Abstract
The chemical universe database GDB-13, which enumerates 977 million organic molecules up to 13 atoms of C, N, O, S and Cl following simple chemical stability and synthetic feasibility rules, represents a vast reservoir for new fragments. GDB-13 was classified using the MQN-system discussed in the preceding paper for the analysis of PubChem fragments. Two hundred and fifty-five subsets of GDB-13 were generated by the combinatorial use of eight restrictive criteria, including fragment-like (“rule of three”) and scaffold-like (no acyclic carbon atoms) filters. Virtual screening for analogs of 15 commercial drugs of 13 non-hydrogen atoms or less shows that retrieving MQN-neighbors of a query molecule from GDB-13 or its subsets provides on average a 38-fold enrichment in structural analogs (Daylight-type substructure fingerprint Tanimoto T SF > 0.7), and a 75-fold enrichment in shape-similar analogs (ROCS TanimotoCombo score > 1.4). An MQN-searchable version of GDB-13 is provided at www.gdb.unibe.ch.
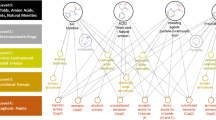
Graphical Abstract









Similar content being viewed by others
References
Coyne AG, Scott DE, Abell C (2010) Drugging challenging targets using fragment-based approaches. Curr Opin Chem Biol 14:299–307
Schulz MN, Hubbard RE (2009) Recent progress in fragment-based lead discovery. Curr Opin Pharmacol 9:615–621
Hartenfeller M, Schneider G (2011) De novo drug design. Methods Mol Biol 672:299–323
Venhorst J, Nunez S, Kruse CG (2010) Design of a high fragment efficiency library by molecular graph theory. ACS Med Chem Lett 1:499–503
Carr RA, Congreve M, Murray CW, Rees DC (2005) Fragment-based lead discovery: leads by design. Drug Discov Today 10:987–992
Rees DC, Congreve M, Murray CW, Carr R (2004) Fragment-based lead discovery. Nat Rev Drug Discov 3:660–672
Hajduk PJ, Greer J (2007) A decade of fragment-based drug design: strategic advances and lessons learned. Nat Rev Drug Discov 6:211–219
Boyd SM, de Kloe GE (2010) Fragment library design: efficiently hunting drugs in chemical space. Drug Discov Today 7:e173–e180
Wang Y, Xiao J, Suzek TO, Zhang J, Wang J, Bryant SH (2009) PubChem: a public information system for analyzing bioactivities of small molecules. Nucleic Acids Res 37:W623–W633
van Deursen R, Blum LC, Reymond JL (2011) Visualisation of the chemical space of fragments, lead-like and drug-like molecules in PubChem. J Comput Aided Mol Des. doi:10.1007/s10822-011-9437-x
Blum LC, Reymond JL (2009) 970 million druglike small molecules for virtual screening in the chemical universe database GDB-13. J Am Chem Soc 131:8732–8733
Fink T, Bruggesser H, Reymond JL (2005) Virtual exploration of the small-molecule chemical universe below 160 daltons. Angew Chem Int Ed Engl 44:1504–1508
Fink T, Reymond JL (2007) Virtual exploration of the chemical universe up to 11 atoms of C, N, O, F: assembly of 26.4 million structures (110.9 million stereoisomers) and analysis for new ring systems, stereochemistry, physicochemical properties, compound classes, and drug discovery. J Chem Inf Model 47:342–353
Nguyen KT, Syed S, Urwyler S, Bertrand S, Bertrand D, Reymond JL (2008) Discovery of NMDA glycine site inhibitors from the chemical universe database GDB. ChemMedChem 3:1520–1524
Nguyen KT, Luethi E, Syed S, Urwyler S, Bertrand S, Bertrand D, Reymond JL (2009) 3-(aminomethyl)piperazine-2, 5-dione as a novel NMDA glycine site inhibitor from the chemical universe database GDB. Bioorg Med Chem Lett 19:3832–3835
Luethi E, Nguyen KT, Burzle M, Blum LC, Suzuki Y, Hediger M, Reymond JL (2010) Identification of selective norbornane-type aspartate analogue inhibitors of the glutamate transporter 1 (GLT-1) from the chemical universe generated database (GDB). J Med Chem 53:7236–7250
