Abstract
Environmental DNA (eDNA) has seen a massive increase in application in freshwater systems with a concurrent growth in protocol developments and a drive to gain a better understanding of the ‘ecology’ of eDNA. This raises the question of whether we are currently still in an early, developmental phase of eDNA-based assessments or already transitioning into a more applied stage for biomonitoring. I conducted a systematic literature review on 381 eDNA-focused studies in freshwater systems targeting macro-organisms over the last 5 years, assessing study goals, methods, target systems and taxa and study design aspects. The results show an increase of biomonitoring-focused studies throughout the years, while the fraction of studies investigating the ‘ecology’ of eDNA decreased. The application of metabarcoding significantly increased while studies applying qPCRs tentatively declined. A geographic inequality was observed concerning study numbers and study goals biased towards the global North. Descriptive studies increased, but the fraction of in-field studies and studies applying eDNA and conventional methods combined revealed no trend. These results show a shift towards application-focused work for eDNA-based assessments but also reveal this field to still be developing. In this transitional phase, practitioners need to ensure consistency and data comparability for long-term monitoring programmes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the ongoing global biodiversity crisis, freshwater habitats have been classified as one of the most vulnerable ecosystems (Dudgeon et al., 2006; WWF, 2020). In order to halt the decline in biodiversity and ecosystem functioning of these habitats, various legislative frameworks, such as the EU Water Framework Directive (WFD), the Clean Water Act of the USA (CWA) or the Water Act 2007 of Australia have been installed. Hereby, all these legislative bodies require solid biodiversity assessment methods in order to ensure the biological integrity of the protected systems (Birk et al., 2012; Bohan et al., 2017; Leese et al., 2018).
The utilization of environmental DNA (or eDNA) has been announced to revolutionize the way we monitor biodiversity, particularly in aquatic systems (Deiner et al., 2017; Cristescu & Hebert, 2018; Pawlowski et al., 2018; Ruppert et al., 2019; Beng & Corlett, 2020). Environmental DNA is defined as DNA that is captured from an environmental sample without pre-isolating any target organisms (Taberlet et al., 2012). The sample can be anything from water, soil, sediments, air, faeces or gut contents or even ancient sources like cores from ice, sediments or permafrost (Taberlet et al., 2012; Thomsen & Willerslev, 2015). With respect to macrobial organisms, this DNA captured from the environment stems from the organisms interacting with their surroundings and permanently shedding genetic material into the environment through, e.g. sloughing of skin cells, hairs or scales, excretion of body fluids or defecation (eDNA sensu stricto Pawlowski et al., 2020).
For freshwater assessments, the striking advantages of eDNA-based methods are among others (1) non-invasiveness—causing little to no disturbance to the target species and target system (when targeting macro-organisms and when sampling eDNA sensu strictu, see above), (2) time and cost efficiency allowing to monitor biodiversity over large spatial and temporal scales, (3) high detection sensitivity, allowing reliable detection of rare or elusive species or early stages of biological invasions and (4) universality of sampling, which allows to assess biodiversity on a wide taxonomic range using the same eDNA sample (Goldberg et al., 2016; Deiner et al., 2017; Shaw et al., 2017; Pawlowski et al., 2018; Beng & Corlett, 2020).
In the case of water samples, eDNA-based approaches first capture and concentrate this shed DNA from the water via filtration, centrifugation or precipitation and then purify the DNA from this concentrate through DNA extraction (Shaw et al., 2017; Ruppert et al., 2019). The next step in the workflow selectively targets the DNA of the investigator’s taxon (or taxon group) of interest. The taxonomic level can hereby range from individual species to whole domains or even ‘total biodiversity’ (Compson et al., 2020) and also dictates the specific method applied for the detection of the target DNA in the ‘mixture’ of an eDNA sample: The first of the two main avenues in this regard encompasses single-species or ‘targeted’ eDNA approaches. These aim to detect DNA of individual (often threatened or invasive) species. A widely applied method for this targeted approach is the usage of real-time or quantitative polymerase chain reaction (qPCR). Hereby, species-specific primers (and sometimes hydrolysis probes) that can be fluorescently labelled are used during PCR amplification. The presence of the target DNA in the eDNA extract is inferred by an increase of the fluorescence signal during PCR (Logan et al., 2009; Doi et al., 2015). In the last few years, digital droplet PCR (ddPCR) has emerged as an alternative targeted DNA detection method. It generates thousands of nanoliter droplets during PCR, ideally each of them containing one or no copy of the target DNA. PCR is carried out in each droplet in parallel using fluorescently labelled probes and the presence of the target PCR is inferred by an end-point measure of the fluorescence signal in all droplets in parallel (Hindson et al., 2011; Doi et al., 2015). ddPCR seems to surpass qPCR concerning sensitivity, inhibitor resistance and DNA quantification (Doi et al., 2015; Hunter et al., 2017), but currently requires higher acquisition and running costs than qPCR. The second avenue encompasses community approaches. Hereby, metabarcoding is the most commonly applied method if multiple species, whole species groups or even complete biodiversity is targeted in an eDNA survey. Hereby, general or universal primers are used to amplify a target marker of the target taxon (group) from the mix of DNA in the eDNA extract, followed by a high-throughput sequencing (HTS) step. The obtained sequences are then usually compared to reference sequences in order to identify the species composition of the target group in the eDNA extract (Taberlet et al., 2012; Shaw et al., 2017; Ruppert et al., 2019). Other methods to characterize the biodiversity contained in an eDNA sample, though less frequently applied, include conventional PCRs (e.g. Wilson et al. 2018; Xu et al. 2018; Cardas et al. 2020), CRISPR-Cas (Williams et al., 2019a), hybridization capture (e.g. Wilcox et al., 2018) and metagenomics (e.g. Bista et al., 2018; Cowart et al., 2018).
The compelling assets of eDNA-based approaches have led to an intense drive to add eDNA-based methods to the standard toolbox for ecosystem monitoring (particularly for aquatic systems) and to include them in the monitoring schemes of the abovementioned legislative bodies (e.g. Leese et al., 2016; Deiner et al., 2017; Hering et al., 2018) but also to continuous research on how to improve workflows and correctly interpret the data. This resulted in an impressive growth of scientific publications on eDNA-based applications in ecology and conservation throughout the last two decades (Beng & Corlett, 2020; Compson et al., 2020). Efforts to improve the workflow can focus on eDNA sampling and sample preservation (Hunter et al., 2019; Ladell et al., 2019; Kumar et al., 2021), DNA extraction (Wong et al., 2020; Anderson & Thompson, 2021), the development of new primers or optimization of laboratory protocols for species-specific assays (e.g. Elberri et al., 2020; Schenekar et al., 2020; Ito and Shibaike, 2021) or for species community assays (Miya et al., 2015; Elbrecht & Leese, 2017) or the development of new software or pipelines for sequence data processing (Sato et al., 2018; Carøe & Bohmann, 2020; Diana et al., 2021; Macher et al., 2021). Advances that have been made in the last decade in the field of waterborne eDNA-based assessments include among others the comparison of differing capture methods of eDNA, with filtering slowly emerging as the most efficient method for many research questions (Bruce et al., 2021), gaining a better understanding of the correlation between sampling effort and DNA recovered, as well as detection probabilities (e.g. Cantera et al., 2019; Furlan et al., 2019), how to identify and deal with inhibited samples (e.g. Bruce et al., 2021) and the development of suitable metabarcoding primers for important bioindicator groups such as diatoms, macroinvertebrates or fishes (e.g. Miya et al., 2015; Elbrecht & Leese, 2017; Vasselon et al., 2017).
eDNA-based assessments have already proven to be effective to answer various ecological questions in freshwater habitats. Such include assessing the presence of elusive, endangered or rare species, like cryptic macroinvertebrates, high-risk fish species or amphibians like the great crested newt Triturus cristatus (Mächler et al., 2014; Biggs et al., 2015; Doi et al., 2017; Roy et al., 2018; Griffiths et al., 2020), monitoring the spread of invasive species in aquatic systems like Asian carp, Dreissenid mussels or the American bullfrog Lithobates catesbeianus (Dejean et al., 2012; Jerde et al., 2013; Ardura et al., 2017), studying spawning ecology of elusive species (Sigsgaard et al., 2015; Maruyama et al., 2018) and species richness assessments of bioindicator taxa (Olds et al., 2016; Deiner et al., 2017).
While for various taxa and habitats, robust and reliable protocols have been developed and have been proven to be equal or even superior to conventional monitoring (see examples above), there are still aspects in the process of eDNA analyses that are not yet fully understood and for certain target taxa or habitats more research and/or protocol development might be needed before we can include these approaches reliably into monitoring schemes.
Areas requiring further research and development (regarding freshwater eDNA analyses) include standardized sampling protocols for habitats with very differing environmental conditions (such as springs, small creeks, large streams, ponds, lakes, swamps or estuaries) as well as the definition of sampling controls to keep the data among studies consistent and comparable (Bruce et al., 2021). Concerning laboratory analyses, open questions on remain concerning PCR inhibition, PCR stochasticity or primer bias (Bruce et al., 2021). Furthermore, the uneven availability of published protocols and/or primer sets for different taxa might pose a significant challenge for eDNA-based assays to be fully integrated into standard biomonitoring schemes (Belle et al., 2019). Well-designed protocols, reported with defined minimum standards, are needed for every bioindicator group or target taxon of interest for freshwater biomonitoring to enable the full implementation of eDNA-based methods into comprehensive biomonitoring frameworks. Thereby, the standardized minimum standards are required to understand the applicability of the respective assays for certain research questions or biodiversity assessments, as well as the associated uncertainties and limitations (such as detection sensitivities) and comparability with others (Thalinger et al., 2021). Analytical aspects like the incorporation of imperfect detection models (such as occupancy modelling), abundance estimations, taxonomic assignment and the assignment of an ecological status are also still a very active field of research and will require harmonization (Beng & Corlett, 2020) with the aim to improve consistency and comparability among surveys. For metabarcoding, the availability of well-curated reference databases is a fundamental prerequisite. The efforts to achieve this are tightly interwoven with the global barcoding initiative of the international Barcode of Life project (iBOL). Inequalities in barcode availability have been identified and researchers are actively working on closing those gaps (Weigand et al., 2019).
