Abstract
Purpose
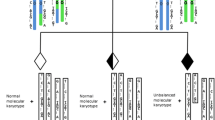
For translocation carriers, preimplantation genetic diagnosis (PGD) provides the opportunity to distinguish between normal/balanced and unbalanced embryos prior to implantation and, as such, increases the likelihood of a successful ongoing pregnancy. The data presented here compares autosomal reciprocal and Robertsonian translocation segregation patterns in day 3 versus day 5/6 IVF-PGD embryos to determine if there is a difference in the chromosome segregation patterns observed at these developmental time points.
Methods
A retrospective analysis on PGD translocation carriers at Monash IVF was performed. Segregation patterns were compared between day 3 and day 5/6 embryos to ascertain whether selection against malsegregants exists.
Results
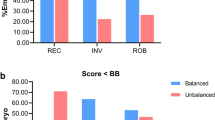
For reciprocal translocations, 1649 day 3 embryos (139 translocations) from 144 couples and 128 day 5/6 embryos (59 translocations) from 60 couples were analysed. Day 3 segregation analysis showed that 22.3% of embryos were normal/balanced (consistent with 2:2 alternate segregation) and 77.7% were unbalanced (malsegregation). Day 5/6 segregation analysis showed that 53.1% of embryos were normal/balanced and 46.9% were unbalanced. For Robertsonian translocations, 847 day 3 embryos (8 translocations) from 54 couples and 193 day 5/6 embryos (6 translocations) from 31 couples were analysed. Day 3 segregation analysis showed that 38.7% of embryos were normal/balanced (consistent with 2:1 alternate segregation) and 61.3% were unbalanced. Day 5/6 segregation analysis showed that 74.1% of embryos were normal/balanced and 25.9% were unbalanced.
Conclusions
This data demonstrates an increase in the proportion of genetically normal/balanced embryos at day 5/6 of development. This suggests a strong natural selection process between day 3 and day 5/6 in favour of normal/balanced embryos. These findings support performing PGD testing on day 5/6 of embryo development.


Similar content being viewed by others
References
Gardner R, Sutherland G, Shaffer L. Chromosome abnormalities and genetic counseling. 4th ed. New York: Oxford University Press; 2012.
Scriven P, Handyside A, Ogilvie C. Chromosome translocations: segregation modes and strategies for preimplantation genetic diagnosis. Prenat Diag. 1998;18:1437–49.
Dahdouh EM, Balayla J, Garcia-Velasco JA. Impact of blastocyst biopsy and comprehensive chromosome screening technology on preimplantation genetic screening: a systematic review of randomized controlled trials. Reprod BioMed Online. 2015;30(3):281–9.
Jones GM, Trounson AO, Gardner DK, Kausche A, Lolatgis N, Wood C. Evolution of a culture protocol for successful blastocyst development and pregnancy. Hum Reprod. 1998;13(1):169–77.
Sandalinas M, Sadowy S, Alikani M, Calderon G, Cohen J, Munné S. Developmental ability of chromosomally abnormal human embryos to develop to the blastocyst stage. Human Repro. 2001;16:1954–8.
Scott R, Upham K, Forman E, Zhao T, Treff N. Cleavage-stage biopsy significantly impairs human embryonic implantation potential while blastocyst biopsy does not: a randomized and paired clinical trial. Fertil Steril. 2013;100(3):624–30.
Grifo JA, Hodes-Wertz B, Lee HL, et al. Single thawed euploid embryo transfer improves IVF pregnancy, miscarriage, and multiple gestation outcomes and has similar implantation rates as egg donation. J Assist Reprod Genet. 2013;30(2):259–64.
Scott RT Jr, Upham KM, Forman EJ, et al. Blastocyst biopsy with comprehensive chromosome screening and fresh embryo transfer significantly increases in vitro fertilization implantation and delivery rates: a randomized controlled trial. Fertil Steril. 2013;100(3):697–703.
Chen M, Wei S, Hu J, Quan S. Can comprehensive chromosome screening technology improve IVF/ICSI outcomes? A meta-analysis. PLOS one. October 15 2015.
