein with 99% identity to the α-amylase Sfamy from S. fibuligera HUT 7212. The S. fibuligera KZ α-amylase (Sfamy KZ) belongs to typical extracellular fungal α-amylases classified in the glycoside hydrolase family 13, subfamily 1, as supported also by clustering observed in the evolutionary tree. Sfamy KZ, in addition to the essential GH13 α-amylase three-domain arrangement (catalytic TIM barrel plus domains B and C), does not contain any distinct starch-binding domain. Sfamy KZ was expressed as a recombinant protein in Saccharomyces cerevisiae and purified to electrophoretic homogeneity. The enzyme had a molecular mass 53 kDa and contained about 2.5% of carbohydrate. The enzyme exhibited pH and temperature optima in the range of 5–6 and 40–50 °C, respectively. Stable adsorption of the enzyme to starch granules was not detected but a low degradation of raw starch in a concentration-dependent manner was observed.






Similar content being viewed by others
Abbreviations
- Sfamy:
-
α-Amylase from Saccharomycopsis fibuligera HUT 7212
- Sfamy KZ:
-
α-amylase from S. fibuligera KZ
- ALP1 :
-
Gene coding for α-amylase from S. fibuligera HUT 7212
- LKA1:
-
α-Amylase 1 from Lipomyces kononenkoae
- LKA2:
-
α-Amylase 2 from Lipomyces kononenkoae
- CBM:
-
Carbohydrate-binding module
- SBD:
-
Starch-binding domain
- GH:
-
Glycoside hydrolase
- AMY1:
-
Barley α-amylase 1 (low pI isozyme)
- AMY2:
-
Barley α-amylase 2 (high pI isozyme)
- CAZy:
-
Carbohydrate-active enzymes
References
Apweiler R, Martin MJ, O’Donovan C, Magrane M, …, Zhang J (2010) Nucleic Acids Res 38 (Database issue): D142–D148
Baeva LF, Kozlov DG, Brevnova EE, Benevolenskij SV (1996) Prikl Biochim Mikrobiol 32:311–314
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) Nucleic Acids Res 28:235–242
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) Nucleic Acids Res 37 (Database issue): D233–D238
Chi Z, Chi Z, Liu G, Wang F, Ju L, Zhang T (2009) Biotechnol Advan 27:423–431
Christiansen C, Abou Hachem M, Janecek S, Viksø-Nielsen A, Blennow A, Svensson B (2009) FEBS J 276:5006–5029
Da Lage JL, Feller G, Janecek S (2004) Cell Mol Life Sci 61:97–109
Dubois M, Gilles GA, Hamilton JK, Rebers PA, Smith F (1956) Anal Chem 28:350–356
Eksteen JM, Steyn AJ, van Rensburg P, Cordero Otero RR, Pretorius IS (2003) Yeast 20:69–78
Eksteen JM, Van Rensburg P, Cordero Otero RR, Pretorius IS (2003) Biotechnol Bioeng 84:639–646
Felsenstein J (1985) Evolution 39:783–791
Galdino AS, Ulhoa CJ, Moraes L, Prates MV, Bloch C, Torres F (2008) FEMS Microbiol Lett 280:189–194
Gašperík J, Hostinová E (1990) Biologia 45:1013–1019
Gašperík J, Hostinová E (1993) Curr Microbiol 27:11–14
Gašperík J, Kováč Ľ, Mináriková O (1991) Int J Biochem 23:21–25
Gietz RD, Woods RA (2002) Methods Enzymol 350:87–96
Gupta R, Gigras P, Mohapatra H, Goswam VK, Chauhan B (2003) Process Biochem 38:1599–1616
Hasan K, Ismaya WT, Kardi I, Andiyana Y, Kusumawidjaya S, Ishmayana S, Subroto T, Soemitro S (2008) Biologia 63:1044–1050
Hostinová E, Balanová J, Zelinka J (1990) Biologia 45:301–306
Hostinová E, Solovicová A, Dvorský R, Gašperík J (2003) Arch Biochem Biophys 411:189–195
Iefuji H, Chino M, Kato M, Iimura Y (1996) Biochem J 318:989–996
Itoh T, Ohtsuki I, Yamashita I, Fukui S (1987) J Bacteriol 169:4171–4176
Itoh T, Yamashita I, Fukui S (1987) FEBS Lett 219:339–342
Janecek S (1994) Eur J Biochem 224:519–524
Janecek S (1997) Progr Biophys Mol Biol 67:67–97
Janecek S (2002) Biologia 57(Suppl. 11):29–41