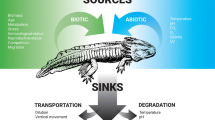
A topic that has become a major focus of investigators in the field of eDNA-based research in the recent years, is the dynamics of eDNA production, distribution, persistence and degradation in the respective study system (Turner et al., 2015; Eichmiller et al., 2016; Tsuji et al., 2017; Allan et al., 2020; Wood et al., 2020). These dynamics, also referred to as the ‘ecology’ of eDNA (Barnes & Turner, 2016) are of decisive importance for estimating power, sensitivity as well as the spatiotemporal scale of the drawn inferences from any eDNA study. For example, depending on various environmental factors, such as temperature, pH, UV radiation or availability of oxygen, eDNA in aquatic systems can degrade within days to the point of non-detectability (Barnes et al., 2014; Barnes & Turner, 2016; Seymour et al., 2018) or persist up to hundreds or even thousands of years in an analyzable state in sediments of freshwater systems (e.g. Giguet-Covex et al., 2019). Likewise, eDNA in lotic systems might be transported downstream only several hundred metres (Jane et al., 2015; Wilcox et al., 2016) or up to dozens of kilometres (Deiner & Altermatt, 2014; Pont et al., 2018). The environmental factors driving these dynamics show great variability among systems, requiring habitat-specific assessments (Harper et al., 2019; Sales et al., 2021), promoting the increase of studies targeting at a better understanding of eDNA ‘ecology’.
Because of the rapid development of eDNA-based assessments in the last two decades, with certain research questions being answered, but new important aspects requiring more research being uncovered, eDNA-based methods are still developing and our understanding of how to optimally conduct and interpret the results of such studies is still progressing. Publications on new or modified protocols in the whole workflow still seem to be published at an equal speed as the application of previously published protocols. However, in order to implement a novel methodology in national or international biomonitoring scheme, the methodology needs to be clearly defined. Furthermore, this workflow has to be proven as effective and robust without the need to adapt it frequently, as protocol modification might result in loss of valuable data comparability among different assessments or in long-term biomonitoring studies.
Therefore, the concurrent and ongoing expansion in eDNA application and advancements in the optimal execution of such studies raises the question of whether we are still at a very early, developmental phase of eDNA-based monitoring with the majority of the research still focusing on methodological advancements and our fundamental understanding of eDNA dynamics, or whether eDNA-based methods are already transforming into a standard tool for freshwater biomonitoring with increased true application of such methods for biodiversity assessments.
The aim of this study was to assess the current state of eDNA-based research on macro-organisms in freshwater ecosystem, with a special emphasis on the main goals of the individual studies reviewed. In particular, I wanted to evaluate whether a relative shift towards true application of eDNA-based methods as a biodiversity monitoring tool can be observed in the most recent years. Furthermore, an overview of the relative frequencies of target taxa and target freshwater systems in such studies is presented and selected aspects of study designs illuminated.
Methods
A literature search was conducted through the Clarivate Web of Science portal on October 30th, 2021. The following search terms and search fields were thereby used: eDNA OR ‘environmental DNA' (All fields) and freshwater OR river$ OR pond$ OR lake$ (All fields) and English (Language) and Article (Document Type). The search was limited to the years 2017 or newer by setting the search field (Publication Date) to Last 5 years (2017–2021). A second search was conducted on November 21st, 2021, using the same search terms but not limiting the search field (Publication Date), allowing to return results from all years of the database (dating back to the year 1900). From this second search, only the number of publications per year was recorded.
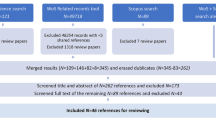
The search results of the first search were exported from Web of Science and imported into Mendeley Reference Manager 1.19.8 (Mendeley Ltd.), whereby the following metadata of the articles were kept: article title, author list, publication year and month, journal, volume, issue, start and end page (or article number), DOI (if available) and abstract. The titles and abstracts were screened to remove out-of-scope documents. Thereby, the following exclusion criteria were used: (1) Articles focusing solely on marine or terrestrial ecosystems and not targeting freshwater habitats. Articles targeting estuarine habitats were kept for analysis, (2) articles solely focusing on microorganisms such as bacteria, viruses, pathogens, protozoans, prokaryotes or fungi, (3) articles focusing solely on ancient DNA in lake sediments, (4) articles utilizing eDNA from freshwater sampling to infer terrestrial biodiversity only, (5) articles focusing solely on diet analysis, (6) articles focusing solely on barcode generation and (7) articles such as reviews, opinion articles or conference papers.
For publications passing this filter, the full articles were downloaded. If the full article could not be accessed, the article was removed from analysis. The remaining articles were subjected to full analysis and the following information was extracted from each publication:
Main goal of the study: The primary purpose for conducting the work, assigned to the following three categories (multiple categories were assigned to a study if it targeted more than one goal with equal emphasis): (1) ‘Biodiversity monitoring’—studies collecting field samples from systems for which the presence of the target taxon was unknown. Studies applying metabarcoding to field samples were always assigned to this category. (2) ‘Development or optimization of protocols’. Studies developing new or optimizing existing protocols in the complete workflow of an eDNA assessment, including sampling, sample preservation, sample processing and data analysis. (3) ‘Understanding the “ecology” of eDNA’—studies aiming to elucidate the dynamics of eDNA production, distribution, transport, fate and/or degradation of eDNA in the target system.
Method applied to detect the target eDNA: Categorized into: (1) ‘qPCR’, (2) ‘ddPCR’, (3) ‘metabarcoding’ and (4) ‘others’ (including conventional PCR, PCR-free approaches such as CRISPR-Cas or metagenomics). Studies were assigned multiple categories if using multiple methods.
Target taxon: The species or species group being the main target of investigation in the study. If the target taxon was not unambiguously defined in the publication, the targeted taxa of the utilized primers were noted. The target taxa were then grouped into the following 13 categories: ‘fishes’ (including lampreys), ‘amphibians’, ‘crustaceans’, ‘reptiles’, ‘mammals’, ‘molluscs’, ‘macroinvertebrates’, ‘macrophytes’, ‘microalgae’, ‘vertebrates’, ‘multiple’ (of the previously listed), ‘metazoa or higher’ and ‘other’. While these groups are at different taxonomic levels, they were chosen to obtain an as homogeneous and consistent classification as possible.
Target system: the freshwater system is targeted by each study. Usually the system from which the field samples originated. For laboratory-based studies, this was the named target system, if explicitly stated. The target systems were then grouped into the following 10 categories: ‘rivers’, ‘lakes/ponds’, ‘lakes/ponds and rivers combined’, ‘artificial channels’, ‘dam reservoirs’, ‘estuaries’, ‘groundwater’, ‘wetlands’, ‘multiple’ (any combination of the previously mentioned categories except lakes/ponds and rivers combined’) and ‘other’.
Country: The country of origin from which field samples were taken. If study was solely laboratory based without utilizing field samples, the country of the first author(s)’ affiliation was listed. Countries were then grouped into the following seven continental assignments: ‘Africa’, ‘Asia’, ‘Australia’, ‘Europe’, ‘North America’, ‘South America’, ‘multiple’ (of the previously listed).
Study design aspects: (a) Study type: Studies were either categorized as ‘manipulative’ (the investigators actively and significantly altered the system of which the eDNA samples originated, with the aim to influence the presence of the target eDNA in this system), or ‘descriptive’ (no significant alterations of the system by the investigators). (b) Study location: Either categorized as ‘in-field’ (eDNA samples were collected from an uncontrolled environment, outside of a laboratory setting) or ‘solely laboratory-based or theoretical’ (eDNA samples were collected from a controlled setting or no eDNA samples were collected at all). If the study was split into two clearly separated sub-studies with different designs, both study types could apply to one study. (c) eDNA versus conventional: For each study targeting biodiversity monitoring, it was listed whether biodiversity monitoring was carried solely using eDNA-based approaches or in combination with alternative or conventional methods.
All data were compiled into a database in Microsoft Excel (Online Resource 1). Statistical analyses and plotting were carried out using R 4.1.2 (R Core Team, 2021) and IBM SPSS v23 (IBM Corp, 2016). Exponential curve fitting of the yearly publication trend was carried out using the nls() function of R and linear modelling to reveal temporal trends of study aspects via the lm() function. Chi-squared tests of homogeneity of methods applied among study goals as well as of study targets and study systems among years were conducted with the chisq.test() function. Figures were finalized using CorelDRAW 2019 (Corel Corporation 2019).
Results
Yearly number of publications
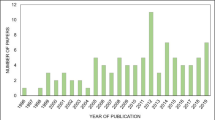
The search encompassing the years 1900–2021 returned 1343 search results. Thereby, a steep increase in publications between 2008 and 2020 can be observed, fitting an exponential curve (Fig. 1). The highest number of studies was published in the year 2020 (221 studies), followed by the year 2021 (196 studies), which was not yet fully completed by the time of the search (November 2021) and 2018 (170 studies).