Brodie D, Beyer CE, Osborne E, Kralevski V, Rasi S, Osianlis T. Preimplantation genetics diagnosis for chromosome rearrangements—one blastomere biopsy versus two blastomere biopsy. J Assist Reprod Genet. 2012;29:821–7.
Munné S, Márquez C, Magli C, Morton P, Morrison L. Scoring criteria for preimplantation genetic diagnosis of numerical abnormalities for chromosomes X, Y, 13, 16, 18 and 21. Mol Hum Repro. 1998;4(9):863–70.
Rabinowitz M, Ryan A, Gemelos G, Hill M, Baner J, Cinnioglu C, et al. Origins and rates of aneuploidy in human blastomeres. Fertil Steril. 2013;97(2):395–401.
BlueGnome. A technical guide to aneuploidy calling with 24sure V3. Version 1.0, 12th July 2012.
BlueGnome. Technical note: 24sure+ translocation detection. 16th November 2010.
Illumina. VeriSeq PGS Library Preparation Guide. February 2014.
Illumina. A technical guide to aneuploidy calling with VeriSeq PGS. September 2014.
Munne S, Grifo J, Wells D. Mosaicism: “survival of the fittest” versus “no embryo left behind”. Fertil Steril. 2016;105(5):1146–9.
Spinella F, Biricik A, Surdo M, Cotroneo E, Bono S, Cursio E, Minasi MG, Greco E, Fiorentino F. Improved detection of chromosomal mosaicism by next generation sequencing (NGS)-based preimplantation genetic screening. Presented to PGDIS conference, Bologna, Italy. Held 10th May 2016.
Fragouli E, Alfarawati S, Spath K, Tarozzi N, Borini A, Wells D. The developmental potential of mosaic embryos. Fertility and Sterility. 2015;104(3) Supplement.
Horak J, Kubicek D, Hornak M, Pesakova M, Travnik, Vesela K. Impact of mosaicism detection on PGS clinical outcome. Presented to PGDIS conference, Bologna, Italy. Held 10th May 2016.
Tormasi S, Capaldi R, Gouw F, Welch C, Munne S, Coates A. Mosaicism rates in embryos resulting in live birth or miscarriage. Presented to PGDIS conference, Bologna, Italy. Held 10th May 2016.
Mastenbroek S, Twisk M, van Echten-Ardens J, Sikkema-Raddatz B, Korevaar JC, Verhoeve HR, et al. In vitro fertilization with preimplantation genetic screening. N Engl J Med. 2007;357(1):9–17.
Dahdouh EM, Balayla J, Garcia-Velasco JA. Impact of blastocyst biopsy and comprehensive chromosome screening technology on preimplantation genetic screening: a systematic review of randomized controlled trials. Reprod BioMed Online. 2007;30(3):281–9.
Beyer CE, Osianlis T, Boekel K, Osborne E, Rombauts L, Catt J, et al. Preimplantation genetic screening outcomes are associated with culture conditions. Hum Reprod. 2009;24:1212–20.
Tan Y, Tan K, Zhang S, Gong F, Cheng D, Xiong B, et al. Single-nucleotide polymorphism microarray-based preimplantation genetic diagnosis is likely to improve the clinical outcome for translocation carriers. Hum Reprod. 2013;28(9):2581–92.
Fragouli E, Alfarawati S, Spath K, Jaroudi S, Sarasa J, Enciso M, et al. The origin and impact of embryonic aneuploidy. Hum Genet. 2013;132(9):1001–13.
Benkhalifa M, Kasakyan S, Clement P, Baldi M, Tachdjian G, Demirol A, et al. Array comparative genomic hybridisation profiling of first-trimester spontaneous abortions that fail to grow in vitro. Prenat Diagn. 2005;25:894–900.
Rubio C, Rodrio L, Mercader A, Mateu E, Buendia P, Pehlivan T, et al. Pellicer. Impact of chromosomal abnormalities on preimplantation embryo development. Prenatal Diag. 2007;27:748–56.