Garcia-Delgado N, Bertrand S, Nguyen KT, van Deursen R, Bertrand D, Reymond J-L (2010) Exploring a7-nicotinic receptor ligand diversity by scaffold enumeration from the chemical universe database GDB. ACS Med Chem Lett 1:422–426
Reymond JL, Van Deursen R, Blum LC, Ruddigkeit L (2010) Chemical space as a source for new drugs. Med Chem Commun 1:30–38
Nguyen KT, Blum LC, van Deursen R, Reymond JL (2009) Classification of organic molecules by molecular quantum numbers. ChemMedChem 4:1803–1805
Pearlman RS, Smith KM (1998) Novel software tools for chemical diversity. Perspect Drug Discov Des 9–11:339–353
Geppert H, Vogt M, Bajorath J (2010) Current trends in ligand-based virtual screening: molecular representations, data mining methods, new application areas, and performance evaluation. J Chem Inf Model 50:205–216
Akella LB, DeCaprio D (2010) Cheminformatics approaches to analyze diversity in compound screening libraries. Curr Opin Chem Biol 14:325–330
Irwin JJ, Shoichet BK (2005) ZINC—A free database of commercially available compounds for virtual screening. J Chem Inf Model 45:177–182
van Deursen R, Blum LC, Reymond JL (2010) A searchable map of PubChem. J Chem Inf Model 50:1924–1934
Huang N, Shoichet BK, Irwin JJ (2006) Benchmarking sets for molecular docking. J Med Chem 49:6789–6801
Willett P, Barnard JM, Downs GM (1998) Chemical similarity searching. J Chem Inf Comput Sci 38:983–996
Rush TS III, Grant JA, Mosyak L, Nicholls A (2005) A shape-based 3-D scaffold hopping method and its application to a bacterial protein-protein interaction. J Med Chem 48:1489–1495
McKay BD (1981) Practical graph isomorphism. Congr Numerantium 30:45–87
Warr WA (1993) Computer-assisted structure elucidation. Part II: indirect database approaches and established systems. Anal Chem 65:1087A–1095A
Steinbeck C (2004) Recent developments in automated structure elucidation of natural products. Nat Prod Rep 21:512–518
Ertl P, Schuffenhauer A (2009) Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J Cheminform 1:8
Congreve M, Carr R, Murray C, Jhoti H (2003) A rule of three for fragment-based lead discovery? Drug Discov Today 8:876–877
Kolb P, Ferreira RS, Irwin JJ, Shoichet BK (2009) Docking and chemoinformatic screens for new ligands and targets. Curr Opin Biotechnol 20:429–436
Willett P (2006) Similarity-based virtual screening using 2D fingerprints. Drug Discov Today 11:1046–1053
Khalifa AA, Haranczyk M, Holliday J (2009) Comparison of nonbinary similarity coefficients for similarity searching, clustering and compound selection. J Chem Inf Model 49:1193–1201
Hawkins PC, Skillman AG, Nicholls A (2007) Comparison of shape-matching and docking as virtual screening tools. J Med Chem 50:74–82
Nicholls A, McGaughey GB, Sheridan RP, Good AC, Warren G, Mathieu M, Muchmore SW, Brown SP, Grant JA, Haigh JA, Nevins N, Jain AN, Kelley B (2010) Molecular shape and medicinal chemistry: a perspective. J Med Chem 53:3862–3886
Schneider G, Neidhart W, Giller T, Schmid G (1999) “Scaffold-hopping” by topological pharmacophore search: a contribution to virtual screening. Angew Chem Int Ed Engl 38:2894–2896
Jolliffe IT (2002) Principal component analysis, 2nd edn. Springer, New York
Acknowledgments
This work was supported financially by the University of Berne, the Swiss National Science Foundation and the Office Fédéral Suisse de l’Education et de la Science.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Blum, L.C., van Deursen, R. & Reymond, JL. Visualisation and subsets of the chemical universe database GDB-13 for virtual screening. J Comput Aided Mol Des 25, 637–647 (2011). https://doi.org/10.1007/s10822-011-9436-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-011-9436-y