Yearly distribution of search results of the two literature searches conducted in this review. Red bars indicate publications retrieved via the ‘all-year’ (1900–2021) search. Cyan bars indicate the number of publications of the 2017–2021 search passing the filtering and curation steps and that have been analysed in this study (total: 381 studies). Black line represents fitted curve of the nonlinear regression model of the red bar data between 2008 and 2020. Statistics of the regression are given in the top left corner. Note that the year 2021 was not fully completed at the time point of the searches (October/November 2021), therefore 2021 and 2022 (containing Early View articles) were excluded for curve fitting
The search limited to the last 5 years returned 876 search results. After screening title and abstract of these 876 publications, 431 (49.2%) out-of-scope articles were removed and 445 (50.7%) were retained for which full articles were downloaded. A total of 11 articles were removed from analysis because the full article was not accessible. During data extraction, an additional 64 were excluded from analysis because they did not meet the inclusion criteria, resulting in a final dataset of 381 publications (43.5% of initial search results). Thereby, the yearly number of publications steadily increased starting at 51 (year 2017) and ending at 108 (year 2021) (Fig. 1).
Main goals of studies
Overall, 247 of the 381 studies in this meta-analysis (64.8%) featured ‘biodiversity monitoring’ either as the sole study goal or in combination with other goals. A total of 220 studies (57.7%) included ‘development or optimization of a protocol’ as a study goal and 76 studies (19.9%) aimed to increase the ‘understanding of the “ecology” of eDNA’. Hereby, an increase in percentage of studies targeting ‘biodiversity monitoring’ was observed between 2017 and 2021 (F-statistic = 39.8, P = 0.008), whereas studies targeting a ‘understanding of the “ecology” of eDNA’ decreased in percentage (F-statistic = 10.5, P = 0.047). No clear trend regarding the percentage of studies targeting ‘protocol development or optimization’ could be observed (F-statistic = 0.06, P = 0.81) (Table 1, Fig. 2).
Temporal trends in relative frequencies of studies featuring ‘biodiversity monitoring’, ‘protocol development or optimization’ and ‘understanding the “ecology” of eDNA’ as their study goal. Dots represent percentage of total studies of the respective publication year. Lines represent regression curves from linear modelling and symbols to the right-hand side of the lines indicate direction and significance of respective slope coefficients (+ : positive, − : negative, n.s.: not significant, P: P value). Shadings indicate 95% confidence intervals of linear models
Geographically, a clear separation could be seen between the global North, namely North America, Europe and Asia, and the global South, namely South America, Africa and Australia (Fig. 3). A total of 337 studies (88.5%) of the 381 assayed studies were conducted in the global North (North America: 136 studies—35.7%, Europe: 110 studies—28.9%, Asia: 91 studies—23.9%) with only a small fraction of the studies (41 studies—10.8%)—targeting systems in the global South (South America: 16 studies—4.2%, Africa: 4 studies—1.0%, Australia 21 studies—5.5%). Three studies (0.8%) were conducted on multiple continents. Within Asia, 19 studies were conducted in China and 58 studies were conducted in Japan; therefore, 84.6% of the studies in Asia were conducted in these two countries (Online Resource 2). Regarding study goals, North America, Europe and Asia revealed overall a lower proportion of studies targeting ‘biodiversity monitoring’ compared to the global South, while studies aiming to gain a better understanding of the ‘ecology' of eDNA were almost exclusively conducted in the global North (Table 2). Only two out of 76 such studies were conducted in the global South, both in Australia. The percentages of studies involving protocol development was relatively similar across continents. However, South America and Africa had the lowest proportions of such studies. Overall, the countries from which most studies originated were USA (112 studies—29.4%), Japan (58 studies—15.2%) and United Kingdom (31 studies—8.1%).
Geographic distribution of the 381 studies surveyed, separated by study goal. Scales next to the respective bar plots indicate total number of studies and study goals are indicated by colour codes of the respective bar. Studies could be assigned more than one study goal. Cartographic basis: (Esri, 2021), visualized using QGIS 3.10.0 (QGIS Development Team, 2019)
Methods applied
Across all 381 studies, qPCRs were the most frequently applied method throughout the 381 studies (210 studies—55.1%), followed by metabarcoding (139 studies—36.5%), ddPCR (18 studies—4.7%) and other methods (34 studies—8.9%). However, the proportion of studies applying metabarcoding significantly increased throughout the last 5 years (Fig. 4, F-statistic = 11.3, P = 0.0437), whereas the application of qPCR showed a slight trend of decrease (F-statistic = 9.23, P = 0.056). No trend was observed concerning the proportion of studies applying ddPCR or other methods (F-statistic = 2.1 and 3.7, respectively, both P > 0.1). For studies targeting ‘biodiversity monitoring’ (248 studies), 122 studies (49.2%) applied qPCRs in their workflow, 110 studies (44.4%) used metabarcoding, 10 studies (4.0%) applied ddPCRs and 21 studies (8.5%) other methods. From the 210 studies targeting ‘protocol development or optimization’, 133 (63.3%) applied qPCRs, 65 studies (30.9%) applied metabarcoding, 9 studies (4.3%) applied ddPCRs and 22 studies (10.5%) applied other methods. Regarding the 76 studies targeting ‘understanding the “ecology” of eDNA’, 52 studies (68.4%) applied qPCRs, 16 studies (21.1%) applied metabarcoding, 8 studies (10.5%) applied ddPCRs and 4 studies (5.3%) applied other methods (Fig. 5), revealing a significant difference in the frequencies of methods applied among the three study goal categories (Pearson's Chi-squared test, χ2 = 27.73, df = 8, P < 0.001).
Temporal trends in the methods applied in the surveyed studies. Dots represent percentage of total studies of the respective publication year. Lines represent regression curves from linear modelling and symbols to the right-hand side of the lines indicate direction and significance of respective slope coefficients (+ : positive, − : negative, n.s.: not significant, P: P value). Shadings indicate 95% confidence intervals of linear models
Target taxa and target systems
The majority of the 381 reviewed studies targeted ‘fishes’ (192 studies—50.4%), followed by ‘molluscs’ (43 studies—11.3%), ‘amphibians’ (27 studies—7.1%) and ‘crustaceans’ (6.8%). The remainder of target taxa all represented less than 5% each in the overall composition of target taxa (Fig. 6a). No significant difference regarding target taxa was observed across the 5 years (Pearson's Chi-squared test, χ2 = 54.9 df = 52, P = 0.365). ‘Biodiversity monitoring’ was the most-frequent study goal in most of the study taxa, except for ‘reptiles’, ‘macroinvertebrates’, ‘molluscs’ and ‘multiple’, for which ‘protocol development or optimization’ was the most-frequent goal. ‘Understanding the “ecology” of eDNA’ constituted a smaller fraction of the study goals for all study taxa, with ‘mammals’ and ‘vertebrates’ featuring this study goal not at all (Fig. S2a, Online Resource 3). Regarding target system of the surveyed studies, the most-frequent systems were ‘rivers’ (190 studies—49.9%), followed by ‘lakes/ponds’ (46 studies—12.1%) and ‘lakes/ponds & rivers combined’ (12.1%). For 20 studies (5.2%) a definite study system was not definable, and the remaining categories all constituted less than 5% each of the overall studies (Fig. 6b). No clear difference regarding composition of target system across the 5 years could be observed (Pearson's Chi-squared test, χ2 = 41.94, df = 40, P = 0.387). Biodiversity monitoring was the most frequently addressed study goal in seven out of the 10 target system categories, whereas ‘protocol development or optimization’ was the most-frequent study goal in three of these categories. ‘Understanding the “ecology” of eDNA’ was the least frequently listed study goal for all systems, with three system categories not addressing this goal at all (Fig. 2b, Online Resource 3).
Study design aspects
On average, the vast majority of the surveyed studies were designed in a descriptive manner (334 studies—87.7%) whereas 62 studies (16.3%) had a manipulative design. There was a weak tendency towards an increase of the proportion of descriptive studies throughout the years (Fig. 7a, F-statistic = 6.59, P = 0.083) as well as a decrease in the proportion of manipulative studies (F-statistic = 5.92, P = 0.093). Most of the 381 studies (344 studies—90.3%) utilized in-field samples whereas only a small fraction (37 studies—9.7%) used samples from a controlled environment or were of purely theoretical nature. No temporal trend could be observed concerning those two proportions (F-statistic = 0.587 and 1.028, respectively, both P > 0.1, Fig. 7b). Finally, out of the 248 studies targeting biodiversity monitoring, 131 studies (52.8%) applied only eDNA-based methods, whereas 117 studies (47.2%) applied both, eDNA-based and conventional or alternative methods in parallel. Again, no temporal trend could be observed concerning those two proportions (both F-statistic = 0.618, both P > 0.1, Fig. 7c).