Janecek S, Leveque E, Belarbi A, Haye B (1999) J Mol Evol 48:421–426
Janecek S, Sevcik J (1999) FEBS Lett 456:119–125
Kato S, Shimizu-Ibuka A, Mura K, Takeuchi A, Tokue C, Arai S (2007) Biosci Biotechnol Biochem 71:3007–3013
Kelley LA, Sternberg MJE (2009) Nat Protoc 4:363–371
Kelly RM, Dijkhuizen L, Leemhuis H (2009) J Biotechnol 140:184–193
MacGregor EA, Janecek S, Svensson B (2001) Biochim Biophys Acta 1546:1–20
Machovic M, Janecek S (2006) Cell Mol Life Sci 63:2710–2724
Matsubara T, Ammar YB, Anindyawati T, Yamamoto S, Ito K, Iizuka M, Minamiura N (2004) J Biochem Mol Biol 37:422–428
Matsui I, Matsui E, Ishikawa K, Miyairi S, Honda K (1990) Agric Biol Chem 54:2009–2015
Matsui I, Yoneda S, Ishikawa K, Miyairi S, Fukui S, Umeyama H, Honda K (1994) Biochemistry 18:451–458
Matsuura Y, Kusunoki M, Harada W, Kakudo M (1984) J Biochem 95:697–770
McCann AK, Barnett JA (1986) Yeast 2:109–115
Nielsen MM, Bozonet S, Seo ES, Motyan JA, Andersen JM, Dilokpimol A, Abou Hachem M, Gyemant G, Naested H, Kandra L, Sigurskjold BW, Svensson B (2009) Biochemistry 18:7686–7697
Page RD (1996) Comput Appl Biosci 12:357–358
Pandey A, Nigam P, Soccol CR, Soccol VT, Singh D, Mohan R (2000) Biotechnol Appl Biochem 31:135–152
Ragunath C, Manuel SGA, Kasinathan C, Ramasubbu N (2008) Biologia 63:1028–1034
Ragunath C, Manuel SG, Venkataraman V, Sait HB, Kasinathan C, Ramasubbu N (2008) J Mol Biol 384:1232–1248
Ramachandran N, Pretorius IS, Cordero Otero RR (2005) Biologia 60(Suppl. 16):103–110
Robert X, Haser R, Gottschalk TE, Ratajczak F, Driguez H, Svensson B, Aghajari N (2003) Structure 11:973–984
Rodriguez-Sanoja R, Oviedo N, Sanches S (2005) Curr Opin Microbiol 8:260–267
Saitou N, Nei M (1987) Mol Biol Evol 4:406–425
Sambrook J, Russel DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Shatsky M, Nussinov R, Wolfson HJ (2004) Proteins 56:143–156
Somogyi M (1952) J Biol Chem 195:19–23
Southall SM, Simpson PJ, Gilbert HJ, Williamson G, Williamson MP (1999) FEBS Lett 447:58–60
Stam MR, Danchin EG, Rancurel C, Coutinho PM, Henrissat B (2006) Protein Eng Des Sel 19:555–562
Steyn AJC, Marmur J, Pretorius IS (1995) Gene 166:65–71
Strikovsky A, Hradil J, Wulff G (2003) React Funct Polymers 54:49–61
Sun H, Zhao P, Ge X, Xia Y, Hao Z, Liu J, Peng M (2010) Appl Biochem Biotechnol 160:988–1003
Ševčík J, Hostinová E, Solovicová A, Gašperík J, Dauter Z, Wilson KS (2006) FEBS J 273:2161–2172
Thompson JD, Higgins D, Gibson TJ (1994) Nucleic Acids Res 22:4673–4680
Tibbot BK, Wong DWS, Robertson GH (2002) Biologia 57(Suppl. 11):229–238
Tranier S, Deville K, Robert X, Bozonnet S, Haser R, Svensson B, Aghajari N (2005) Biologia (Suppl. 16):37–46
van der Kaaij RM, Janecek S, van der Maarel MJ, Dijkhuizen L (2007) Microbiology 153:4003–4015
van der Maarel M, van der Veen B, Uitdehaag J, Leemhuis H, Dijkhuizen L (2002) J Biotechnol 94:137–155
Vernet T, Dignard D, Thomas DY (1987) Gene 52:225–233
Vujicic-Zagar A, Dijkstra BW (2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62:716–721
Acknowledgments
This work was supported in part by the grant No. 2/0114/08 from the Slovak grant agency VEGA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hostinová, E., Janeček, Š. & Gašperík, J. Gene Sequence, Bioinformatics and Enzymatic Characterization of α-Amylase from Saccharomycopsis fibuligera KZ. Protein J 29, 355–364 (2010). https://doi.org/10.1007/s10930-010-9260-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-010-9260-6