Abstract
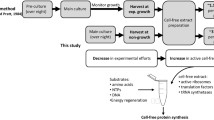
Cell-free protein synthesis is suitable for stable-isotope labeling of proteins for NMR analysis. The Escherichia coli cell-free system containing potassium acetate for efficient translation (KOAc system) is usually used for stable-isotope labeling, although it is less productive than other systems. A system containing a high concentration of potassium l-glutamate (l-Glu system), instead of potassium acetate, is highly productive, but cannot be used for stable-isotope labeling of Glu residues. In this study, we have developed a new cell-free system that uses potassium d-glutamate (d-Glu system). The productivity of the d-Glu system is approximately twice that of the KOAc system. The cross peak intensities in the 1H–15N HSQC spectrum of the uniformly stable-isotope labeled Ras protein, prepared with the d-Glu system, were similar to those obtained with the KOAc system, except that the Asp intensities were much higher for the protein produced with the d-Glu system. These results indicate that the d-Glu system is a highly productive cell-free system that is especially useful for stable-isotope labeling of proteins.



Similar content being viewed by others
Abbreviations
- CAT:
-
Chloramphenicol acetyltransferase
- HSQC:
-
Heteronuclear single quantum coherence
- Ras(Y32W)/d-Glu:
-
Ras(Y32W) protein produced by the d-Glu system
- Ras(Y32W)/KOAc:
-
Ras(Y32W) protein produced by the KOAc system
References
Bax A (1994) Multidimensional nuclear magnetic resonance methods for protein studies. Curr Opin Struct Biol 4:738–744
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Jewett MC, Swartz JR (2004) Substrate replenishment extends protein synthesis with an in vitro translation system designed to mimic the cytoplasm. Biotechnol Bioeng 87:465–472
Johnson BA (2004) Using NMR View to visualize and analyze the NMR spectra of macromolecules. Methods Mol Biol 278:313–352
Kigawa T, Muto Y, Yokoyama S (1995) Cell-free synthesis and amino acid-selective stable isotope labeling of proteins for NMR analysis. J Biomol NMR 6:129–134
Kigawa T, Yabuki T, Yoshida Y, Tsutsui M, Ito Y, Shibata T, Yokoyama S (1999) Cell-free production and stable-isotope labeling of milligram quantities of proteins. FEBS Letters 442:15–19
Kigawa T, Yabuki T, Matsuda N, Matsuda T, Nakajima R, Tanaka A, Yokoyama S (2004) Preparation of Escherichia coli cell extract for highly productive cell-free protein expression. J Struct Funct Genomics 5:63–68
Kim DM, Kigawa T, Choi CY, Yokoyama S (1996) A highly efficient cell-free protein synthesis system from Escherichia coli. Eur J Biochem 239:881–886
Li H, Inoue M, Yabuki T, Aoki M, Seki E, Matsuda T, Nunokawa E, Motoda Y, Kobayashi A, Terada T, Shirouzu M, Koshiba S, Lin YJ, Guntert P, Suzuki H, Hayashizaki Y, Kigawa T, Yokoyama S (2005) Solution structure of the mouse enhancer of rudimentary protein reveals a novel fold. J Biomol NMR 32:329–334
Maeda T, Inoue M, Koshiba S, Yabuki T, Aoki M, Nunokawa E, Seki E, Matsuda T, Motoda Y, Kobayashi A, Hiroyasu F, Shirouzu M, Terada T, Hayami N, Ishizuka Y, Shinya N, Tatsuguchi A, Yoshida M, Hirota H, Matsuo Y, Tani K, Arakawa T, Carninci P, Kawai J, Hayashizaki Y, Kigawa T, Yokoyama S (2004) Solution structure of the SEA domain from the murine homologue of ovarian cancer antigen CA125 (MUC16). J Biol Chem 279:13174–13182
Matsuda T, Kigawa T, Koshiba S, Inoue M, Aoki M, Yamasaki K, Seki M, Shinozaki K, Yokoyama S (2006) Cell-free synthesis of zinc-binding proteins. J Struct Funct Genomics, (in press). DOI 10.1007/s10969-006-9012-1
Ozawa K, Headlam MJ, Schaeffer PM, Henderson BR, Dixon NE, Otting G (2004) Optimization of an Escherichia coli system for cell-free synthesis of selectively N-labelled proteins for rapid analysis by NMR spectroscopy. Eur J Biochem 271:4084–4093
Spirin AS, Baranov VI, Ryabova LA, Ovodov SY, Alakhov YB (1988) A continuous cell-free translation system capable of producing polypeptides in high yield. Science 242:1162–1164
Vinarov DA, Lytle BL, Peterson FC, Tyler EM, Volkman BF, Markley JL (2004) Cell-free protein production and labeling protocol for NMR-based structural proteomics. Nature Methods 1:149–153
Wu PS, Ozawa K, Jergic S, Su XC, Dixon NE, Otting G (2006) Amino-acid type identification in 15N-HSQC spectra by combinatorial selective 15N-labelling. J. Biomol. NMR 34:13–21
Yamasaki K, Shirouzu M, Muto Y, Fujita-Yoshigaki J, Koide H, Ito Y, Kawai G, Hattori S, Yokoyama S, Nishimura S, Miyazawa T (1994) Site-directed mutagenesis, fluorescence, and two-dimensional NMR studies on microenvironments of effector region aromatic residues of human c-Ha-Ras protein. Biochemistry 33:65–73
Yamasaki K, Kigawa T, Inoue M, Tateno M, Yamasaki T, Yabuki T, Aoki M, Seki E, Matsuda T, Tomo Y, Hayami N, Terada T, Shirouzu M, Tanaka A, Seki M, Shinozaki K, Yokoyama S (2005) Solution structure of an Arabidopsis WRKY DNA binding domain. Plant Cell 17:944–956
Yokoyama S (2003) Protein expression systems for structural genomics and proteomics. Curr Opin Chem Biol 7:39–43
Acknowledgements
We are grateful to Aya Kurosawa, Natsumi Suzuki, Yasuko Tomo, Yukiko Fujikura, Satoko Yasuda, Fumiko Hiroyasu, and Masaomi Ikari for their technical assistance. We thank Naohiro Kobayashi and Tadashi Tomizawa for help with the NMR analysis, and Jun Yokoyama for valuable suggestions and comments. We also thank Azusa Ishii, Kiyomi Yajima, and Tomoko Nakayama for expert secretarial assistance. This work was supported by the RIKEN Structural Genomics/Proteomics Initiative (RSGI) of the National Project on Protein Structural and Functional Analyses, Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Matsuda, T., Koshiba, S., Tochio, N. et al. Improving cell-free protein synthesis for stable-isotope labeling. J Biomol NMR 37, 225–229 (2007). https://doi.org/10.1007/s10858-006-9127-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-006-9127-5