Temporal trends concerning study design aspects in the surveyed studies. Dots represent percentage of total studies in the respective publication year. Lines represent regression curves from linear modelling and symbols to the right-hand side of the lines indicate direction and significance of respective slope coefficients (+ : positive, − : negative, n.s.: not significant, P: P value). Shadings indicate 95% confidence intervals of linear models. a Descriptive versus manipulative study design, b samples stem from a laboratory setting versus samples stem from in-field sampling, c within ‘biodiversity monitoring’ studies only: Methods applied were solely eDNA-based versus a combined approach also including conventional methods
Discussion
The number of publications meeting the search criteria increased throughout the last 5 years reflecting the well-documented ongoing growth in the field of eDNA research (e.g. Compson et al., 2020; Beng and Corlett, 2020; Mathieu et al., 2020). ‘Biodiversity monitoring’ was the most-frequent study goal in the publications reviewed. The relative frequency of studies targeting this goal significantly increased throughout the years surveyed, potentially indicating a shift towards more true application of eDNA-based methods in freshwater monitoring. In parallel, the absolute number of studies featuring ‘understanding the “ecology” of eDNA’ as a study goal remained relatively constant, causing a significant decrease in the relative frequency of this goal. Given that this aspect of eDNA research has gained interest only recently and has been advocated as an important issue to be addressed in eDNA-based assessments (Barnes & Turner, 2016; Harrison et al., 2019; Jo et al., 2021a), this decrease should not be interpreted as a loss of interest in this facet of eDNA research but potentially as a risk of this aspect falling behind in the fast-growing field of eDNA-based assessments. Jo et al. (2021b) not only describe an increase of studies investigating eDNA dynamics in their literature review but also highlight the low frequency of such studies compared to eDNA studies in general. As the application of eDNA-based methods continue to expand also into new study systems, the lag of studies investigating eDNA dynamics in these systems might hamper our ability to draw meaningful conclusions in such systems. Therefore, active research on this aspect is still highly needed. Finally, studies targeting ‘protocol development or optimization’ grew at a similar rate as the total number of publications in general, revealing a constant relative rate of methodological advancements. However, the proportion of studies incorporating protocol development varied greatly among target taxa and study systems (see Online Resource 3). More than half of the studies featuring ‘protocol development or optimization’ (119 studies—56.7%) combined these efforts with a biodiversity monitoring assay, very often in the form of developing a taxon-specific qPCR protocol and then applying this protocol in a survey to detect the taxon in the field. As the application of eDNA-based methods is still expanding across the tree of life, this trend is expected to continue in the future.
The geographic bias towards the global North of the returned studies of the literature search has been described in similar contexts before (Belle et al., 2019; Coble et al., 2019; Weigand et al., 2019) and was suggested to result from socioeconomic constraints and potentially the lack of the required infrastructure complicating eDNA implementation in global South countries (Belle et al., 2019). Additionally, this study revealed that studies targeting ‘protocol development or optimization’ or ‘understanding the “ecology” of eDNA’ are even more severely underrepresented in the global South or tropics. As mentioned before, the dynamics of eDNA might differ drastically among study systems and across climatic zones, and therefore, target-system-specific assessments are required (Harper et al., 2019; Sales et al., 2021). Such studies are more likely to be financed by fundamental research funds, and therefore, a bias for such research towards well-funded research centres in temperate climates in the global North may have evolved. However, it is essential to steer such research also towards yet understudied aquatic systems to better support biodiversity monitoring via eDNA-based assessments in these systems.
Among all eDNA-based approaches, metabarcoding has been particularly emphasized for its immense potential in biodiversity monitoring (Deiner et al., 2017; Ruppert et al., 2019; Compson et al., 2020) and the significant increase of studies utilizing metabarcoding described in this review might already reflect this. Furthermore, HTS costs continue to decrease (Hall, 2013; Wetterstrand, 2021) and new, smaller HTS sequencing platforms (such as the illumina iSeq™ or MiniSeq™ or Oxford Nanopore’s MinION) are being developed that could furthermore reduce total costs per survey for end-users. These advancements are likely to push metabarcoding for application in this field additionally.
In parallel, the application of qPCR showed a slight trend of decrease in frequency. Whether this tendency will get statistically better supported in the upcoming years needs to be evaluated. A potential increase in ddPCR application (e.g. if acquisition and running costs become competitive to those of qPCR) might restrain qPCR application in the future. However, in this study, the application of ddPCR did not reveal a statistically significant increase in frequency, yet. Other methods that are just being explored for application in eDNA-based research, such as CRISPR-Cas or metagenomics might need more time to be picked up for true application in biodiversity monitoring, but initial trials reveal already exciting prospects (Won et al., 2017; Garlapati et al., 2019; Williams et al., 2019b; Schroeter et al., 2020; Duran-Vinet et al., 2021).
The relative frequency of methods applied clearly varied among study goals. Metabarcoding was most frequently applied for ‘biodiversity monitoring’, probably once again underpinning its particular suitability for this goal. In contrast to that, qPCRs were more frequently applied in studies targeting ‘protocol development or optimization’ and ‘understanding the “ecology” of eDNA’. The former simply results from qPCRs protocols being mostly species specific, requiring the development of more protocols, in comparison to metabarcoding protocols, which are designed for whole species communities. Studies investigating the ‘ecology’ of eDNA often require quantification of DNA. Absolute quantification of target DNA in an eDNA sample still remains a challenging task in eDNA research. However, qPCR and ddPCR approaches are emerging to have a higher potential for reliable quantification, as metabarcoding has additional method-inherent complications for such an estimate (Evans et al., 2016; Sassoubre et al., 2016; Valentini et al., 2016; Deiner et al., 2017; Thalinger et al., 2018). The higher reliability of ddPCR for quantification might also be reflected in the relatively high frequency of this method in studies targeting ‘understanding of the “ecology” of eDNA’ compared to the other two study goals.
In this review, fishes were by far the most-frequent study taxa (being targeted by approximately half of the 381 studies surveyed), as described also in other meta-analyses (Belle et al., 2019; Coble et al., 2019). Drivers for this pattern, being observed also in freshwater research and conservation in general (Di Marco et al., 2017; Pérez-Espona, 2017), are undoubtedly socioeconomic interests but also numerous fish species either representing iconic flagship species, globally invasive species or keystone species for ecosystem service (Geist, 2015; Belle et al., 2019). The economic interests have potentially provided extra fuel for the rapid development of protocols targeting fishes (both, for metabarcoding and also species-specific assays) leading to more available protocols to date that allow a large application of such assays in biomonitoring studies. Furthermore, abundance estimations of fishes are often key parameters for many ecological indices calculated for freshwater habitats. This presumably led to an additional effort to improve protocols for density or abundance estimations of fishes.
The second most-frequent target taxon was molluscs. For these, the study motivation was either rooted in their susceptibility to habitat deterioration (e.g. Carlsson et al., 2017; Rodgers et al., 2020; Stoeckle et al., 2021) or in their role in ongoing biological invasions (e.g. De Ventura et al., 2017; Pie et al., 2017; Trebitz et al., 2019). The genus Dreissena, encompassing the highly invasive zebra and quagga mussels, was the most frequently targeted genus within molluscs (16 studies—37.2%) highlighting the effort to monitor and detect these invasive neobiota in many systems.
Regarding study systems, there was a clear domination of lotic systems, with rivers being the targeted system in approximately 50% of the surveyed studies. This predominance mirrors the general pattern of riverine systems being more frequently targeted in freshwater research or conservation in general (Wurtsbaugh et al., 2014).
It shall be noted that the trends described in this review are limited to the years 2017–2021, and therefore, previous research on specific taxa or systems (such as protocols being developed already earlier) might have influenced this pattern. However, other reviews encompassing larger timeframes (Beng & Corlett, 2020; Compson et al., 2020) show agreeing patterns in comparable aspects regarding target taxa and systems suggestion similar long-term trends. Fishes continue to be the most researched taxon group. This is expected to continue in the future due socioeconomic interets but also due to their key role in many monitoring schemes. However, as eDNA-based approaches do seem to be expanding more broadly across the tree of life, we might see a flattening of that monopoly in the future (which would be a desireable trend, see below).
The tendency towards more descriptive study designs throughout the last 5 years might also indicate a shift towards more frequent true application of eDNA-based studies is aquatic biomonitoring. Since biodiversity assessments are usually of descriptive design, this would be an expected concurring trend with an increase of studies featuring ‘biodiversity monitoring’ as a study goal. Interestingly, the frequency of purely laboratory-based or theoretical studies did not significantly decrease which would also be expected in line with this trend.
Among the studies featuring ‘biodivesity monitoring’ as their study goal, the fraction of studies applying both, eDNA-based and conventional methods did not change over time, remaining at around 50% of the studies. This might suggest a relative high frequency of ‘proof-of-concept’ studies that verify results retrieved by eDNA methodology using conventional methods (such as electrofishing or kicknet sampling and morphological identification). However, as been highlighted previously, eDNA-based or molecular tools in general are not intended to completely replace established or conventional methods but to complement them (Hering et al., 2018; Harper et al., 2019; Ruppert et al., 2019; Beng & Corlett, 2020). Both, molecular and conventional methods can elucidate different aspects of biodiversity that the other cannot. For example, eDNA can have higher detection rates for invasive species in biological invasions (Dejean et al., 2012; Bylemans et al., 2016), but so far eDNA-based methods are not able to give meaningful information about important demographic parameters of a population, such as age structure (Stoeckle et al., 2016; Pont et al., 2021). Therefore, a parallel application of molecular and conventional methods for biodiversity assessments is expected to continue also in the future.
Conclusion
This systematic literature review confirmed the ongoing rapid growth of eDNA-focused literature in freshwaters systems. Furthermore, the relative frequency of true application of eDNA-based methods for biodiversity monitoring did increase significantly over the last 5 years, potentially indicating a current phase of transition towards these methods becoming a standard tool for biodiversity monitoring. As fishes and riverine systems are the most frequently target taxa and systems, respectively, it is expected that true application of eDNA-based methods will first happen on a large scale for these systems with the other taxa and systems following after them.
