Abstract
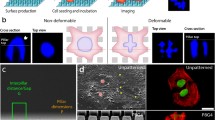
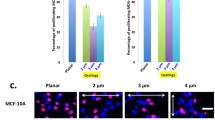
Osteosarcoma-derived cell lines (SaOs-2, MG63) have recently been shown to deform their nucleus considerably in response to surface topography. Such a deformation had not been described previously. Here we present results on additional cell lines, including cancerous (OHS4, U2OS), immortalized (F/STRO-1+A and FHSO6) and healthy cells (HOP). The cancerous cells were found to deform extensively, the immortalized cells showed small deformations, whereas the healthy cells showed deformation only at short incubation times. These results suggest a strong link between the malignant transformation of cells and the state of the cytoskeletal network. We propose mechanisms to explain the deformation in which the cytoskeleton either pushes down on the nucleus during spreading or pulls it down upon adhesion to the pillars.




Similar content being viewed by others
References
Su WT, Chu IM, Yang JY, Lin CD. The geometric pattern of a pillared substrate influences the cell-process distribution and shapes of fibroblasts. Micron. 2006;37:699–706.
Tan JL, Tien J, Pirone DM, Gray DS, Bhadriraju K, Chen CS. Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc Natl Acad Sci. 2003;100:1484–9.
Steinberg T, Schultz S, Spatz JP, Grabe N, Mussig E, Kohl A, et al. Early keratinocyte differentiation on micropillar interfaces. Nano Lett. 2007;7:287–94.
Thery M, Racine V, Piel M, Pépin A, Dimitrov A, Chen Y, et al. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. PNAS. 2006;103:19771–6.
Bischofs IB, Schmidt SS, Schwarz US. Effect of adhesion geometry and rigidity on cellular force distributions. Phys Rev Lett. 2009;103:048101.
Anselme K, Bigerelle M, Noël B, Loison I, Hardouin P. Kinetic study of the expression of β-catenin, actin and vinculin during osteoblastic adhesion on grooved titanium substrates. Biomed Mater Eng. 2004;14:545–56.
Tsai WB, Ting YC, Yang JY, Lai JY, Liu HL. Fibronectin modulates the morphology of osteoblast-like cells (MG-63) on nano-grooved substrates. J Mater Sci Mater Med. 2009;20:1367–78.
Dalby MJ, Riehle MO, Yarwood SJ, Wilkinson CD, Curtis AS. Nucleus alignment and cell signaling in fibroblasts: response to a micro-grooved topography. Exp Cell Res. 2003;284:274–82.
Davidson P, Özçelik H, Hasirci V, Reiter G, Anselme K. Micro-structured surfaces cause severe but non-detrimental deformation of the cell nucleus. Adv Mater. 2009;21:3586–90.
Ingber DE. Tensegrity: the architectural basis of cellular mechanotransduction. Annu Rev Physiol. 1997;59:575–99.
Taddei A, Hediger F, Neumann FR, Gasser SM. The function of nuclear architecture: a genetic approach. Annu Rev Genet. 2004;38:305–45.
Reddy KL, Zullo JM, Bertolino E, Singh H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature. 2008;452:243–7.
Ben-Ze’ev A. The cytoskeleton in cancer cells. Biochim Biophys Acta. 1985;780:197–212.
Ali SH, DeCaprio JA. Cellular transformation by SV40 large T antigen: interaction with host proteins. Semin Cancer Biol. 2001;11:15–23.
Guck J, Schinkinger S, Lincoln B, Wottawah F, Ebert S, Romeyke M, et al. Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence. Biophys J. 2005;88:3689–98.
Anselme K, Broux O, Noël B, Bouxin B, Bascoulergue G, Dudermel A-F, et al. In vitro control of human bone marrow stromal cells for bone tissue engineering. Tissue Eng. 2002;8:941–53.
Oyajobi BO, Lomri A, Hott M, Marie PJ. Isolation and characterization of human clonogenic osteoblast progenitors immunoselected from fetal bone marrow stroma using STRO-1 monoclonal antibody. J Bone Miner Res. 1999;14:351–61.
Fromigue O, Kheddoumi N, Lomri A, Marie PJ, Body JJ. Breast cancer cells release factors that induced apoptosis in human bone marrow stromal cells. J Bone Miner Res. 2001;16:1600–10.
Rodan GA, Majeska RJ. Phenotypic maturation of osteoblastic osteosarcoma cells in culture. Prog Clin Biol Res. 1982;110 Pt B:249–59.
Bidwell JP, Alvarez M, Feister H, Onyia J, Hock J. Nuclear matrix proteins and osteoblast gene expression. J Bone Miner Res. 1998;13:155–67.
Wang N, Tytell JD, Ingber DE. Mechanotransduction at a distance: mechanically coupling the extracellular matrix with the nucleus. Nat Rev Mol Cell Biol. 2009;10:75–82.
Frolov VA, Zimmerberg J. Shaping biological matter. Nat Mater. 2009;8:173–4.
Comer KA, Dennis PA, Armstrong L, Catino JJ, Kastan MB, Kumar CC. Human smooth muscle alpha-actin gene is a transcriptional target of the p53 tumor suppressor protein. Oncogene. 1998;16:1299–308.
Heessen S, Leonchiks A, Issaeva N, Sharipo A, Selivanova G, Masucci MG, et al. Functional p53 chimeras containing the Epstein-Barr virus Gly-Ala repeat are protected from Mdm2- and HPV-E6-induced proteolysis. Proc Natl Acad Sci U S A. 2002;99:1532–7.
Ochalek T, Nordt FJ, Tullberg K, Burger MM. Correlation between cell deformability and metastatic potential in B16–F1 melanoma cell variants. Cancer Res. 1988;48:5124–8.
Docheva D, Padula D, Popov C, Mutschler W, Clausen-Schaumann H, Schieker M. Researching into the cellular shape, volume and elasticity of mesenchymal stem cells, osteoblasts and osteosarcoma cells by atomic force microscopy. J Cell Mol Med. 2008;12:537–52.
Acknowledgments
Financial support provided through the European Community’s ‘‘Marie Curie Actions’’ under contract MRTN-CT-2004-005516 (BioPolySurf). V. H. also acknowledges the support of TUBITAK (the Scientific and Technical Research Council of Turkey) through the project Nanobiomat (TBAG 105T508). We also acknowledge Prof. A. Aydinli of Bilkent University for the patterned templates.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Davidson, P.M., Fromigué, O., Marie, P.J. et al. Topographically induced self-deformation of the nuclei of cells: dependence on cell type and proposed mechanisms. J Mater Sci: Mater Med 21, 939–946 (2010). https://doi.org/10.1007/s10856-009-3950-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10856-009-3950-7