Abstract
Monogenic autoinflammatory diseases (mAIDs) are a heterogeneous group of diseases affecting primarily innate immunity, with various genetic causes. Genetic diagnosis of mAIDs can assist in the patient’s management and therapy. However, a large number of sporadic and familial cases remain genetically uncharacterized. Deficiency in ELF4, X-linked (DEX) is recently identified as a novel mAID. Here, we described a pediatric patient suffering from recurrent viral and bacterial respiratory infection, refractory oral ulcer, constipation, and arthritis. Whole-exome sequencing found a hemizygous variant in ELF4 (chrX:129205133 A > G, c.691 T > C, p.W231R). Using cells from patient and point mutation mice, we showed mutant cells failed to restrict viral replication effectively and produced more pro-inflammatory cytokines. RNA-seq identified several potential critical antiviral and anti-inflammation genes with decreased expression, and ChIP-qPCR assay suggested mutant ELF4 failed to bind to the promoters of these genes. Thus, we presented the second report of DEX.






Similar content being viewed by others
Data Availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
Code Availability
Not applicable.
Abbreviations
- mAIDs:
-
Monogenic autoinflammatory diseases
- ChIP-qPCR:
-
Chromatin immunoprecipitation-quantitative real-time PCR IEI
- IEI:
-
Inborn errors of immunity
- PIDs:
-
Primary immunodeficiency diseases
- WES:
-
Whole-exome sequencing
- NF-kB:
-
Nuclear factor-kB
- PBMC:
-
Peripheral blood mononuclear cells
- BMDM:
-
Bone marrow-derived macrophage
- VSV:
-
Vesicular stomatitis virus
- HSV-1:
-
Herpes simplex virus 1
- URT:
-
Upper respiratory tract
- WBC:
-
White blood cell
- IVIG:
-
Intravenous immune globulin
- NSAIDs:
-
Non-steroidal anti-inflammatory drugs
- ANCA:
-
Anti-neutrophil cytoplasmic antibody
- BCG:
-
Bacille Calmette-Guérin
- DPT:
-
Diphtheria-pertussis-tetanus
- WT:
-
Wild type
- MUT:
-
Mutant
- DEGs:
-
Differentially expressed genes
- DEX:
-
Deficiency in ELF4, X-linked
References
Martinon F, Aksentijevich I. New players driving inflammation in monogenic autoinflammatory diseases. Nat Rev Rheumatol. 2015;11(1):11–20. https://doi.org/10.1038/nrrheum.2014.158.
Park H, Bourla AB, Kastner DL, Colbert RA, Siegel RM. Lighting the fires within: the cell biology of autoinflammatory diseases. Nat Rev Immunol. 2012;12(8):570–80. https://doi.org/10.1038/nri3261.
Tyler PM, Bucklin ML, Zhao M, Maher TJ, Rice AJ, Ji W, et al. Human autoinflammatory disease reveals ELF4 as a transcriptional regulator of inflammation. Nat Immunol. 2021. https://doi.org/10.1038/s41590-021-00984-4.
Suico MA, Shuto T, Kai H. Roles and regulations of the ETS transcription factor ELF4/MEF. J Mol Cell Biol. 2017;9(3):168–77. https://doi.org/10.1093/jmcb/mjw051.
Lacorazza HD, Miyazaki Y, Di Cristofano A, Deblasio A, Hedvat C, Zhang J, et al. The ETS protein MEF plays a critical role in perforin gene expression and the development of natural killer and NK-T cells. Immunity. 2002;17(4):437–49. https://doi.org/10.1016/s1074-7613(02)00422-3.
Yamada T, Park CS, Mamonkin M, Lacorazza HD. Transcription factor ELF4 controls the proliferation and homing of CD8+ T cells via the Kruppel-like factors KLF4 and KLF2. Nat Immunol. 2009;10(6):618–26. https://doi.org/10.1038/ni.1730.
Mamonkin M, Puppi M, Lacorazza HD. Transcription factor ELF4 promotes development and function of memory CD8(+) T cells in Listeria monocytogenes infection. Eur J Immunol. 2014;44(3):715–27. https://doi.org/10.1002/eji.201343775.
Lee PH, Puppi M, Schluns KS, Yu-Lee LY, Dong C, Lacorazza HD. The transcription factor E74-like factor 4 suppresses differentiation of proliferating CD4+ T cells to the Th17 lineage. J Immunol. 2014;192(1):178–88. https://doi.org/10.4049/jimmunol.1301372.
You F, Wang P, Yang L, Yang G, Zhao YO, Qian F, et al. ELF4 is critical for induction of type I interferon and the host antiviral response. Nat Immunol. 2013;14(12):1237–46. https://doi.org/10.1038/ni.2756.
Du H, Xia H, Liu T, Li Y, Liu J, Xie B, et al. Suppression of ELF4 in ulcerative colitis predisposes host to colorectal cancer. 2021;102169.
Chometon TQ, Siqueira MDS, Sant Anna JC, Almeida MR, Gandini M, Martins de Almeida Nogueira AC, et al. A protocol for rapid monocyte isolation and generation of singular human monocyte-derived dendritic cells. PLoS One. 2020;15(4):e0231132. https://doi.org/10.1371/journal.pone.0231132.