A remaining challenge for broad implementation of these methods into legislative frameworks will be the need of standardization among methods, reliable abundance estimations and incorporation of demographic parameters in such assessments, all of which are currently being worked on actively (Goldberg et al., 2016; Pawlowski et al., 2018; Shu et al., 2020; Bohmann et al., 2021).
A critical pattern revealed in this study is the great variability in research efforts directed at different taxonomic groups, with fishes standing out as the main group of interest. However, for holistic biodiversity assessments, all target taxa need to be detected with equal certainty, and therefore, equal effort should be put into the development of methodology and detection assays. Thus, targeted promotion of the research into less tested, but often more biodiverse bioindicator groups is highly needed.
Finally, resilience is important for any monitoring approach. As shown, eDNA protocol development is still ongoing in all target taxa and systems. There is, thus, a risk that protocols being currently adopted in biomonitoring schemes will potentially be proven inferior to newer protocols at a later stage. As mentioned above, any change in methodology during an ongoing biomonitoring assessment might compromise data comparability. Therefore, managers and researchers should ensure that the chosen protocol meets certain minimum standards required for their research question or monitoring goal and can, therefore, be maintained for longer timeframes.
Taken together, eDNA-based approaches are a powerful addition to the biomonitoring toolbox and their uptake for biodiversity assessments is evident and increasing. However, stakeholders should be aware that new insights are still being gained and advances are continually made. In this transitional phase, it is then particularly critical for practitioners to ensure data consistency and comparability for long-term and/or large-scale biomonitoring assessments.
Combined with the positive assets of established methods, eDNA-based methods will definitely boost our efficiency to monitor freshwater biodiversity, enabling managers to quickly react to negative trends in these systems and to ensure healthy and functioning freshwater habitats also in the future.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Allan, E. A., A. F. Govindarajan, W. G. Zhang & A. C. Lavery, 2020. Environmental DNA shedding and decay rates from diverse animal forms and thermal regimes. Environmental DNA. https://doi.org/10.1002/edn3.141.
Anderson, S. R. & L. R. Thompson, 2021. Optimizing an enclosed bead beating extraction method for microbial and fish environmental DNA. Environmental DNA. https://doi.org/10.1002/edn3.251.
Ardura, A., A. Zaiko, Y. J. Borrell, A. Samuiloviene & E. Garcia-Vazquez, 2017. Novel tools for early detection of a global aquatic invasive, the zebra mussel Dreissena polymorpha. Aquatic Conservation-Marine and Freshwater Ecosystems 27: 165–176.
Barnes, M. A. & C. R. Turner, 2016. The ecology of environmental DNA and implications for conservation genetics. Conservation Genetics 17: 1–17.
Barnes, M. A., C. R. Turner, C. L. Jerde, M. A. Renshaw, W. L. Chadderton & D. M. Lodge, 2014. Environmental conditions influence eDNA persistence in aquatic systems. Environmental Science and Technology 48: 1819–1827.
Belle, C. C., B. C. Stoeckle & J. Geist, 2019. Taxonomic and geographical representation of freshwater environmental DNA research in aquatic conservation. Aquatic Conservation: Marine and Freshwater Ecosystems. https://doi.org/10.1002/aqc.3208.
Beng, K. C. & R. T. Corlett, 2020. Applications of environmental DNA (eDNA) in ecology and conservation: opportunities, challenges and prospects. Biodiversity and Conservation 29: 2089–2121.
Biggs, J., N. Ewald, A. Valentini, C. Gaboriaud, T. Dejean, R. A. Griffiths, J. Foster, J. W. Wilkinson, A. Arnell, P. Brotherton, P. Williams & F. Dunn, 2015. Using eDNA to develop a national citizen science-based monitoring programme for the great crested newt (Triturus cristatus). Biological Conservation 183: 19–28.
Birk, S., W. Bonne, A. Borja, S. Brucet, A. Courrat, S. Poikane, A. Solimini, W. Van De Bund, N. Zampoukas & D. Hering, 2012. Three hundred ways to assess Europe’s surface waters: an almost complete overview of biological methods to implement the Water Framework Directive. Ecological Indicators 18: 31–41.
Bista, I., G. R. Carvalho, M. Tang, K. Walsh, X. Zhou, M. Hajibabaei, S. Shokralla, M. Seymour, D. Bradley, S. Liu, M. Christmas & S. Creer, 2018. Performance of amplicon and shotgun sequencing for accurate biomass estimation in invertebrate community samples. Molecular Ecology Resources 18: 1020–1034.
Bohan, D. A., C. Vacher, A. Tamaddoni-Nezhad, A. Raybould, A. J. Dumbrell, & G. Woodward, 2017. Next-generation global biomonitoring: large-scale, automated reconstruction of ecological networks. Trends in Ecology & Evolution 32: 477–487, https://www.sciencedirect.com/science/article/pii/S0169534717300605.
Bohmann, K., P. Chua, L. E. Holman & C. Lynggaard, 2021. DNAqua-Net conference unites participants from around the world with the quest to standardize and implement DNA-based aquatic biomonitoring. Environmental DNA. https://doi.org/10.1002/edn3.207.
Bruce, K., R. Blackman, S. J. Bourlat, A. M. Hellström, J. Bakker, I. Bista, K. Bohmann, A. Bouchez, R. Brys, K. Clark, V. Elbrecht, S. Fazi, V. Fonseca, B. Hänfling, F. Leese, E. Mächler, A. R. Mahon, K. Meissner, K. Panksep, J. Pawlowski, P. Schmidt Yáñez, M. Seymour, B. Thalinger, A. Valentini, P. Woodcock, M. Traugott, V. Vasselon, & K. Deiner, 2021. A practical guide to DNA-based methods for biodiversity assessment. Advanced Books. Pensoft Publishers, https://ab.pensoft.net/article/68634/.
Bylemans, J., E. M. Furlan, L. Pearce, T. Daly & D. M. Gleeson, 2016. Improving the containment of a freshwater invader using environmental DNA (eDNA) based monitoring. Biological Invasions 18: 3081–3089.
Cantera, I., K. Cilleros, A. Valentini, A. Cerdan, T. Dejean, A. Iribar, P. Taberlet, R. Vigouroux & S. Brosse, 2019. Optimizing environmental DNA sampling effort for fish inventories in tropical streams and rivers. Scientific Reports. https://doi.org/10.1038/s41598-019-39399-5.
Cardas, J. B., D. Deconinck, I. Marquez, P. P. Torre, E. Garcia-Vazquez & G. Machado-Schiaffino, 2020. New eDNA based tool applied to the specific detection and monitoring of the endangered European eel. Biological Conservation. https://doi.org/10.1016/j.biocon.2020.108750.
Carlsson, J. E. L., D. Egan, P. C. Collins, E. D. Farrell, F. Igoe & J. Carlsson, 2017. A qPCR MGB probe based eDNA assay for European freshwater pearl mussel (Margaritifera margaritifera L.). Aquatic Conservation-Marine and Freshwater Ecosystems 27: 1341–1344.
Carøe, C. & K. Bohmann, 2020. Tagsteady: A metabarcoding library preparation protocol to avoid false assignment of sequences to samples. Molecular Ecology Resources 20: 1620–1631.
Coble, A. A., C. A. Flinders, J. A. Homyack, B. E. Penaluna, R. C. Cronn & K. Weitemier, 2019. eDNA as a tool for identifying freshwater species in sustainable forestry: a critical review and potential future applications. Science of the Total Environment 649: 1157–1170.
Compson, Z. G., B. McClenaghan, G. A. C. Singer, N. A. Fahner & M. Hajibabaei, 2020. Metabarcoding from microbes to mammals: comprehensive bioassessment on a global scale. Frontiers in Ecology and Evolution 8: 1–19.
Cowart, D. A., K. R. Murphy, & C.-H. C. Cheng, 2018. Metagenomic sequencing of environmental DNA reveals marine faunal assemblages from the West Antarctic Peninsula. Marine Genomics 37: 148–160, https://www.sciencedirect.com/science/article/pii/S1874778717302581.
Cristescu, M. E. & P. D. N. Hebert, 2018. Uses and misuses of environmental DNA in biodiversity science and conservation. Annual Review of Ecology, Evolution, and Systematics 49: 209–230.
De Ventura, L., K. Kopp, K. Seppaelae & J. Jokela, 2017. Tracing the quagga mussel invasion along the Rhine river system using eDNA markers: early detection and surveillance of invasive zebra and quagga mussels. Management of Biological Invasions 8: 101–112.
Deiner, K. & F. Altermatt, 2014. Transport distance of invertebrate environmental DNA in a natural river. PLoS ONE. https://doi.org/10.1371/journal.pone.0088786.
Deiner, K., H. M. Bik, E. Mächler, M. Seymour, A. Lacoursière-Roussel, F. Altermatt, S. Creer, I. Bista, D. M. Lodge, N. de Vere, M. E. Pfrender & L. Bernatchez, 2017. Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Molecular Ecology 26: 5872–5895.
Dejean, T., A. Valentini, C. Miquel, P. Taberlet, E. Bellemain & C. Miaud, 2012. Improved detection of an alien invasive species through environmental DNA barcoding: the example of the American bullfrog Lithobates catesbeianus. Journal of Applied Ecology 49: 953–959.
Di Marco, M., S. Chapman, G. Althor, S. Kearney, C. Besancon, N. Butt, J. M. Maina, H. P. Possingham, K. Rogalla von Bieberstein, O. Venter, & J. E. M. Watson, 2017. Changing trends and persisting biases in three decades of conservation science. Global Ecology and Conservation 10: 32–42, https://www.sciencedirect.com/science/article/pii/S2351989417300148.
