Abstract
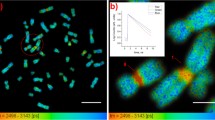
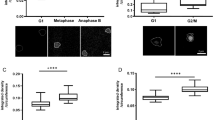
Studies on the physical nature of the structural heterogeneity of chromatin in their native states are few. The eukaryotic chromatin as observed by dye staining studies is of heterogeneous intensity when observed by fluorescent stains, where less and more bright regions apparently correspond to euchromatin and heterochromatin respectively. These are also associated with differential gene expression where it is believed that euchromatin is transcriptionally more active due to increased flexibility. Unfixed squashed preparations of polytene chromosomes of Drosophila were stained with a dsDNA specific dye PicoGreen and fluorescence lifetimes as well as fluorescence anisotropy decay kinetics were measured. Here we report a positive correlation between fluorescence lifetimes and fluorescence intensities, and show that less bright regions corresponding to euchromatin have shorter lifetimes, whereas more bright regions corresponding to heterochromatin have longer lifetimes. We interpret this as less bright regions being more dynamic, a conclusion also supported by fluorescence anisotropy decay kinetics. We infer that the comparatively higher flexibility associated with euchromatin can be directly measured by fluorescence lifetimes and fluorescence anisotropy decay kinetics.



Similar content being viewed by others
References
Heitz E (1928) Das heterochromatin der moose. I Jahrb Wiss Botanik 69:762–818
Dillon N, Festenstein R (2002) Unravelling heterochromatin:competition between positive and negative factors regulates accessibility. Trends Genet 18:252–258
Smith CD, Shu S, Mungall CJ, Karpen GH (2007) The Release 5.1 annotation of Drosophila melanogaster heterochromatin. Science 316:1586–1591
Biosystems Arabidopsis Sequencing Consortium (2000) Cell 100:377–386
Henikoff S (1998) Conspiracy of silence among repeated transgenes. Bioessays 20:532–535
Sun FL, Cuaycong MH, Elgin SC (2001) Long-range nucleosome ordering is associated with gene silencing in Drosophila melanogaster pericentric heterochromatin. Mol Cell Biol 21:2867–2879
Schotta G, Ebert A, Dorn R, Reuter G (2003) Position-effect variegation and the genetic dissection of chromatin regulation in Drosophila. Semin Cell Dev Biol 14:67–75
Banerjee B, Bhattacharya D, Shivashankar GV (2006) Chromatin structure exhibits spatio-temporal heterogeneity within the cell nucleus. Biophys J 91:2297–2303
Roopa T, Shivashankar GV (2006) Direct measurement of local chromatin fluidity using optical trap modulation force spectroscopy. Biophys J 91:4632–4637
Murata S, Herman P, Lin HJ, Lakowicz JR (2000) Fluorescence lifetime imaging of nuclear DNA: effect of fluorescence resonance energy transfer. Cytometry 41:178–185
Sailer BL, Valdez JG, Steinkamp JA, Crissman HA (1998) Apoptosis induced with different cycle-perturbing agents produces differential changes in the fluorescence lifetime of DNA bound ethidium bromide. Cytometry 31:208–216
Zipper H, Brunner H, Bernhagen J, Vitzthum F (2004) Investigations on DNA intercalation and surface binding by SYBR Green I, its structure determination and methodological implications. Nucleic Acids Res 32:e103
Singer VL, Jones LJ, Yue ST, Haugland RP (1997) Characterization of PicoGreen reagent and development of a fluorescence-based solution assay for double-stranded DNA quantitation. Anal Biochem 249:228–238
Cosa G, Vinette AL, McLean JR, Scaiano JC (2002) DNA damage detection technique applying time-resolved fluorescence measurements. Anal Chem 74:6163–6169
Nag N, Ramreddy T, Kombrabail M, Krishna Mohan PM, D’souza J, Rao BJ, Duportail G, Mely Y, Krishnamoorthy G (2006) Dynamics of DNA and Protein-DNA complexes viewed through time-domain fluorescence. In: Geddes CD, Lakowicz JR (eds) Reviews in Fluorescence, vol 3. Springer, US, pp 311–340
Cosa G, Focsaneanu K-S, Scaiano JC, McLean JRN (2000) Direct determination of single-to-double stranded DNA ratio in solution applying time-resolved fluorescence measurements of dye-DNA complexes. Chem Commun 689–690
Pardue ML (1994) Looking at polytene chromosomes. Methods Cell Biol 44:333–351
Mazumder A, Shivashankar GV (2007) Gold-nanoparticleassisted laser perturbation of chromatin assembly reveals unusual aspects of nuclear architecture within living cells. Biophys J 93:2209–2216
Nag N, Rao BJ, Krishnamoorthy G (2007) Altered dynamics of DNA bases adjacent to a mismatch: a cue for mismatch recognition by MutS. J Mol Biol 374:39–53
Acknowledgements
We thank Dr. Anoop Saxena and Swagata Chakraborthy for their help in experiments with TCSPC-microscope and Prof. N. Periasamy for software used in the analysis of fluorescence decay kinetics.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Noothi, S.K., Kombrabail, M., Rao, B.J. et al. Fluorescence Characterization of the Structural Heterogeneity of Polytene Chromosomes. J Fluoresc 20, 37–41 (2010). https://doi.org/10.1007/s10895-009-0519-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-009-0519-2