Fiorentino F, Biricik A, Bono S, Spizzichino L, Cotroneo E, Cottone G, et al. Development and validation of a nextgeneration sequencing-based protocol for 24-chromosome aneuploidy screening of embryos. Fertil Steril. 2014;101:1375–82.
Fiorentino F, Bono S, Biricik A, Nuccitelli A, Cotroneo E, Cottone G, et al. Application of next-generation sequencing technology for comprehensive aneuploidy screening of blastocysts in clinical preimplantation genetic screening cycles. Hum Reprod. 2014;29:2802–13.
Yang Z, Lin J, Zhang J, Fong WI, Li P, Zhao R, et al. Randomized comparison of next-generation sequencing and array comparative genomic hybridization for preimplantation genetic screening: a pilot study. BMC Med Genet. 2015;8:30.
Yin X, Tan K, Vajta G, Jiang H, Tan Y, Zhang C, et al. Massively parallel sequencing for chromosomal abnormality testing in trophectoderm cells of human blastocysts. Biol Reprod. 2013;88:69.
Munné S. Preimplantation genetic diagnosis for aneuploidy and translocations using array comparative genomic hybridization. Curr Genomics. 2012;13:463–70.
Colls P, Escudero T, Fischer J, Cekleniak NA, Ben-Ozer S, Meyer B, et al. Validation of array comparative genome hybridization for diagnosis of translocations in preimplantation human embryos. Reprod BioMed Online. 2012;24:621–9.
Treff NR, Northrop LE, Kasabwala K, Su J, Levy B, Scott RT Jr. Single nucleotide polymorphism microarray-based concurrent screening of 24-chromosome aneuploidy and unbalanced translocations in preimplantation human embryos. Fertil Steril. 2011;95:1606–1612.e1–2. 107.
Johnson DS, Hill M, Abae M, Frederick J, Swanson M, Rabinowitz M. First clinical application of DNA microarrays for translocations and inversions. Fertil Steril. 2010;93:S13–4.
Fragouli E, Alfarawati S, Spath K, Wells D. Morphological and cytogenetic assessment of cleavage and blastocyst stage embryos. Mol Hum Repro. 2014;20(2):117–26.
Burgoyne PS, Holland K, Stephens R. Incidence of numerical chromosome anomalies in human pregnancy estimation from induced and spontaneous abortion data. Hum Reprod. 1991;6:555–65.
Yusuf R, Naeem R. Cytogenetic abnormalities in products of conception: a relationship revisited. Am J Repro Immunology. 2004;52(1):88–96.
Azmanov D, Milachich T, Zaharieve B, Michailova G, Dimitrova V, Karagiozova Z, Maznejkova V, Chernev T, Toncheva D. Profile of chromosomal aberrations in different gestational age spontaneous abortions detected by comparative genomic hybridisation. Eur J Obstet Gynecol Reprod Biol 2007;131:127–31.
McCoy R, Demko Z, Ryan A, Banjevic M, Hill M, Sigurjonsson S, Rabinowitz M, Petrov D. Evidence of selection against complex mitotic-origin aneuploidy during preimplantation development. PLOS Genetics. October 22 2015.
Acknowledgements
The authors would like to thank all the embryology staff at Monash IVF for performing the embryo biopsy procedures and the genetics staff at Monash IVF for their genetic analysis and contributions towards this paper.
Author information
Authors and Affiliations
Contributions
Claire E. Beyer participated in study concept and design, data acquisition, study execution, analysis, manuscript drafting, critical discussion and interpretation of data.
Elissa Willats participated in study design, manuscript drafting, critical discussion and interpretation of data.
Corresponding author
Ethics declarations
This study was approved by the Monash Surgical Private Hospital Human Research Ethics Committee (approval number: 07078).
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Supplementary Table S1
(DOC 159 kb).
Rights and permissions
About this article
Cite this article
Beyer, C.E., Willats, E. Natural selection between day 3 and day 5/6 PGD embryos in couples with reciprocal or Robertsonian translocations. J Assist Reprod Genet 34, 1483–1492 (2017). https://doi.org/10.1007/s10815-017-1009-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-017-1009-0