Diana, A., E. Matechou, J. E. Griffin, A. S. Buxton & R. A. Griffiths, 2021. An RShiny app for modelling environmental DNA data: accounting for false positive and false negative observation error. Ecography 44: 1–7.
Doi, H., I. Katano, Y. Sakata, R. Souma, T. Kosuge, M. Nagano, K. Ikeda, K. Yano & K. Tojo, 2017. Detection of an endangered aquatic heteropteran using environmental DNA in a wetland ecosystem. Royal Society Open Science 4(7): 170568.
Doi, H., T. Takahara, T. Minamoto, S. Matsuhashi, K. Uchii & H. Yamanaka, 2015. Droplet digital polymerase chain reaction (PCR) outperforms real-time PCR in the detection of environmental DNA from an invasive fish species. Environmental Science and Technology 49: 5601–5608.
Dudgeon, D., A. H. Arthington, M. O. Gessner, Z. I. Kawabata, D. J. Knowler, C. Lévêque, R. J. Naiman, A. H. Prieur-Richard, D. Soto, M. L. J. Stiassny & C. A. Sullivan, 2006. Freshwater biodiversity: Importance, threats, status and conservation challenges. Biological Reviews of the Cambridge Philosophical Society 81: 163–182.
Duran-Vinet, B., K. Araya-Castro, T. C. Chao, S. A. Wood, V. Gallardo, K. Godoy & M. Abanto, 2021. Potential applications of CRISPR/Cas for next-generation biomonitoring of harmful algae blooms: a review. Harmful Algae. https://doi.org/10.1016/j.hal.2021.102027.
Eichmiller, J. J., S. E. Best & P. W. Sorensen, 2016. Effects of temperature and trophic state on degradation of environmental DNA in lake water. Environmental Science and Technology 50: 1859–1867.
Elberri, A. I., A. Galal-Khallaf, S. E. Gibreel, S. F. El-Sakhawy, I. El-Garawani, S.E.-S.H. Elnabi & K. Mohammed-Geba, 2020. DNA and eDNA-based tracking of the North African sharptooth catfish Clarias gariepinus. Molecular and Cellular Probes. https://doi.org/10.1016/j.mcp.2020.101535.
Elbrecht, V. & F. Leese, 2017. Validation and development of COI metabarcoding primers for freshwater macroinvertebrate bioassessment. Frontiers in Environmental Science 5: 1–11.
Esri, 2021. World Continents. https://www.arcgis.com/home/item.html?id=a3cb207855b348a297ab85261743351d.
Evans, N. T., B. P. Olds, M. A. Renshaw, C. R. Turner, Y. Li, C. L. Jerde, A. R. Mahon, M. E. Pfrender, G. A. Lamberti & D. M. Lodge, 2016. Quantification of mesocosm fish and amphibian species diversity via environmental DNA metabarcoding. Molecular Ecology Resources 16: 29–41.
Furlan, E. M., D. G. Eeson, C. Wisniewski, J. Yick & R. P. Duncan, 2019. eDNA surveys to detect species at very low densities: a case study of European carp eradication in Tasmania, Australia. Journal of Applied Ecology 56: 2505–2517.
Garlapati, D., B. Charankumar, K. Ramu, P. Madeswaran & M. V. Ramana Murthy, 2019. A review on the applications and recent advances in environmental DNA (eDNA) metagenomics. Reviews in Environmental Science and Bio/technology 18: 389–411.
Geist, J., 2015. Seven steps towards improving freshwater conservation. Aquatic Conservation: Marine and Freshwater Ecosystems John Wiley & Sons, Ltd 25: 447–453.
Giguet-Covex, C., G. F. Ficetola, K. Walsh, J. Poulenard, M. Bajard, L. Fouinat, P. Sabatier, L. Gielly, E. Messager, A. L. Develle, F. David, P. Taberlet, E. Brisset, F. Guiter, R. Sinet & F. Arnaud, 2019. New insights on lake sediment DNA from the catchment: importance of taphonomic and analytical issues on the record quality. Scientific Reports 9: 14676.
Goldberg, C. S., C. R. Turner, K. Deiner, K. E. Klymus, P. F. Thomsen, M. A. Murphy, S. F. Spear, A. McKee, S. J. Oyler-McCance, R. S. Cornman, M. B. Laramie, A. R. Mahon, R. F. Lance, D. S. Pilliod, K. M. Strickler, L. P. Waits, A. K. Fremier, T. Takahara, J. E. Herder & P. Taberlet, 2016. Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods in Ecology and Evolution 7: 1299–1307.
Griffiths, N. P., J. D. Bolland, R. M. Wright, L. A. Murphy, R. K. Donnelly, H. V. Watson & B. Hänfling, 2020. Environmental DNA metabarcoding provides enhanced detection of the European eel Anguilla anguilla and fish community structure in pumped river catchments. Journal of Fish Biology 97: 1375–1384.
Hall, N., 2013. After the gold rush. Genome Biology 14: 115.
Harper, L. R., A. S. Buxton, H. C. Rees, K. Bruce, R. Brys, D. Halfmaerten, D. S. Read, H. V. Watson, C. D. Sayer, E. P. Jones, V. Priestley, E. Mächler, C. Múrria, S. Garcés-Pastor, C. Medupin, K. Burgess, G. Benson, N. Boonham, R. A. Griffiths, L. Lawson Handley & B. Hänfling, 2019. Prospects and challenges of environmental DNA (eDNA) monitoring in freshwater ponds. Hydrobiologia 826: 25–41.
Harrison, J. B., J. M. Sunday & S. M. Rogers, 2019. Predicting the fate of eDNA in the environment and implications for studying biodiversity. Proceedings of the Royal Society b: Biological Sciences Royal Society. https://doi.org/10.1098/rspb.2019.1409.
Hering, D., A. Borja, J. I. Jones, D. Pont, P. Boets, A. Bouchez, K. Bruce, S. Drakare, B. Hänfling, M. Kahlert, F. Leese, K. Meissner, P. Mergen, Y. Reyjol, P. Segurado, A. Vogler & M. Kelly, 2018. Implementation options for DNA-based identification into ecological status assessment under the European Water Framework Directive. Water Research 138: 192–205.
Hindson, B. J., K. D. Ness, D. A. Masquelier, P. Belgrader, N. J. Heredia, A. J. Makarewicz, I. J. Bright, M. Y. Lucero, A. L. Hiddessen, T. C. Legler, T. K. Kitano, M. R. Hodel, J. F. Petersen, P. W. Wyatt, E. R. Steenblock, P. H. Shah, L. J. Bousse, C. B. Troup, J. C. Mellen, D. K. Wittmann, N. G. Erndt, T. H. Cauley, R. T. Koehler, A. P. So, S. Dube, K. A. Rose, L. Montesclaros, S. Wang, D. P. Stumbo, S. P. Hodges, S. Romine, F. P. Milanovich, H. E. White, J. F. Regan, G. A. Karlin-Neumann, C. M. Hindson, S. Saxonov, & B. W. Colston, 2011. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Analytical chemistry American Chemical Society 83: 8604–8610. https://pubmed.ncbi.nlm.nih.gov/22035192.
Hunter, M. E., R. M. Dorazio, J. S. S. Butterfield, G. Meigs-Friend, L. G. Nico & J. A. Ferrante, 2017. Detection limits of quantitative and digital PCR assays and their influence in presence–absence surveys of environmental DNA. Molecular Ecology Resources 17: 221–229.
Hunter, M. E., J. A. Ferrante, G. Meigs-Friend & A. Ulmer, 2019. Improving eDNA yield and inhibitor reduction through increased water volumes and multi-filter isolation techniques. Scientific Reports 9: 1–9.
Corp, I. B. M., 2016. IBM SPSS Statistics for Windows, IBM Corp., Armonk, NY:
Ito, K. & H. Shibaike, 2021. Use of environmental DNA to survey the distribution of the invasive mussel Limnoperna fortunei in farm ponds. Plankton & Benthos Research 16: 100–108.
Jane, S. F., T. M. Wilcox, K. S. Mckelvey, M. K. Young, M. K. Schwartz, W. H. Lowe, B. H. Letcher & A. R. Whiteley, 2015. Distance, flow and PCR inhibition: EDNA dynamics in two headwater streams. Molecular Ecology Resources 15: 216–227.
Jerde, C. L., W. L. Chadderton, A. R. Mahon, M. A. Renshaw, J. Corush, M. L. Budny, S. Mysorekar & D. M. Lodge, 2013. Detection of Asian carp DNA as part of a Great Lakes basin-wide surveillance program. Canadian Journal of Fisheries and Aquatic Sciences 70: 522–526.
Jo, T., S. Ikeda, A. Fukuoka, T. Inagawa, J. Okitsu, I. Katano, H. Doi, K. Nakai, H. Ichiyanagi & T. Minamoto, 2021. Utility of environmental DNA analysis for effective monitoring of invasive fish species in reservoirs. Ecosphere. https://doi.org/10.1002/ecs2.3643.
Jo, T., K. Takao & T. Minamoto, 2021b. Linking the state of environmental DNA to its application for biomonitoring and stock assessment: Targeting mitochondrial/nuclear genes, and different DNA fragment lengths and particle sizes. Environmental DNA. https://doi.org/10.1002/edn3.253.
Kumar, G., E. Farrell, A. M. Reaume, J. A. Eble & M. R. Gaither, 2021. One size does not fit all: tuning eDNA protocols for high- and low-turbidity water sampling. Environmental DNA. https://doi.org/10.1002/edn3.235.