Kao SH, Wang WL, Chen CY, Chang YL, Wu YY, Wang YT, et al. Analysis of protein stability by the cycloheximide chase assay. Bio Protoc. 2015;5(1):e1374. https://doi.org/10.21769/BioProtoc.1374.
Ding Y, Zhou L, Xia Y, Wang W, Wang Y, Li L, et al. Reference values for peripheral blood lymphocyte subsets of healthy children in China. J Allergy Clin Immunol. 2018;142(3):970-3 e8. https://doi.org/10.1016/j.jaci.2018.04.022.
Takeda JI, Nanatsue K, Yamagishi R, Ito M, Haga N, Hirata H, et al. InMeRF: prediction of pathogenicity of missense variants by individual modeling for each amino acid substitution. NAR Genom Bioinform. 2020;2(2):lqaa038. https://doi.org/10.1093/nargab/lqaa038.
Sharrocks AD. The ETS-domain transcription factor family. Nat Rev Mol Cell Biol. 2001;2(11):827–37. https://doi.org/10.1038/35099076.
Rodrigues CH, Pires DE, Ascher DB. DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res. 2018;46(W1):W350–5. https://doi.org/10.1093/nar/gky300.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021. https://doi.org/10.1038/s41586-021-03819-2.
Reva B, Antipin Y, Sander C. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res. 2011;39(17):e118. https://doi.org/10.1093/nar/gkr407.
Wang L, Zhou Y, Chen Z, Sun L, Wu J, Li H, et al. PLCbeta2 negatively regulates the inflammatory response to virus infection by inhibiting phosphoinositide-mediated activation of TAK1. Nat Commun. 2019;10(1):746. https://doi.org/10.1038/s41467-019-08524-3.
Coers J, Brown HM, Hwang S, Taylor GA. Partners in anti-crime: how interferon-inducible GTPases and autophagy proteins team up in cell-intrinsic host defense. Curr Opin Immunol. 2018;54:93–101. https://doi.org/10.1016/j.coi.2018.06.008.
Tretina K, Park ES, Maminska A, MacMicking JD. Interferon-induced guanylate-binding proteins: guardians of host defense in health and disease. J Exp Med. 2019;216(3):482–500. https://doi.org/10.1084/jem.20182031.
Xiong Q, Huang H, Wang N, Chen R, Chen N, Han H, et al. Metabolite-sensing G protein coupled receptor TGR5 protects host from viral infection through amplifying type I interferon responses. Front Immunol. 2018;9:2289. https://doi.org/10.3389/fimmu.2018.02289.
Huang H, Zhang N, Xiong Q, Chen R, Zhang C, Wang N, et al. Elimination of GPR146-mediated antiviral function through IRF3/HES1-signalling pathway. Immunology. 2017;152(1):102–14. https://doi.org/10.1111/imm.12752.
Zheng R, Wan C, Mei S, Qin Q, Wu Q, Sun H, et al. Cistrome data browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res. 2019;47(D1):D729–35. https://doi.org/10.1093/nar/gky1094.
McDermott MF, Aksentijevich I, Galon J, McDermott EM, Ogunkolade BW, Centola M, et al. Germline mutations in the extracellular domains of the 55 kDa TNF receptor, TNFR1, define a family of dominantly inherited autoinflammatory syndromes. Cell. 1999;97(1):133–44. https://doi.org/10.1016/s0092-8674(00)80721-7.
Hollenhorst PC, McIntosh LP, Graves BJ. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu Rev Biochem. 2011;80:437–71. https://doi.org/10.1146/annurev.biochem.79.081507.103945.
Acknowledgements
We thank the patient and his family for their cooperation, especially for the patient’s mother whose perseverance in caring for her child gives us strength to pursue a definitive genetic diagnosis and next a more effective personalized treatment. We also thank all other members of You lab (Peking University Health Science Center) for insightful discussion of the results and practical technical tips.
Funding
This study is funded by Sanming Project of Medicine in Shenzhen (SZSM201812002), Livelihood Project of Chongqing, China (cstc2018jscxmsybX0005), National Natural Science Foundation of China (81974255), National Natural Science Foundation of China (81900136), Postdoc Fund at Chongqing Medical University, and Ministry of Education Key Laboratory Foundation at Children’s Hospital of Chongqing Medical University.
Author information
Authors and Affiliations
Contributions
HQ. D and XD. Z conceived and supervised the project, reviewed and revised the manuscript; HQ. D designed the experiments, analyzed data, and wrote the manuscript. G. S performed the experiments, analyzed data, and followed the patient. FP. Y provided plasmids, viruses, experimental expertise, and scientific sights in designing the project and revising the manuscript. HW. X constructed ELF4 mutant plasmids. LL. C performed ChIP-qPCR experiment. Others provided essential help for clinical management and follow-up of the patient and valuable clinical expertise in reviewing the manuscript.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
The ethics committee of Children’s Hospital of Chongqing Medical University approved the study. Blood samples collection were performed in accordance with the Declaration of Helsinki. Written informed consent for participation in the study were obtained from patients’ parents.