Ladell, B. A., L. R. Walleser, S. G. McCalla, R. A. Erickson & J. J. Amberg, 2019. Ethanol and sodium acetate as a preservation method to delay degradation of environmental DNA. Conservation Genetics Resources 11: 83–88.
Leese, F., F. Altermatt, A. Bouchez, T. Ekrem, D. Hering, K. Meissner, P. Mergen, J. Pawlowski, J. Piggott, F. Rimet, D. Steinke, P. Taberlet, A. Weigand, K. Abarenkov, P. Beja, L. Bervoets, S. Björnsdóttir, P. Boets, A. Boggero, A. Bones, Á. Borja, K. Bruce, V. Bursić, J. Carlsson, F. Čiampor, Z. Čiamporová-Zatovičová, E. Coissac, F. Costa, M. Costache, S. Creer, Z. Csabai, K. Deiner, Á. DelValls, S. Drakare, S. Duarte, T. Eleršek, S. Fazi, C. Fišer, J.-F. Flot, V. Fonseca, D. Fontaneto, M. Grabowski, W. Graf, J. Guðbrandsson, M. Hellström, Y. Hershkovitz, P. Hollingsworth, B. Japoshvili, J. Jones, M. Kahlert, B. Kalamujic Stroil, P. Kasapidis, M. Kelly, M. Kelly-Quinn, E. Keskin, U. Kõljalg, Z. Ljubešić, I. Maček, E. Mächler, A. Mahon, M. Marečková, M. Mejdandzic, G. Mircheva, M. Montagna, C. Moritz, V. Mulk, A. Naumoski, I. Navodaru, J. Padisák, S. Pálsson, K. Panksep, L. Penev, A. Petrusek, M. Pfannkuchen, C. Primmer, B. Rinkevich, A. Rotter, A. Schmidt-Kloiber, P. Segurado, A. Speksnijder, P. Stoev, M. Strand, S. Šulčius, P. Sundberg, M. Traugott, C. Tsigenopoulos, X. Turon, A. Valentini, B. van der Hoorn, G. Várbíró, M. Vasquez Hadjilyra, J. Viguri, I. Vitonytė, A. Vogler, T. Vrålstad, W. Wägele, R. Wenne, A. Winding, G. Woodward, B. Zegura, & J. Zimmermann, 2016. DNAqua-Net: Developing new genetic tools for bioassessment and monitoring of aquatic ecosystems in Europe. Research Ideas and Outcomes Pensoft Publishers 2: e11321. http://riojournal.com/articles.php?id=11321.
Leese, F., A. Bouchez, K. Abarenkov, F. Altermatt, Á. Borja, K. Bruce, T. Ekrem, F. Čiampor, Z. Čiamporová-Zaťovičová, F. O. Costa, S. Duarte, V. Elbrecht, D. Fontaneto, A. Franc, M. F. Geiger, D. Hering, M. Kahlert, B. Kalamujić Stroil, M. Kelly, E. Keskin, I. Liska, P. Mergen, K. Meissner, J. Pawlowski, L. Penev, Y. Reyjol, A. Rotter, D. Steinke, B. van der Wal, S. Vitecek, J. Zimmermann & A. M. Weigand, 2018. Why we need sustainable networks bridging countries, disciplines, cultures and generations for Aquatic Biomonitoring 2.0: a perspective derived from the DNAqua-Net COST action. Advances in Ecological Research 58: 63–99.
Logan, J., J. M. J. Logan, K. J. Edwards, & N. A. Saunders, 2009. Real-time PCR: Current Technology and Applications. Caister Academic Press, https://books.google.at/books?id=YxGKpOg8TuQC.
Macher, T.-H., A. J. Beermann & F. Leese, 2021. TaxonTableTools: a comprehensive, platform-independent graphical user interface software to explore and visualise DNA metabarcoding data. Molecular Ecology Resources 21: 1705–1714.
Mächler, E., K. Deiner, P. Steinmann & F. Altermatt, 2014. Utility of environmental DNA for monitoring rare and indicator macroinvertebrate species. Freshwater Science 33: 1174–1183.
Maruyama, A., K. Sugatani, K. Watanabe, H. Yamanaka & A. Imamura, 2018. Environmental DNA analysis as a non-invasive quantitative tool for reproductive migration of a threatened endemic fish in rivers. Ecology and Evolution 8: 11964–11974.
Mathieu, C., S. M. Hermans, G. Lear, T. R. Buckley, K. C. Lee & H. L. Buckley, 2020. A systematic review of sources of variability and uncertainty in eDNA data for environmental monitoring. Frontiers in Ecology and Evolution 8: 1–14.
Miya, M., Y. Sato, T. Fukunaga, T. Sado, J. Y. Poulsen, K. Sato, T. Minamoto, S. Yamamoto, H. Yamanaka, H. Araki, M. Kondoh & W. Iwasaki, 2015. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. Royal Society Open Science. https://doi.org/10.1098/rsos.150088.
Olds, B. P., C. L. Jerde, M. A. Renshaw, Y. Li, N. T. Evans, C. R. Turner, K. Deiner, A. R. Mahon, M. A. Brueseke, P. D. Shirey, M. E. Pfrender, D. M. Lodge & G. A. Lamberti, 2016. Estimating species richness using environmental DNA. Ecology and Evolution 6: 4214–4226.
Pawlowski, J., L. Apothéloz-Perret-Gentil & F. Altermatt, 2020. Environmental DNA: What’s behind the term? Clarifying the terminology and recommendations for its future use in biomonitoring. Molecular Ecology 29: 4258–4264.
Pawlowski, J., M. Kelly-Quinn, F. Altermatt, L. Apothéloz-Perret-Gentil, P. Beja, A. Boggero, A. Borja, A. Bouchez, T. Cordier, I. Domaizon, M. J. Feio, A. F. Filipe, R. Fornaroli, W. Graf, J. Herder, B. van der Hoorn, J. Iwan Jones, M. Sagova-Mareckova, C. Moritz, J. Barquín, J. J. Piggott, M. Pinna, F. Rimet, B. Rinkevich, C. Sousa-Santos, V. Specchia, R. Trobajo, V. Vasselon, S. Vitecek, J. Zimmerman, A. Weigand, F. Leese & M. Kahlert, 2018. The future of biotic indices in the ecogenomic era: Integrating (e)DNA metabarcoding in biological assessment of aquatic ecosystems. Science of the Total Environment 637–638: 1295–1310.
Pérez-Espona, S., 2017. Conservation genetics in the European Union – biases, gaps and future directions. Biological Conservation 209: 130–136, https://www.sciencedirect.com/science/article/pii/S0006320716305183.
Pie, M. R., P. R. Stroher, A. O. Agostinis, R. Belmonte-Lopes, M. Z. Tadra-Sfeir & A. Ostrensky, 2017. Development of a real-time PCR assay for the detection of the golden mussel (Limnoperna fortunei, Mytilidae) in environmental samples. Anais da Academia Brasileira de Ciencias 89: 1041–1045.
Pont, D., M. Rocle, A. Valentini, R. Civade, P. Jean, A. Maire, N. Roset, M. Schabuss, H. Zornig & T. Dejean, 2018. Environmental DNA reveals quantitative patterns of fish biodiversity in large rivers despite its downstream transportation. Scientific Reports 8: 1–13.
Pont, D., A. Valentini, M. Rocle, A. Maire, O. Delaigue, P. Jean & T. Dejean, 2021. The future of fish-based ecological assessment of European rivers: from traditional EU Water Framework Directive compliant methods to eDNA metabarcoding-based approaches. Journal of Fish Biology 98: 354–366.
QGIS Development Team, 2019. QGIS. QGIS Geographic Information System. Open Source Geospatial Foundation Project., http://qgis.osgeo.org.
R Core Team, 2021. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria., https://www.r-project.org/.
Rodgers, T. W., J. C. Dysthe, C. Tait, T. W. Franklin, M. K. Schwartz & K. E. Mock, 2020. Detection of 4 imperiled western North American freshwater mussel species from environmental DNA with multiplex qPCR assays. Freshwater Science 39: 762–772.
Roy, M., V. Belliveau, N. E. Mandrak & N. Gagné, 2018. Development of environmental DNA (eDNA) methods for detecting high-risk freshwater fishes in live trade in Canada. Biological Invasions 20: 299–314.
Ruppert, K. M., R. J. Kline & M. S. Rahman, 2019. Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: A systematic review in methods, monitoring, and applications of global eDNA. Global Ecology and Conservation 17: 1–29.
Sales, N., O. Wangensteen, D. Carvalho, K. Deiner, K. Præbel, I. Coscia, A. D. Mcdevitt & S. Mariani, 2021. Space-time dynamics in monitoring neotropical fi sh communities using eDNA metabarcoding. Science of the Total Environment. https://doi.org/10.1016/j.scitotenv.2020.142096.
Sassoubre, L. M., K. M. Yamahara, L. D. Gardner, B. A. Block & A. B. Boehm, 2016. Quantification of environmental DNA (eDNA) shedding and decay rates for three marine fish. Environmental Science and Technology 50: 10456–10464.
Sato, Y., M. Miya, T. Fukunaga, T. Sado & W. Iwasaki, 2018. MitoFish and MiFish Pipeline: a mitochondrial genome database of fish with an analysis pipeline for environmental DNA metabarcoding. Molecular Biology and Evolution 35: 1553–1555.
Schenekar, T., M. Schletterer & S. J. Weiss, 2020. Development of a TaqMan qPCR protocol for detectingAcipenser ruthenusin the Volga headwaters from eDNA samples. CONSERVATION GENETICS RESOURCES 12: 395–397.
Schroeter, J. C., A. P. Maloy, C. B. Rees & M. L. Bartron, 2020. Fish mitochondrial genome sequencing: expanding genetic resources to support species detection and biodiversity monitoring using environmental DNA. Conservation Genetics Resources 12: 433–446.
Seymour, M., I. Durance, B. J. Cosby, E. Ransom-Jones, K. Deiner, S. J. Ormerod, J. K. Colbourne, G. Wilgar, G. R. Carvalho, M. de Bruyn, F. Edwards, B. A. Emmett, H. M. Bik & S. Creer, 2018. Acidity promotes degradation of multi-species environmental DNA in lotic mesocosms. Communications Biology Springer, US 1: 1–8.
Shaw, J. L. A., L. Weyrich & A. Cooper, 2017. Using environmental (e)DNA sequencing for aquatic biodiversity surveys: a beginner’s guide. Marine and Freshwater Research 68: 20–33.
Shu, L., A. Ludwig & Z. Peng, 2020. Standards for methods utilizing environmental dna for detection of fish species. Genes 11: 296.
Sigsgaard, E. E., H. Carl, P. R. Møller, & P. F. Thomsen, 2015. Monitoring the near-extinct European weather loach in Denmark based on environmental DNA from water samples. Biological Conservation 183: 46–52, https://www.sciencedirect.com/science/article/pii/S0006320714004480.
Stoeckle, B. C., S. Beggel, R. Kuehn & J. Geist, 2021. Influence of stream characteristics and population size on downstream transport of freshwater mollusk environmental DNA. Freshwater Science 40: 191–201.
Stoeckle, B. C., R. Kuehn & J. Geist, 2016. Environmental DNA as a monitoring tool for the endangered freshwater pearl mussel (Margaritifera margaritifera L.): a substitute for classical monitoring approaches? Aquatic Conservation: Marine and Freshwater Ecosystems 26: 1120–1129.
Taberlet, P., E. Coissac, M. Hajibabaei, & L. H. Rieseberg, 2012. Environmental DNA. Molecular Ecology 21: 1789–1793, http://onlinelibrary.wiley.com/doi/10.1111/j.1365-294X.2012.05542.x/full%5Cnpapers2://publication/uuid/3E2B6116-3B64-435F-A8EF-EFD5323DFD46.
Thalinger, B., D. Sint, C. Zeisler, D. Kirschner, R. Schwarzenberger, C. Moritz & M. Traugott, 2018. Quantification of fish populations using eDNA in alpine streams. Wasserwirtschaft 108: 30–34.
Thalinger, B., K. Deiner, L. R. Harper, H. C. Rees, R. C. Blackman, D. Sint, M. Traugott, C. S. Goldberg & K. Bruce, 2021. A validation scale to determine the readiness of environmental DNA assays for routine species monitoring. Environmental DNA 3: 823–836.
Thomsen, P. F. & E. Willerslev, 2015. Environmental DNA – an emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation 183: 4–18.
Trebitz, A. S., C. L. Hatzenbuhler, J. C. Hoffman, C. S. Meredith, G. S. Peterson, E. M. Pilgrim, J. T. Barge, A. M. Cotter & M. J. Wick, 2019. Dreissena veligers in western Lake Superior – inference from new low-density detection. Journal of Great Lakes Research 45: 691–699.
Tsuji, S., M. Ushio, S. Sakurai, T. Minamoto & H. Yamanaka, 2017. Water temperature-dependent degradation of environmental DNA and its relation to bacterial abundance. PLOS ONE. https://doi.org/10.1371/journal.pone.0176608.
Turner, C. R., K. L. Uy & R. C. Everhart, 2015. Fish environmental DNA is more concentrated in aquatic sediments than surface water. Biological Conservation 183: 93–102.
Valentini, A., P. Taberlet, C. Miaud, R. Civade, J. Herder, P. F. Thomsen, E. Bellemain, A. Besnard, E. Coissac, F. Boyer, C. Gaboriaud, P. Jean, N. Poulet, N. Roset, G. H. Copp, P. Geniez, D. Pont, C. Argillier, J. M. Baudoin, T. Peroux, A. J. Crivelli, A. Olivier, M. Acqueberge, M. Le Brun, P. R. Møller, E. Willerslev & T. Dejean, 2016. Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Molecular Ecology 25: 929–942.
Vasselon, V., F. Rimet, K. Tapolczai & A. Bouchez, 2017. Assessing ecological status with diatoms DNA metabarcoding: scaling-up on a WFD monitoring network (Mayotte island, France). Ecological Indicators Elsevier 82: 1–12.
Weigand, H., A. J. Beermann, F. Čiampor, F. O. Costa, Z. Csabai, S. Duarte, M. F. Geiger, M. Grabowski, F. Rimet, B. Rulik, M. Strand, N. Szucsich, A. M. Weigand, E. Willassen, S. A. Wyler, A. Bouchez, A. Borja, Z. Čiamporová-Zat’ovičová, S. Ferreira, K. Dijkstra, U. Eisendle, J. Freyhof, P. Gadawski, W. Graf, A. Haegerbaeumer, B. B. van der Hoorn, B. Japoshvili, L. Keresztes, E. Keskin, F. Leese, J. Macher, T. Mamos, G. Paz, V. Pešić, D. M. Pfannkuchen, M. A. Pfannkuchen, B. W. Price, B. Rinkevich, M. A. L. Teixeira, G. Várbíró, & T. Ekrem, 2019. DNA barcode reference libraries for the monitoring of aquatic biota in Europe: Gap-analysis and recommendations for future work. bioRxiv, https://www.biorxiv.org/content/10.1101/576553v1.
Wetterstrand, K., 2021. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP), www.genome.gov/sequencingcostsdata.
Wilcox, T. M., K. S. McKelvey, M. K. Young, A. J. Sepulveda, B. B. Shepard, S. F. Jane, A. R. Whiteley, W. H. Lowe & M. K. Schwartz, 2016. Understanding environmental DNA detection probabilities: a case study using a stream-dwelling char Salvelinus fontinalis. Biological Conservation 194: 209–216.
Wilcox, T. M., K. E. Zarn, M. P. Piggott, M. K. Young, K. S. McKelvey & M. K. Schwartz, 2018. Capture enrichment of aquatic environmental DNA: a first proof of concept. Molecular Ecology Resources 18: 1392–1401.
Williams, M.-A., J. O’Grady, B. Ball, J. Carlsson, E. de Eyto, P. McGinnity, E. Jennings, F. Regan & A. Parle-McDermott, 2019a. The application of CRISPR-Cas for single species identification from environmental DNA. Molecular Ecology Resources 19: 1106–1114.
Williams, M.-A., J. O’Grady, B. Ball, J. Carlsson, E. de Eyto, P. McGinnity, E. Jennings, F. Regan & A. Parle-McDermott, 2019b. The application of CRISPR-Cas for single species identification from environmental DNA. Molecular Ecology Resources 19: 1106–1114.
Wilson, J.-J., K.-W. Sing, P.-N. Chen & A. Zieritz, 2018. Tracking the southern river terrapin (Batagur affinis) through environmental DNA: prospects and challenges. Mitochondrial DNA Part A 29: 862–866.
Won, N. I., K. H. Kim, J. H. Kang, S. R. Park & H. J. Lee, 2017. Exploring the impacts of anthropogenic disturbance on seawater and sediment microbial communities in Korean Coastal waters using metagenomics analysis. International Journal of Environmental Research and Public Health 14: 130.
Wong, M.K.-S., M. Nakao & S. Hyodo, 2020. Field application of an improved protocol for environmental DNA extraction, purification, and measurement using Sterivex filter. Scientific Reports. https://doi.org/10.1038/s41598-020-77304-7.
Wood, S. A., L. Biessy, J. L. Latchford, A. Zaiko, U. von Ammon, F. Audrezet, M. E. Cristescu & X. Pochon, 2020. Release and degradation of environmental DNA and RNA in a marine system. Science of the Total Environment. https://doi.org/10.1016/j.scitotenv.2019.13531.
Wurtsbaugh, W. A., N. A. Heredia, B. G. Laub, C. S. Meredith, H. E. Mohn, S. E. Null, D. A. Pluth, B. B. Roper, W. Carl Saunders, D. K. Stevens, R. H. Walker & K. Wheeler, 2014. Approaches for studying fish production: do river and lake researchers have different perspectives? Canadian Journal of Fisheries and Aquatic Sciences 72: 149–160.
WWF, 2020. Living Planet Report 2020 – bending the curve of biodiversity loss. Gland, Switzerland.
Xu, N., B. Zhu, F. Shi, K. Shao, Y. Que, W. Li, W. Li, W. Jiao, H. Tian, D. Xu & J. Chang, 2018. Monitoring seasonal distribution of an endangered anadromous sturgeon in a large river using environmental DNA. Science of Nature. https://doi.org/10.1007/s00114-018-1587-4.
Acknowledgements
I would like to thank Pooja Singh, Tarik Bodasing, Till Macher and an anonymous reviewer for their constructive comments on previous versions of this manuscript.
Funding
Open access funding provided by University of Graz.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no known conflict of interest with any organization concerning the content in this manuscript.
Additional information
Guest editors: Koen Martens, Sidinei M. Thomaz, Diego Fontaneto & Luigi Naselli-Flores / Emerging Trends in Aquatic Ecology IV
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schenekar, T. The current state of eDNA research in freshwater ecosystems: are we shifting from the developmental phase to standard application in biomonitoring?. Hydrobiologia 850, 1263–1282 (2023). https://doi.org/10.1007/s10750-022-04891-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-022-04891-z