Consent for Publication
Written informed consent for publication of the study was obtained from patients’ parents.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10875_2022_1243_MOESM1_ESM.jpg
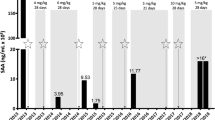
Supplementary Fig.1 Mutant cells failed to restrain viral replication effectively. A Plaque-formation assays of culturesupernatant of peripheral monocytes from healthy control (HC) and patient (P)after 12 h infection with VSV or HSV-1-GFP. B mRNA quantification of VSV in peripheral monocytes from healthycontrol (HC) and patient (P) after 12 h infection with VSV. C Microscopy analysis (left images,magnification, 5×) of HeLa cells transfected with human WT ELF4 (hWT ELF4),human W231R mutant ELF4 (hMut ELF4), mouse WT ELF4 (mWT ELF4) or mouse W231Rmutant ELF4 (mMut ELF4) after 12 h infection with HSV-1-GFP. Right graph,quantification of HSV-1-GFP infected cells indicated by GFP expression, 2-3random fields were counted. DMicroscopy analysis (left images, magnification, 5×) of BMDM from WT mice (WT)and point mutation mice (Mut) after 6h and 12 h infection with HSV-1-GFP. Rightgraph, quantification of HSV-1-GFP infected cells indicated by GFP expression,2-3 random fields were counted. Data were pooled from at least 3 independent experimentsand represented as mean±SD. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.Unpaired two-sided Student’s t test (JPG 854 KB)
10875_2022_1243_MOESM3_ESM.jpg
Supplementary Fig. 3Loss of function of ELF4 leads to increased virus-induced inflammation. A mRNA quantification of indicatedinterferons of peripheral monocytes from healthy control (HC) and patient (P)after 12 h infection with VSV or HSV-1-GFP. B IFN-b mRNA quantification of HeLa cells transfected with human WTELF4 (hWT ELF4), human W231R mutant ELF4 (hMut ELF4), mouse WT ELF4 (mWT ELF4)or mouse W231R mutant ELF4 (mMut ELF4) after 12 h infection with HSV-1-GFP. C mRNA quantification of indicatedinterferons of BMDM from WT mice (WT) and point mutation mice (Mut) after 6 hinfection with VSV or HSV-1-GFP. DmRNA quantification of indicated cytokines of peripheral monocytes from healthycontrol (HC) and patient (P) after 12 h infection with VSV or HSV-1-GFP. E ELISA quantification of indicatedcytokines in peripheral monocytes isolated from healthy control (HC) andpatient (P) after 12 h infection with VSV or HSV-1-GFP. F mRNA quantification of indicated cytokines and ELISAquantification of IL-6 in peritoneal macrophage from WT mice (WT) and pointmutation mice (Mut) after 6 h infection with VSV or HSV-1-GFP. Data were pooledfrom at least 3 independent experiments and represented as mean±SD. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.Unpaired two-sided Student’s t test (JPG 1540 KB)
10875_2022_1243_MOESM4_ESM.jpg
Supplementary Fig. 4 A Representative image of flow cytometryanalysis of IL-17A producing CD4+T cells, gated on CD3+CD4+T cells. B Flow cytometry analysis showing percentageof IL-17A producing CD4+T cells differentiated from naïve CD4+ Th17 in vitrounder non-pathogenic conditions (TGFβ + IL-6), cells were restimulated with PMAand Ionomycin. WT n = 3, Mut n = 3. CMouse IL-17A ELISA analysis of Th17 culture supernatant after restimulation. WTn = 3, Mut n = 3. Data were represented as mean±SD.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Unpairedtwo-sided Student’s t test (JPG 449 KB)
10875_2022_1243_MOESM5_ESM.jpg
Supplementary Fig. 5 MutantELF4 fails to activate potential antiviral or anti-inflammatory genes. A mRNA quantification of indicatedgenes in BMDM from WT mice (WT) and point mutation mice (Mut) after 6 hinfection with VSV or HSV-1-GFP. B mRNAquantification of indicated genes in peripheral monocytes from healthy control (HC)and patient (P) after 6 h infection with VSV or HSV-1-GFP. C ChIP-qPCR experiments showing Flag-ELF4 binding to promoters ofindicated genes in HEK293T cells transfected with empty vector, WT ELF4, W231Rmutant ELF4 or W251A mutant ELF4. Data were pooled from at least 3 independent experimentsand represented as mean±SD. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.Unpaired two-sided Student’s t test (JPG 515 KB)
10875_2022_1243_MOESM6_ESM.jpg
Supplementary Fig. 6Cistrome database mining. Elf4 antibody binding peaks in the promoters ofindicated genes. Raw data were derived from Cistrome database (JPG 1821 KB)
Rights and permissions
About this article
Cite this article
Sun, G., Qiu, L., Yu, L. et al. Loss of Function Mutation in ELF4 Causes Autoinflammatory and Immunodeficiency Disease in Human. J Clin Immunol 42, 798–810 (2022). https://doi.org/10.1007/s10875-022-01243-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-022-01243-3